-
The
stimulusschema is a self-contained application that generates and presents visual stimuli using Psychtoolbox and records conditions and trials in a DataJoint database. -
See the Element Visual Stimulus documentation for the background information and development timeline.
-
For more information on the DataJoint Elements project, please visit https://elements.datajoint.org. This work is supported by the National Institutes of Health.
Click to expand details
Requirements: Some of the stimuli require MATLAB R2016b+.
Although the following steps steps can be executed manually, they are typically automated and thus serve as the application interface for the experiment control software.
>> stimulus.open
Stimulus trials are generated and queued by the scripts in the +stimulus/+conf directory. You need to know which configuration script needs to be run.
For example, to prepare the grate stimulus, execute
>> stimulus.conf.grate
While the stimulus is loaded, you will see a sequence of dots . and asterisks *, which respectively indicate whether the conditions are computed anew or are loaded from the database. Some stimuli take a long time to compute and you might like to run the configuration before you begin the experiment. On subsequent runs, the computed stimuli will be loaded from the database and will not take as long.
The stimulus must be run for a specific scan in the experiment.Scan table.
Table experiment.Scan contains a dummy entry that can be used for testing. Its primary key is struct('animal_id', 0, 'session', 0, 'scan_idx', 0). During the experiment, the correct scan identification must be provided.
The following command will run the queued stimulus trials for the dummy scan.
>> stimulus.run(struct('animal_id', 0, 'session', 0, 'scan_idx', 0))
While the stimulus is playing, you can interrupt with Ctrl+c. The stimulus program will handle this event, cancel the ongoing trial, and clear the screen. To resume the stimulus, repeat the stimulus.run call above. Or to queue a new set of trials, run the configuration script again.
To close the stimulus program,
>> stimulus.close
Click to expand details
The stimulus configuration and playback are written and executed in MATLAB. However, the control software in our lab is written in Python.
First configure the MATLAB Engine API for Python as described at https://www.mathworks.com/help/matlab/matlab_external/install-the-matlab-engine-for-python.html.
Upon installation, you can reproduce the steps above in Python as
import matlab.engine as eng
mat = eng.start_matlab()
# step 1: Initialize screen
mat.stimulus.open(nargout=0)
# step 2: intialize conditions and queue trials
mat.stimulus.conf.grate(nargout=0)
# step 3: run stimulus for the specific scan
f = mat.stimulus.run(dict(animal_id=0, session=0, scan_idx=0), nargout=0, async=True)
# step 4. Interrupt and resume stimulus
f.cancel() # interrupt
f = mat.stimulus.run(dict(animal_id=0, session=0, scan_idx=0), nargout=0, async=True) # resume
# step 5. Exit
f.done() # True if stimulus is done
f.result() # waits until the stimulus is done
f.stimulus.close(nargout=0) # close the stimulus screen Click to expand details
The schema diagram below depicts the stimulus schema.
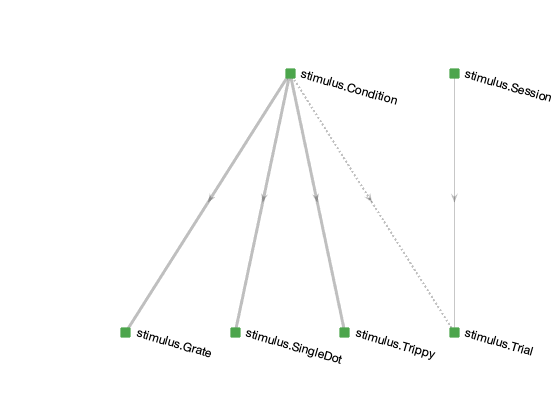
The central table is stimulus.Condition, which enumerates all possible stimulus conditions to be presented.
It is populated before the stimulus is presented for the first time.
The specialization tables below it contain parameters that are specific to each type of stimulus.
For example, stimulus.Monet2 contains parameters that are specific to a single stimulus condition of the type Monet2.
For each tuple in stimulus.Condition, exactly one of the specialization tables contains the corresponding entry.
The name of the specialization table is indicated in each row of stimulus.Condition in field stimulus_type.
