Trained-Rank-Pruning
PyTorch code for "Trained Rank Pruning for Efficient Neural Networks"
Our code is built based on bearpaw
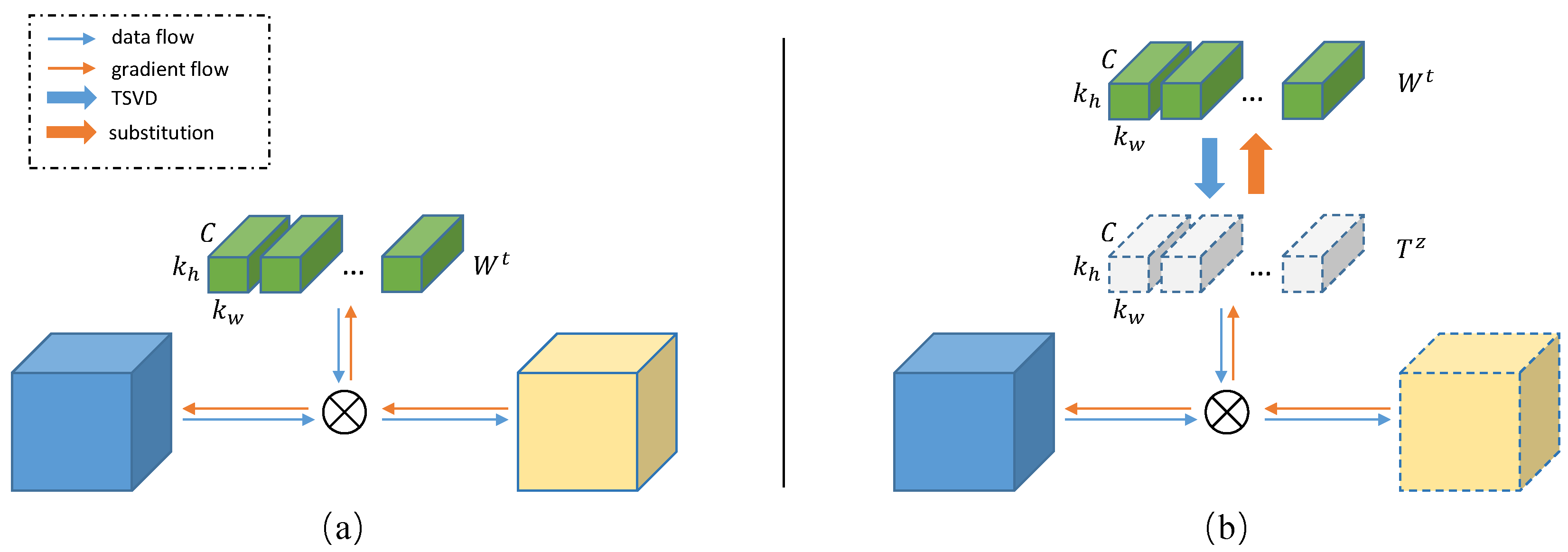
What's in this repo so far:
- TRP code for CIFAR-10 experiments
- Nuclear regularization code for CIFAR-10 experiments
Simple Examples
optional arguments:
-a model
--depth layers
--epoths training epochs
-c path to save checkpoints
--gpu-id specifiy using GPU or not
--nuclear-weight nuclear regularization parameterTraining ResNet-20 baseline:
python cifar-nuclear-regularization.py -a resnet --depth 20 --epochs 164 --schedule 81 122 --gamma 0.1 --wd 1e-4 --checkpoint checkpoints/cifar10/resnet-20
Training ResNet-20 with nuclear norm:
python cifar-nuclear-regularization.py -a resnet --depth 20 --epochs 164 --schedule 81 122 --gamma 0.1 --wd 1e-4 --checkpoint checkpoints/cifar10/resnet-20 --nuclear-weight 0.0003
Training ResNet-20 with TRP:
python cifar-TRP.py -a resnet --depth 20 --epochs 164 --schedule 81 122 --gamma 0.1 --wd 1e-4 --checkpoint checkpoints/cifar10/resnet-20 --nuclear-weight 0.0003
Decompose the trained model without retraining:
python cifar-nuclear-regularization.py.py -a resnet --depth 20 --resume checkpoints/cifar10/resnet-20/model_best.pth.tar --evaluate
Decompose the trained model with retraining:
python cifar-nuclear-regularization.py.py -a resnet --depth 20 --resume checkpoints/cifar10/resnet-20/model_best.pth.tar --evaluate --retrain
Notes
During decomposition, TRP using value threshold(very small value to truncate singular values) based strategy because the trained model is in low-rank format. Other methods including Channel or spatial-wise decomposition baseline methods use energy threshold.
Citation
If you think this work is helpful for your own research, please consider add following bibtex config in your latex file
@article{xu2018trained,
title={Trained Rank Pruning for Efficient Deep Neural Networks},
author={Xu, Yuhui and Li, Yuxi and Zhang, Shuai and Wen, Wei and Wang, Botao and Qi, Yingyong and Chen, Yiran and Lin, Weiyao and Xiong, Hongkai},
journal={arXiv preprint arXiv:1812.02402},
year={2018}
}