serofoi is an R package to estimate the Force-of-Infection (FoI) of a given pathogen from age-disaggregated population-based cross-sectional serosurveys, using a Bayesian framework. The package provides a set of features for assessing model fitting, convergence and visualisation.
serofoi relies on the
rstan package, which
provides an R interface for the Stan programming language for
statistical Bayesian modelling. Particularly, serofoi relies on
the use of a Hamiltonian Monte Carlo (HMC) algorithm implemented by
Stan for Markov chain Monte Carlo (MCMC) sampling. The implemented
methods are outlined in (Cucunubá et al. 2017) and
(Carrera et al. 2020) (see FoI
Models
for further details)
serofoi is part of the Epiverse Initiative.
You can install the development version of serofoi from GitHub with:
if(!require("pak")) install.packages("pak")
pak::pak("epiverse-trace/serofoi")serofoi provides a minimal serosurvey dataset, serodata, that
can be used to test out the package.
# Load example dataset chagas2012 included with the package
data(chagas2012)
head(chagas2012, 5)
#> survey total counts age_min age_max tsur country test antibody
#> 1 COL-035-93 34 0 1 1 2012 COL ELISA IgG anti-T.cruzi
#> 2 COL-035-93 25 0 2 2 2012 COL ELISA IgG anti-T.cruzi
#> 3 COL-035-93 35 1 3 3 2012 COL ELISA IgG anti-T.cruzi
#> 4 COL-035-93 29 0 4 4 2012 COL ELISA IgG anti-T.cruzi
#> 5 COL-035-93 36 0 5 5 2012 COL ELISA IgG anti-T.cruziThe function prepare_serodata will prepare the entry data for the use
of the modelling module; this function computes the sample size, the
years of birth and the binomial confidence interval for each age group
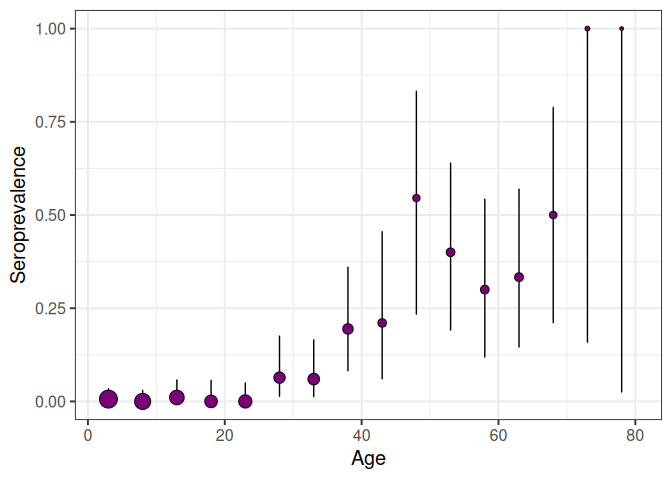
in the provided dataset. A visualisation of the prepared seroprevalence
data can be obtained using the function plot_seroprev:
serodata_test <- prepare_serodata(chagas2012)
plot_seroprev(serodata_test, size_text = 15)Contributors to the project include:
-
Zulma M. Cucunubá (author, maintainer)
-
Nicolás Torres (author)
-
Ben Lambert (author)
-
Pierre Nouvellet (author)
-
Geraldine Gómez (contributor)
-
Jaime A. Pavlich-Mariscal (contributor)
-
Miguel Gamez (contributor)
-
Hugo Gruson (contributor)
-
David Santiago Quevedo (contributor)
-
Everlyn Kamau (contributor)
-
Richard Creswell (contributor)
-
Sumali Bajaj (contributor)
More details on how to use serofoi can be found in the online documentation as package vignettes, under Get Started, An Introduction to FoI Models and Real-life Use Cases for serofoi
To report a bug please open an issue.
Contributions to serofoi are welcomed. Please follow the package contributing guide.
Please note that the serofoi project is released with a Contributor Code of Conduct. By contributing to this project, you agree to abide by its terms.
Carrera, Jean-Paul, Zulma M. Cucunubá, Karen Neira, Ben Lambert, Yaneth Pittí, Jesus Liscano, Jorge L. Garzón, et al. 2020. “Endemic and Epidemic Human Alphavirus Infections in Eastern Panama: An Analysis of Population-Based Cross-Sectional Surveys.” The American Journal of Tropical Medicine and Hygiene 103 (6): 2429–37. https://doi.org/10.4269/ajtmh.20-0408.
Cucunubá, Zulma M, Pierre Nouvellet, Lesong Conteh, Mauricio Javier Vera, Victor Manuel Angulo, Juan Carlos Dib, Gabriel Jaime Parra -Henao, and María Gloria Basáñez. 2017. “Modelling Historical Changes in the Force-of-Infection of Chagas Disease to Inform Control and Elimination Programmes: Application in Colombia.” BMJ Global Health 2 (3): e000345. https://doi.org/10.1136/bmjgh-2017-000345.