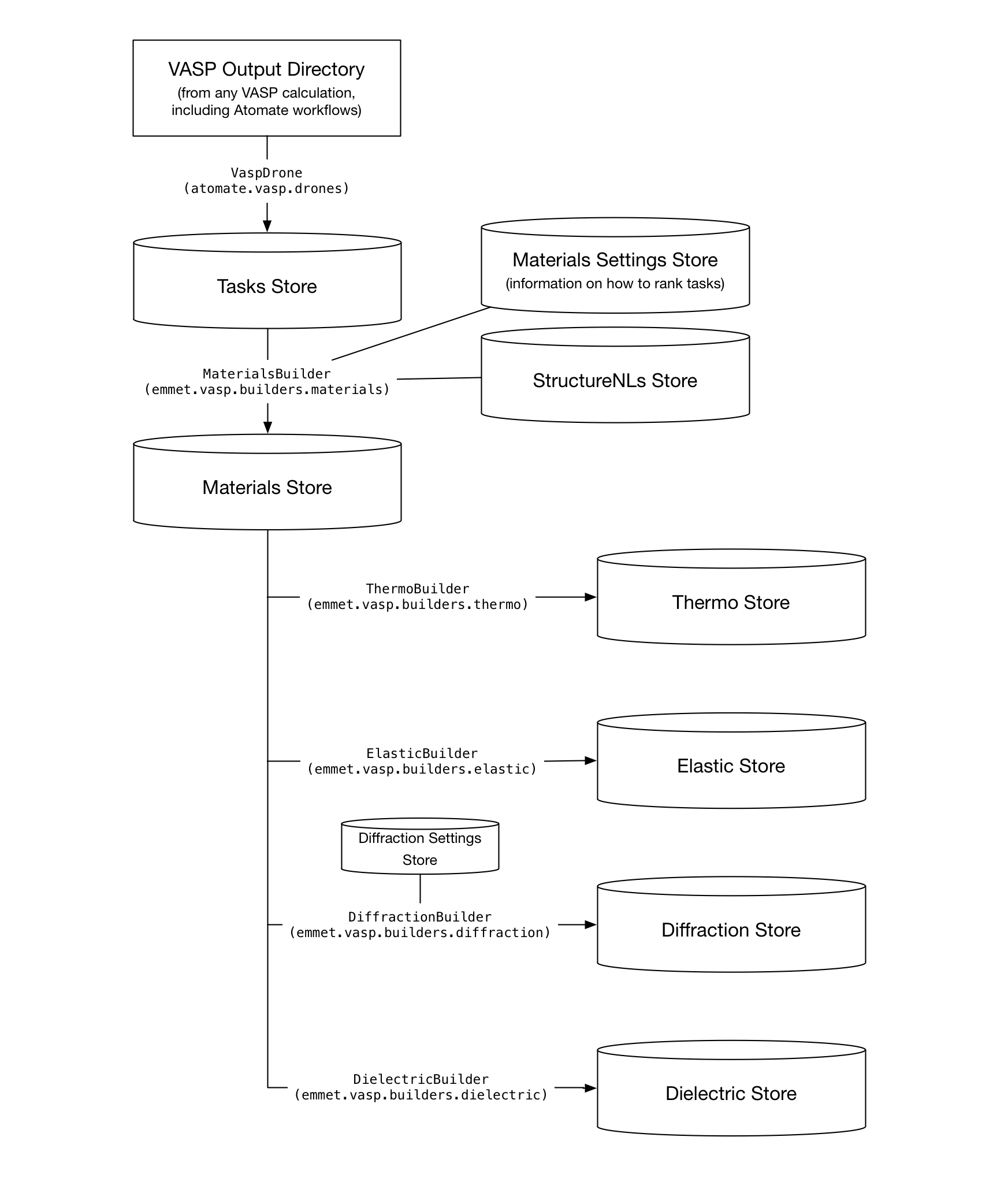
The purpose of Emmet is to 'build' collections of materials properties from the output of computational materials calculations. Currently, the effective purpose of Emmet is to take the output of VASP electronic structure calculations to build MongoDB collections that back the Materials Project website and its apps.
Emmet uses Maggma, our more general aggregation framework which abstracts away the behind-the-scenes machinery: Maggma provides our Builder class and a general interface to Stores, which can be MongoDB collections or plain JSON files.
The Builder takes source Store(s), processes items in that store, and then builds results to target Store(s).
To ease debugging, in Emmet data flows in one direction only: this means that each Store is only built by a specific builder, and will not then be modified by successive builders.
Builders are designed to be run periodically and automatically: as such, Stores have a 'last updated' filter (lu_filter) so that we only attempt to process new entries in the Store.
