EPIGRAPHHUB
Health Data Hub & Data Analysis Ecosystem for Rapid Epidemic Outbreak Response
Explore the docs »
View Platform
·
Report Bug
·
Request Feature
Table of Contents
About The Project
 EpigraphHub is an entirely open source Data integration platform focused on Epidemiological analyses.
EpigraphHub is an entirely open source Data integration platform focused on Epidemiological analyses.
Built With
EpigraphHub is built on top of the industry-leading opensource BI platform Apache Superset enriched by our own analytical libraries in Python and R.
Getting Started
To get a feel of EpigraphHub as a user, please check it out at Epigraphhub.org.
Prerequisites
Installation
Usage
Roadmap
See the open issues for a list of proposed features (and known issues).
Check our wiki for complementary informations
Contributing
Contributions are what make the open source community such an amazing place to be learn, inspire, and create. Any contributions you make are greatly appreciated. Contact us to get involved in the project and join our team on discord
- Fork the Project
- Create your Feature Branch (
git checkout -b feature/ProposedFeature) - Commit your Changes (
git commit -m 'Add some ProposedFeature') - Push to the Branch (
git push origin feature/ProposedFeature) - Open a Pull Request
Development
To create a new environment for EpiGraphHub, you can use it using conda or docker.
Also, ensure you have the .env file with the correct configuration.
You can copy its structure from .env.tpl.
Conda
If you don't have conda installed, you can install it from here. Micromamba is a miniconda version with mamba, a powerful installer that is much faster than the conda installer.
Create your new environtment running the following command:
$ mamba env create -n conda/prod.yamlTo activate your new environment with all the packages installed, run:
$ conda activate epigraphhubContainers
The project provides three compose files, where one is the base
definition (containers/compose-base.yaml) and the others are one for
development (containers/compose-dev.yaml) and other for production
(containers/compose-prod.yaml). Additionally, there is a Makefile
file that provides make targets that helps to manage the docker services:
containers-build: builds docker imagescontainers-start: starts docker servicescontainers-stop: stops docker servicescontainers-restart: restarts docker servicescontainers-logs: shows dockercontainers-dev-prepare-db: prepares the development version of the databasecontainers-run-cron: run the cron taskscontainers-bash: opens the docker service bash
This make targets runs by default docker for development. For production,
it is necessary to pass the argument ENV=prod , for example:
make containers-buildA common workflow to prepare the system would be:
- Build the docker image:
$ make containers-build- Start the services:
$ make containers-startBefore moving forward, check if the services are working properly:
$ ./containers/healthcheck.sh epigraphhub- Now, prepare the development database (skip it for production):
$ make containers-prepare-dbAIRFLOW
Create a FERNET KEY:
openssl rand -base64 32MinIO
EpiGraphHub uses MinIO as a cloud storage. MinIO is s3-ish local storage that offers an AWS S3 API. For development, it is already created by the containers stack.
In order to configure it locally, run:
make dev-create-s3-credentialsNote: it needs the AWS env variable on the environment (or from .env file).
Deployment
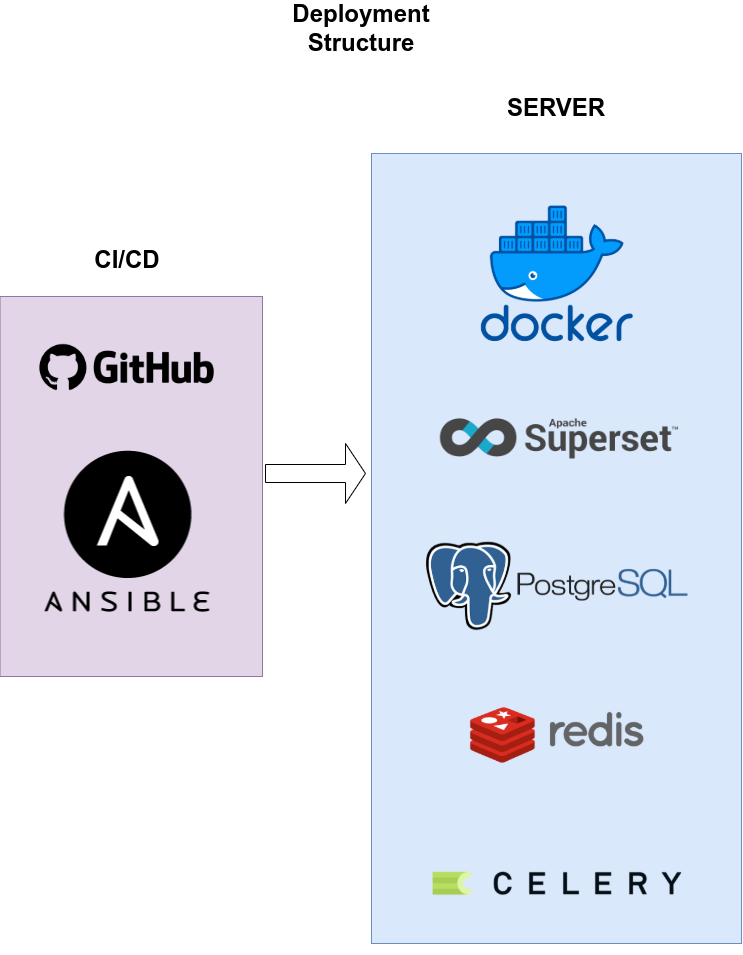
The deployment is executed by Ansible and triggered by GitHub Actions. When a new release is triggered, on GitHub Actions, there is a step there that uses Ansible to connect to the EpiGraphHub server and deploy the new version.
NOTE: Currently, the deployment process is under development.
The deployment is designed in the following structure:
All the services are being migrated to docker containers. It will allows us to isolate the services and keep the host server safer.
License
Distributed under the MIT License. See LICENSE for more information.
Contact
Project Link: https://github.com/thegraphnetwork/GRAPHnetEpiApp
Linked-in : https://www.linkedin.com/company/the-graph-network
Acknowledgements
TBD