dwindling-reads [--csv] --stage <stage1> <files> [--stage <stage2> <files> [--stage <stage3> <files>]
dwindling-reads --help
This program collates summary information from a set of next-gen sequencing paired-end reads at each stage of your pipeline.
Each stage is specified by --stage, followed by two to four arguments. The first argument is an arbitrary name of your choosing to identify that stage. Both FastQ (uncompressed and gzipped) and BAM files are supported. FastQ stages must specify at least two files for forward and reverse reads. An optional third file may be specified for single-end orphans, such as those commonly produced by quality trimmers. BAM stages must specify only a single .bam file.
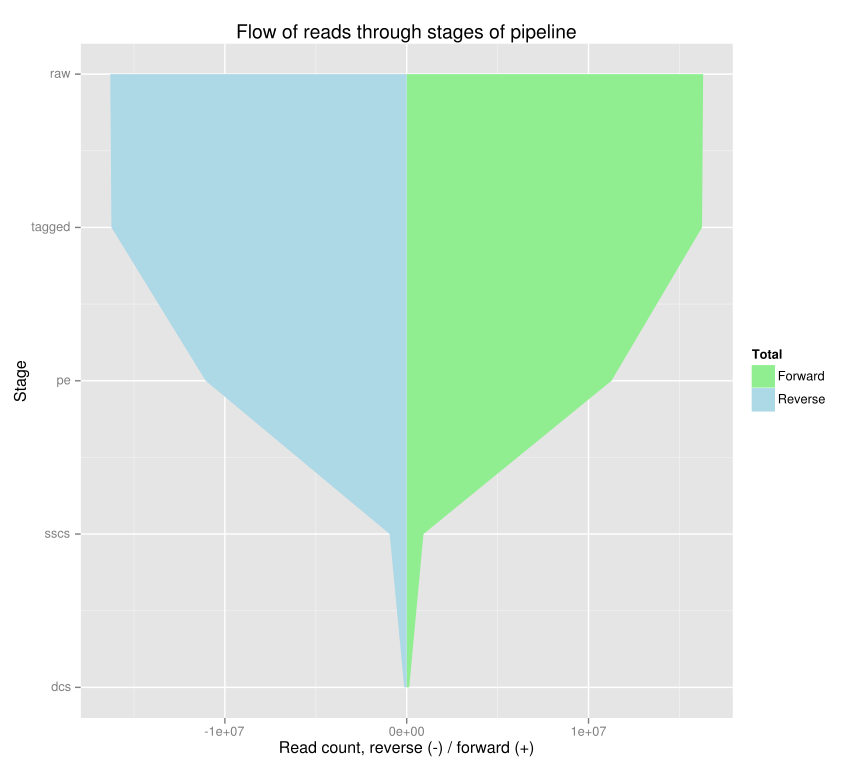
CSV output may be piped into dwindling-plot to produce an SVG plot of the read counts through the various stages.
--csv output CSV instead of formatted text
--help print usage message and exit
dwindling-reads --stage raw raw/ABC_R1.fq.gz raw/ABC_R2.fq.gz \
--stage sickle trimmed/ABC_R1.fq trimmed/ABC_R2.fq trimmed/ABC_orphans.fq \
--stage mapped ABC.bam
196300 raw = 98150 forward + 98150 reverse
161640 sickle = 80820 forward + 80820 reverse (= 82.34% of raw)
6278 single-end orphans (= 3.20% of raw)
34660 discarded, including orphans (= 17.66% of raw)
161910 mapped = 80820 forward + 80820 reverse (= 100.17% of sickle)
-270 discarded (= -0.17% of sickle)
(In this case, bwa mem found 270 reads with secondary mappings so the reads are growing not dwindling!)
Requires:
- Perl 5.14 or newer
- zcat (distributed with gzip)
- wc (distributed with coreutils)
- samtools
The first three are generally pre-installed on most Linux/Unix operating systems.
In addition, several Perl libraries from CPAN are needed and
described in the cpanfile. The quickest way to satisfy the Perl
prerequisites is by running:
make bundle
from within a clone of the git repository. This command will download
cpanm and use it to install the
required dependencies into the inc/ directory. The dwindling-reads program
will automatically use libraries in this directory if they exist.
Requires:
You can generally just install the two R packages like so:
Rscript -e 'install.packages(commandArgs(T))' ggplot2 Cairo
To install them system-wide, use sudo Rscript.