Emmet is currently primarily an internal Materials Project tool so if you're reading this you might have just joined the group, in which case: welcome! :-)
Emmet is on PyPI, so pip install emmet should work. However, it is currently in very active development, so an editable install is recommended for now:
pip install -e git+git@github.com:materialsproject/emmet.git#egg=emmet --src .Here is a sample script for running the MaterialsBuilder. Replace database information as appropriate (this assumes a test database running on localhost with a pre-populated tasks collection, with mat.json in the working directory).
#!/usr/bin/env python
from maggma.runner import Runner
from maggma.stores import MongoStore, JSONStore
from emmet.vasp.builders.materials import MaterialsBuilder
from emmet.vasp.builders.thermo import ThermoBuilder
tasks_store = MongoStore(database="test",
collection_name="materials",
host="localhost",
port=27017,
lu_field="last_updated")
materials_settings_store = JSONStore("mat.json")
materials_store = MongoStore(database="test",
collection_name="tasks",
host="localhost",
port=27017)
materials_builder = MaterialsBuilder(tasks_store,
materials_settings_store,
materials_store,
lu_field="last_updated")
runner = Runner([materials_builder])
runner.run()Take care to set the lu_field correctly: this is the key that the builder looks for to see when the document was last updated, and thus which new documents to build from. This field does not exist by default in MongoDB.
To run more than one builder, add:
thermo_store = MongoStore(database="test",
collection="thermo",
host="localhost",
port=27017)
thermo_builder = ThermoBuilder(materials_store,
thermo_store)and change runner = Runner([materials_builder]) to runner = Runner([materials_builder, thermo_builder]).
The list of builders can be provided in any order: their dependencies will be resolved intelligently and the Runner will run the builders in the correct order and in parallel if supported by the system.
Sub-class the Builder base class and implement the following methods:
get_items()– get your items to process, e.g. as a result of a running a query on your source(s)process_item()– for each of your items, do something, e.g. calculate a diffraction patternupdate_targets()– update your target(s) with your processed datafinalize()– optional, perform any final clean up (close database connections etc., the base class can handle this)
The DiffractionBuilder is a nice simple builder to copy from to get started.
The VASP builders all operate on a tasks Store which is parsed from any VASP calculation folder by Atomate's VaspDrone. Once the tasks Store has been created, Emmet's builders take over.
Source(s) tasks (typically tasks collection), material_settings (typically mat.json), snls (optional)
Target(s) materials (typically materials collection)
-
Filters to only include tasks that completed successfuly.
-
Groups tasks into those for the same structure.
Structure matching first only selects materials that have the same chemical formula, and then uses pymatgen's
StructureMatcherto perform symmetry analysis. -
For each property, ranks tasks for a given structure according to those that are expected to predict the property more accurately (for example, a band gap from a band structure calculation is ranked higher than a band gap from a generic calculation). This value is then picked as the canonical value for that property.
The
task_typeis already determined and comes from the tasks store, and the rankings are specified inmat.json. No attempt is made to rank which task of the sametask_typeis best; in this case it is assumed that the most recent calculation takes precendence. -
(Optional) The Structure Notation Language (or 'SNLs') provide a way to bundle a structure and its metadata (such as bibtex references for where the structure came from) within the Materials Project. This will lookup if there are existing SNL(s) for the structure, and assign an SNL accordingly.
Source(s) materials
Target(s) thermo
-
Groups materials into those in the same chemical system (that is, materials whose crystal structure contain the same elements).
-
Filters out materials that can not be directly compared to each other, e.g. they've been calculated by different methods such that their total energies are on different scales.
By default, this is done by using
MaterialsProjectCompatibility('Advanced')in pymatgen, which intelligently mixes GGA and GGA+U calculations depending on the elements present, and performs corrections to the total energy as appropriate. -
Uses pymatgen's
phasediagrampackage to calculate the energy above hull for each material and, if the material is unstable, its decomposition pathway.
Source(s) materials
Target(s) elastic
- Selects an initial structure from materials
- Finds deformed instances of this initial structure from materials, and calculates the deformation matrix
- If 6 independent deformations are found, calculates the elastic tensor using pymatgen's
ElasticTensor
Source(s) materials, xrd_settings (typically xrd.json)
Target(s) diffraction
- For each structure, calculates its ideal X-ray diffraction pattern for a variety of X-ray targets (specified in
xrd.json)
Source(s) tasks, materials
Target(s) toplogy, bader
-
For each structure in materials, calculates bonding from the material's crystal structure using a variety of methods (pymatgen's local_env and critic2's sum of atomic charge densities).
-
It then finds the task corresponding to a static calculation.
-
If
AECCAR0,AECCAR2,CHGCARare present, performs attempts to find bonding information using critic2 and also performs a bader analysis that is stored separately.