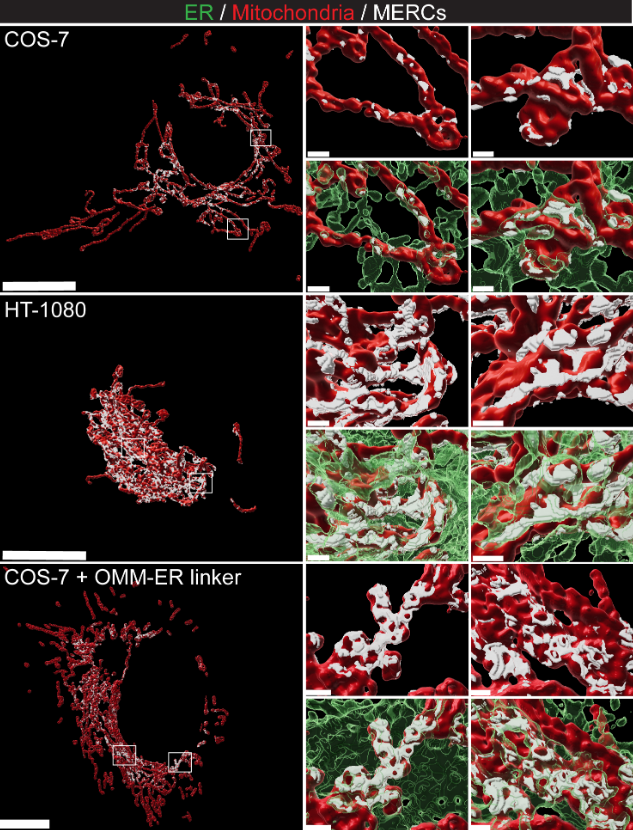
Detects sub-precision contacts between subcellular organelles in 2 and 3D STED (precision ~ 50-150nm) superresolution microscopy, for example endoplasmum reticulum and mitochondria (contacts ~ 20-100nm).
Where a pixel precise segmentation is not feasible due to the precision of the microscope, and colocalization does not describe the interface in a meaningful way, SubPrecisionContactDetection can reconstruct the plausible interface between the organelles.
An example rendering of the postprocessed contact zones (white) between endoplasmum reticulum (green) and mitochondria (red) is shown here (source):
- Fast: using multiple threads, and Julia's fast LLVM JIT code
- Reproducible: tests ensure backwards compatibility
- Configurable: Can process deconvolved or raw images, with optional extra denoising
- Rich: provides interpretable features for each detected contact
This project is developed using Julia. For ease of use and to maximize reproducibility we also provide container images using Singularity.
This project was developed on Linux, and deployed on scientific computing clusters running Linux. The Singularity workflow ensures both local and cluster computations run exactly the same. The automated tests at CircleCI](https://dl.circleci.com/status-badge/redirect/gh/bencardoen/SubPrecisionContactDetection.jl/tree/main) run the exact same environment.
This cannot be guaranteed across different OS'es (e.g. Windows, MacOs). While there are no technical reasons preventing the code from working on any OS, you may run into issues as it is not something we actively use ourselves.
You can use an optimized Singularity image, which has all dependencies pre-installed.
If you do not have Singularity, please see the documentation for detailed installation instructions.
The below steps are examples, but may not be complete for each platform, for the reference instructions, please visit installation instructions.
Fedora/RPM
sudo dnf install singularityTo run Singularity on Windows, set up WSL2 or refer to installation instructions.
See instructions.
Download the image as mcsdetect.sif. For example, using wget (Linux), you could do:
wget -O mcsdetect.sif http://vault.sfu.ca/index.php/s/QJ4Evcet4oVWXPL/downloadOn MacOS you can install wget using:
brew install wgetFirst, make sure execute permissions are set:
chmod u+x mcsdetect.sif./mcsdetect.sifExpected output:
chmod u+x mcsdetect.sif
./mcsdetect.sif -e 'using SubPrecisionContactDetection;'Expected output:
chmod u+x mcsdetect.sif
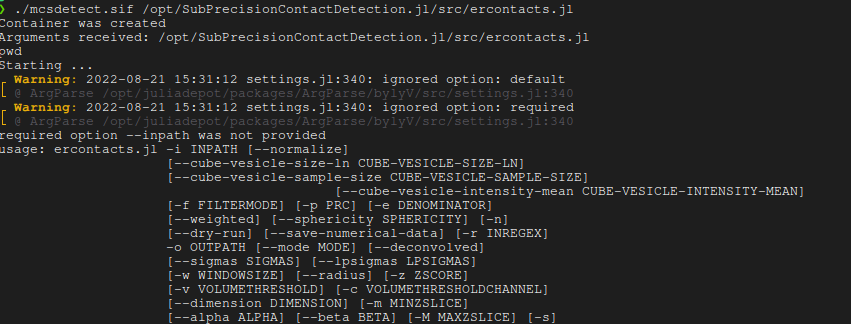
./mcsdetect.sif /opt/SubPrecisionContactDetection.jl/scripts/ercontacts.jl ARGSWhere you'd replace ARGS with arguments to the script as documented in scripts/ercontacts.jl. Run it without arguments to get the help prompt.
