Vivarium E. coli is a port of the Covert Lab's E. coli Whole Cell Model (wcEcoli) to the Vivarium framework.
As in wcEcoli, raw experimental data (reconstruction/ecoli/flat) is first processed
by the parameter calculator or Parca (reconstruction/ecoli/fit_sim_data_1.py) to calculate
model parameters (e.g. transcription probabilities).
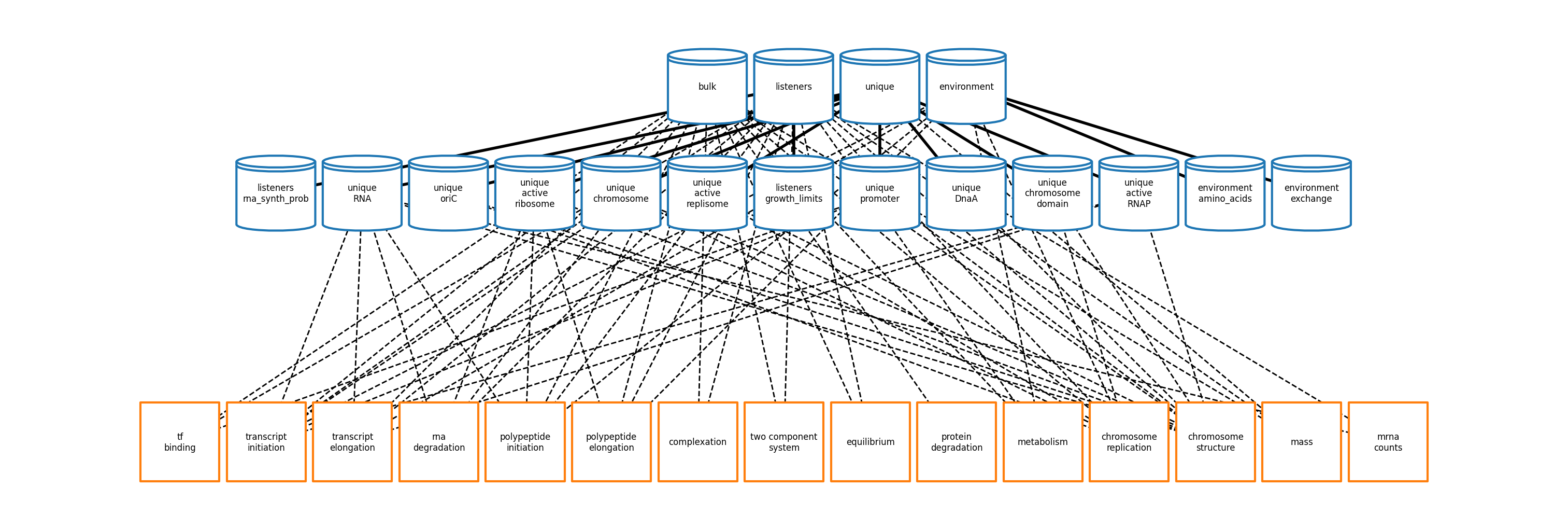
These parameters are used to configure processes (ecoli/processes) that are linked together
into a complete simulation in accordance with the topology generated by
ecoli/experiments/ecoli_master_sim.py.
For more details, refer to the user guide.
Note: The following instructions assume a Linux or MacOS system. Windows users can attempt to follow the same instructions after setting up Windows Subsystem for Linux.
pyenv lets you install and switch between multiple Python releases and multiple "virtual environments", each with its own pip packages. Using pyenv, create a virtual environment and install Python 3.11.3. For a tutorial on how to create a virtual environment, follow the instructions here and stop once you reach the "Create the 'wcEcoli3' python virtual environment" step. Then, run the following command in your terminal:
$ pyenv virtualenv 3.11.3 viv-ecoli && pyenv local viv-ecoli
Update pip, setuptools and wheel to avoid issues with these:
$ pip install --upgrade pip setuptools wheel
Now, install numpy (check requirements.txt for the exact version):
$ pip install numpy
Then install the remaining requirements:
$ pip install -r requirements.txt
And build the Cython components:
$ make clean compile
In order to save simulation data for downstream analyses, install and start MongoDB.
To run the simulation, set the PYTHONPATH environment variable to the cloned repository and run
ecoli_master_sim.py. If you are at the top-level of the cloned repository, invoke:
$ export PYTHONPATH=.
$ python ecoli/experiments/ecoli_master_sim.py
For details on configuring simulations through either the command-line interface or .json files, see the Ecoli-master configurations readme.
The main motivation behind the migration is to modularize the processes and allow them to be run in new contexts or reconfigured or included in different biological models.
There are three main aspects to the migration:
-
Decoupling each process from
sim_dataso they can be instantiated and run independently of the assumptions made in the structure of thesim_dataobject. This allows each process to be reconfigured into new models as well as ensuring each process has the ability to run on its own. Each process now has parameters declared for each class that can be provided at initialization time. -
Refactoring access to
Bulk-andUniqueMoleculesinto theports_schema()declaration used by Vivarium. This turns each process's state interactions into explicit declarations of properties and attributes. These declarations allow the processes to be recombined and to be expanded with new properties as needed. It also frees the processes from having to conform to theBulk/Uniquedichotomy and provides a means for other kinds of state (previously implemented by reading from listeners or accessing other process class's attributes), further promoting decoupling. This declared schema is then used as a structure to provide the current state of the simulation to each process'snext_update(timestep, states)function during the simulation. -
Translating all state mutation from inside the processes into the construction of a Vivarium
updatedictionary which is returned from thenext_update(timestep, states). The structure of this update mirrors the providedstatesdictionary and the declaredports_schema()structure. This allows each process to operate knowing that values will not be mutated by other processes before it sees them, a fundamental requirement for modular components.
The way Vivarium state updates work, all states are provided as they appear at the beginning of each process's update timestep, and then are applied at the end of that timestep. This ensures all states are synchronized between processes.
Note: Must be a member of the
CovertLabGitHub organization.
The Causality visualization tool can be used to manually examine the model's
output (molecule counts, reaction fluxes, etc.). First, run a simulation ensuring that data is set to be emitted to MongoDB (e.g. emitter is database in configuration). Then, build the Causality network with the following command:
python ecoli/analysis/buildCausalityNetwork.py
Then, clone the Causality repository into the same folder as the vivarium-ecoli repository (e.g. /dev/causality and /dev/vivarium-ecoli) and follow the installation instructions.
Finally, start the Causality server, which will open an interactive webpage in your web browser:
cd /dev/causality
python site/server.py ../vivarium-ecoli/out/seriesOut/