A preview of the stimulus.Condition:
>> stimulus.Condition
ans =
Object stimulus.Condition
:: stimulus condition ::
CONDITION_HASH stimulus_type stimulus_version
______________________ ___________________ ________________
'+0cObnxIHpoB5RKZJVYj' 'stimulus.Matisse2' '1'
'+3o2cquPfKnts4Gmjwr4' 'stimulus.Matisse2' '1'
'+9mOEvwZHyV2MiwRBsMy' 'stimulus.Varma' '1'
'+9nMtSVLIPAj/VEmey+6' 'stimulus.Matisse' '2'
'+9OAgABcltcbRZUw77Kt' 'stimulus.Matisse2' '1'
'+A8FfGEWQNonM6RMmrTk' 'stimulus.Matisse2' '1'
'+C/KYdzQvzn0jzQScSGy' 'stimulus.Matisse2' '1'
'+cI6EqAdQgh2tyJ1eMzy' 'stimulus.Matisse' '2'
'+eFINMa+jF58wHzuk9qQ' 'stimulus.Monet2' 'dimitri-1'
'+eK4n7czWTGRVKKh4EJO' 'stimulus.Matisse2' '1'
'+f+o1UeO1AtHWPTo3vlc' 'stimulus.Matisse2' '1'
'+h1WWj2NG6mjGFobRphN' 'stimulus.Matisse2' '1'
'+HeQV7jovoXvymyCqYCP' 'stimulus.Matisse2' '1'
...
The table stimulus.Trial contains the information about the presentation of a condition during a specific scan (from experiment.Scan).
Any number of conditions of any type can be presented during a scan and each condition may be displayed multiple times.
>> stimulus.Trial
ans =
Object stimulus.Trial
:: visual stimulus trial ::
ANIMAL_ID SESSION SCAN_IDX TRIAL_IDX condition_hash last_flip trial_ts flip_times
_________ _______ ________ _________ ______________________ _________ _____________________ __________
0 0 0 0 'Qjz5gJN2igKvsonApHO1' 21322 '2017-04-21 16:23:40' '=BLOB='
0 0 0 1 'KMk2le1nd79vP4uhW+lG' 21324 '2017-04-21 16:23:42' '=BLOB='
0 0 0 2 'd3TMSkOO74Y2QzRngY9r' 21325 '2017-04-21 16:23:43' '=BLOB='
0 0 0 3 'EvIYjxUYNs2QOjjiXoFo' 21327 '2017-04-21 16:23:45' '=BLOB='
0 0 0 4 '8hPjQGXtiY7VJdmWBJhz' 21328 '2017-04-21 16:23:46' '=BLOB='
0 0 0 5 'koXklHGOKSXzqG4vgeKw' 21330 '2017-04-21 16:23:48' '=BLOB='
0 0 0 6 '9vYRXrkmZd1mi6oFcBAC' 21332 '2017-04-21 16:23:49' '=BLOB='
0 0 0 7 'Yj+hNG8q+V2Icr+GW5WT' 21333 '2017-04-21 16:23:51' '=BLOB='
0 0 0 8 'idP7ku8g2U51r28Hb6Nb' 21335 '2017-04-21 16:23:52' '=BLOB='
0 0 0 9 'mzrmJBasIquICcmTK8K7' 21336 '2017-04-21 16:23:54' '=BLOB='
0 0 0 10 'hp9b+CKb/QgV1O6B5gk+' 21338 '2017-04-21 16:23:55' '=BLOB='
0 0 0 11 'lA4QZVp/vl9JxiH3JH8Y' 21339 '2017-04-21 16:23:57' '=BLOB='
0 0 0 12 'H+w/IuPACeCuzUziOIfi' 21341 '2017-04-21 16:23:58' '=BLOB='
...
If the language is unspecified, the queries run in both MATLAB and Python.
visualScans = experiment.Scan() & stimulus.Trial()
monetScans = experiment.Scan() & (stimulus.Trial() * stimulus.Monet())
or
monetScans = experiment.Scan() & (stimulus.Trial() * stimulus.Condition() & 'stimulus_type="stimulus.Monet"')
# python
session_key = dict(session=7302)
scan_conditions = stimulus.Condition() & (stimulus.Trial() & session_key)% matlab
sessionKey = struct('session', 7302, 'session', 1, 'scan_idx', 3);
scanConditions = stimulus.Condition & (stimulus.Trial & sessionKey);-
If your work uses DataJoint and DataJoint Elements, please cite the respective Research Resource Identifiers (RRIDs) and manuscripts.
-
DataJoint for Python or MATLAB
-
Yatsenko D, Reimer J, Ecker AS, Walker EY, Sinz F, Berens P, Hoenselaar A, Cotton RJ, Siapas AS, Tolias AS. DataJoint: managing big scientific data using MATLAB or Python. bioRxiv. 2015 Jan 1:031658. doi: https://doi.org/10.1101/031658
-
DataJoint (RRID:SCR_014543) - DataJoint for
<Select Python or MATLAB>(version<Enter version number>)
-
-
DataJoint Elements
-
Yatsenko D, Nguyen T, Shen S, Gunalan K, Turner CA, Guzman R, Sasaki M, Sitonic D, Reimer J, Walker EY, Tolias AS. DataJoint Elements: Data Workflows for Neurophysiology. bioRxiv. 2021 Jan 1. doi: https://doi.org/10.1101/2021.03.30.437358
-
DataJoint Elements (RRID:SCR_021894) - Element Visual Stimulus (version
<Enter version number>)
-