Note due to a bug with conda MacOS installations will have some tests failing, the module itself is functional
You can either add to the global Julia installation:
julia -e 'using Pkg;Pkg.add(url="https://github.com/bencardoen/ERGO.jl.git");Pkg.add(url="https://github.com/bencardoen/SPECHT.jl.git");Pkg.add(url="https://github.com/bencardoen/SubPrecisionContactDetection.jl.git")'
julia -e 'using Pkg; Pkg.build("SubPrecisionContactDetection");Pkg.test("SubPrecisionContactDetection")'Or create a new environment and install it there:
mkdir -p test
cd test
julia --project=. -e 'using Pkg;Pkg.add(url="https://github.com/bencardoen/ERGO.jl.git");Pkg.add(url="https://github.com/bencardoen/SPECHT.jl.git");Pkg.add(url="https://github.com/bencardoen/SubPrecisionContactDetection.jl.git")'
julia --project=. -e 'using Pkg; Pkg.build("SubPrecisionContactDetection");Pkg.test("SubPrecisionContactDetection")'In both cases, you should see that all tests pass:
git clone https://github.com/bencardoen/SubPrecisionContactDetection.jl.git
cd SubPrecisionContactDetection.jl
julia --project=. installlocal.jlThis should result in output similar to this screenshot:
The command line interface does the heavy lifting for you:
Using the singularity image not only saves you from dependency tracking, it also is precompiled, making it x5 - x10 faster. This is especially true on clusters where the speedup can be even larger.
./mcsdetect.sif opt/SubPrecisionContactDetection/scripts/ercontacts.jl --inpath ./in -r "*[1,2].tif" -w 2 --deconvolved --sigmas 2.5-2.5-1.5 --outpath ./out --alpha 0.01 --beta 0.01 -c 1 -v 2000 --mode=deconjulia --project=. ./scripts/ercontacts.jl --inpath ./in -r "*[1,2].tif" -w 2 --deconvolved --sigmas 2.5-2.5-1.5 --outpath ./out --alpha 0.01 --beta 0.01 -c 1 -v 2000 --mode=decon 2>&1 | tee -a log_test.txtWhere:
- --{in|out}path : directories where tif files can be found
- -r : regex to tif files, e.g. *[1,2].tif indicates channel 1 and 2 will have filenames ending in 1,2.tif respectively.
- -w : windowsize, >1 or higher
- --sigmas : smoothing Gaussian, set < precision
- --alpha : max false positive rate (p-value), 0.05 is a common value.
- --beta : max false negative rate (stat. power) 0.05 implies 95% stat power.
- -c 1: postprocess channel 1
- -v 2000: drop all contacts touching objects in channel 1 with volume < 2000
- --mode=decon : input are non deconvolved tiff files
- 2>&1 | tee -a log_test.txt : save any output to log.txt (in addition to showing it in stdout)
The output should look like:
- skeleton_contacts.tif
- channel_[1,2].tif
- (non)_vesicle_contacts: mitochondria (channel 1) with volume < 2000 are considered vesicles, split contacts so you can visualize them separately
- channel_1_(non_)vesicle.tif : channel 1 objects (mitochondria) split into < 2000 and > 2000 objects
- raw|gradient|eroded.tif : stages of progressively computed contacts, all but 'eroded' are debug output
- *.csv : features for each contact
In scripts/run_cube_sampling_on_dataset.jl you'll find a script that samples contacts with a sliding window, to avoid long tail statistics dominating the conclusion of any analysis. The paper goes into more depth why this is beneficial.
See hpcscripts/arraysbatch.sh for an example parameter sweep on a large set of cells. Assuming you created inlists.txt and outlists.txt, you'd submit to SLURM.
sbatch hpcscripts/arraysbatch.shPlease edit and revise before you submit, e.g. your email and cluster account need to change at a minimum.
If you find this project useful, please cite
@article {Cardoen2022.06.23.497346,
author = {Cardoen, Ben and Gao, Guang and Vandevoorde, Kurt R. and Alan, Parsa and Liu, William and Vogl, A. Wayne and Hamarneh, Ghassan and Nabi, Ivan R.},
title = {Automatic sub-precision membrane contact site detection identifies convoluted tubular riboMERCs},
elocation-id = {2022.06.23.497346},
year = {2022},
doi = {10.1101/2022.06.23.497346},
publisher = {Cold Spring Harbor Laboratory},
URL = {https://www.biorxiv.org/content/early/2022/06/26/2022.06.23.497346},
eprint = {https://www.biorxiv.org/content/early/2022/06/26/2022.06.23.497346.full.pdf},
journal = {bioRxiv}
}If you have any issues, please create an issue.
Make sure to include:
- include OS, Julia version
- description of steps to reproduce
- be concise yet complete
Yes, if you clone the repository, and are using Linux, you need to do 2 things
- edit singularity_recipes/recipe.def
- execute buildimage # Needs sudo
./buildimage.shThis will rebuild the image, first checking out the latest version of the code.
Expected RAM usage for images of sizes 500x500x20 ~ 5GB RAM, 2000x2000x70: ~ 50GB RAM, and so on. By default, all of JULIA_NUM_THREADS cores will be used to run in parallel. > 8 is overkill, so set to 4-8 at most:
export JULIA_NUM_THREADS=4On desktops this is unlikely to be an issue, but on a cluster node with > 64 cores you will probably get a slowdown if you exceed 8-12 cores.
First, make sure you install and clone in a clean environment:
mdkir mydir
cd mydir
julia
julia> ]
(@v1.x) pkg> activate .
(@v1.x) pkg> updateDo not use Julia < 1.7, there's no guarantee that deprecated APIs will still work, and performance and user friendliness of the e.g. the package manager alone make 1.7 the ideal baseline.
Current memory usage is higher than it strictly needs to be because we generate a lot of intermediate steps. In principle we could reduce usage by x2 or more, but it would come at the cost of debugging/interpretability.
MacOS + Conda has a bug where a certificate error triggers a cascade of errors. The errors can be ignored, including the failing tests, this is an optional part of the module. When the bug in conda is resolved, this issue should be resolved as well.