The objective is to create permutations of 10, 100 or 1000 colours. Similar colours being kept as close as possible. To do so, we will calculate the Euclidean distance of adjacent colours (using RGB values) and sum it. Our goal will be to minimise this sum.
We tried 4 heuristic algorithms in order to find the best solution:
- Random search: There is no move operator required for random search as we are not making any improvements, just generating random solution every time and keeping track of the minimum solution found till the end.
- Hill climber using a two-opt move: The two-opt move operator gets us close to the local optima in a very short period of time. This operator also minimises the number of iterations and computation time.
- Iterated local search using 2 two-opt moves: For my algorithm we performed two two opt move which is even more powerful. It increases a great number of chances for escaping the local optima and moving closer to Global optima.
- Evolutionary algorithm: The move operators that I chose in the algorithm are one point crossover for generating two children where we copy the first parts into both the children and the rest we inherit from the parents in each of the child skipping the values they already inherit. This move creates a necessary diversity in the children which gives novelty to every child. I chose to use two opt move for the mutation of the children.
- Write functions to read the file and calculate the sum of the Euclidean distance
- Write the algorithms as well as the perturbation functions (two-opt move, perturbation and crossover)
- Run each algorithm 20 times and report the mean, standard deviation and best fitness found + plot boxplots
- Plot line plots of the best fitness found depending on the number of iterations
- Visualize the sets of colours
- We can then make conclusions about the best algorithm depending on the requirements (processing time for example)
- Permutations for 100 colours and 2 algorithms
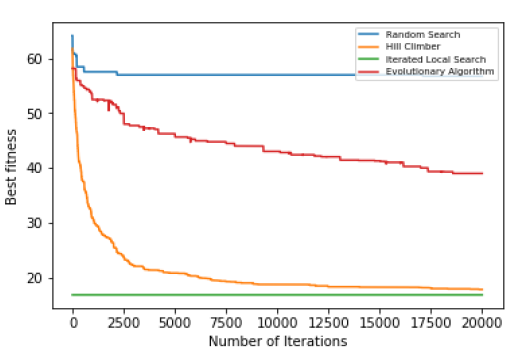
- Line plot for 100 colours
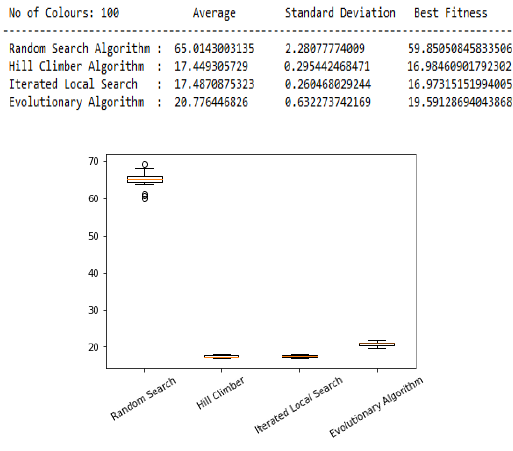
- Box plots for 100 colours
'''
Reading the file and calculate the sum of the Euclidean distance
'''
import matplotlib.pyplot as plt
import numpy as np
import re
from scipy.spatial import distance
import random
# The below function is to read the file and returns all the colours and size of the colurs
def readFile(name,s):
colours1 = []
with open(name) as f:
data=f.read().splitlines() # Reading the lines
for i in data:
if not re.search('[a-zA-Z]',i): # excluding the lines with text in it
d = i.split(' ')
d = [float(x) for x in d] # Converting the string values into float
colours1.append(d) # getting every colour and appending it in a list
f = s+1 #Getting input from the user
colours = colours1[1:f] # creating a list according to user input
size = len(colours); # calculating the sie of the file
return colours,size # returning the list of colours and its size
# The below function generates a list of random numbers according to the size
def rand(size):
return random.sample(range(0, size), size)
# The below function takes input the file with colours and permutation of index for which the distance has to be calculated
def evaluate(col,permutation):
Distance= []
for i in range(0,len(col)-1):
dst = distance.euclidean(col[permutation[i]],col[permutation[i+1]]) # calculate the distance according to the permutation
Distance.append(dst) # appending the eucledian distance into a list
allsum = sum(Distance) # calculating the sum of all the eucledian distances in the solution
return allsum # returning the sum of the distances'''
Perturbation functions
'''
# The below function returns a mutated permutation list
def two_opt_move(val):
mutated_lst = val[:] # make a copy
s = len(mutated_lst)
t1,t2 = 0, 0 # initialising two random points
while (t1 == t2): # ensure the two points are not the same
t1, t2 = random.randrange(0, s), random.randrange(0, s)
if t2 < t1: # ensure we always have t1<t2
t1, t2 = t2, t1
t2 = t2+1
reverse = list(reversed(mutated_lst[t1:t2])) # reversing the list
mutated_lst[t1:t2] = reverse
return mutated_lst # returning the reversed list
# The below function is used for swapping two index in a single term
def pertubation(lst):
pos = random.sample(range(0, len(lst)), 4) # generate 4 unique integers
lst[pos[0]],lst[pos[1]]=lst[pos[1]],lst[pos[0]] # swapping the first two
lst[pos[2]],lst[pos[3]]=lst[pos[3]],lst[pos[2]] # swapping the next two
return lst # returning the swapped list
# The below function performs the two_opt_move twice
def pertub_two_opt_move(val):
mutated_lst = val[:] # make a copy
d = int(len(mutated_lst)/2) # dividing the list into two parts so that the indexes don't coincide
lst1 = mutated_lst[:d] # storing the first half
lst2 = mutated_lst[d:] # storng the seconf half of the list
rand = random.sample(range(0,len(lst1)),2) # generating two unique random integers
t = rand[0]
t1 = rand[1]
rand1 = random.sample(range(0,len(lst2)),2) #generating two unique random integers
t2 = rand1[0]
t3 = rand1[1]
if t1 < t: # ensure we always have t<t1
t, t1 = t1, t
if t3 < t2: # ensure we always have t<t1
t2,t3 = t3,t2
t1 = t1+1
t3 = t3+1
reverse = list(reversed(lst1[t:t1])) # reversing the list from first part
reverse1 = list(reversed(lst2[t2:t3])) # reversing the list from the second part
lst1[t:t1] = reverse
lst2[t2:t3] = reverse1
mutated_lst = lst1+lst2 # concatinating both the reversed lists
return mutated_lst # returning the mutated solution
# The below function is for performing one-point crossover in the genetic algorithm
def crossover(dad, mom): # getting the input as dad and mom
xp =random.randint(0,len(dad)-1) # generating on random number for crossover
child1 = dad[:xp] # storing the first part of mom in first child
child2 = mom[:xp] # storing the first part of dad in second child
d = mom[xp:] # storing the next part of dad in first child
d1 = dad[xp:] # storing the next part of mom in second child
# All the below loops are to prevent the duplication of value by skipping if the child1 already has the value
# if child1 doesn't have the value it inherit it from mom and dad
for i in d:
if i not in child1:
child1.append(i)
for i in dad:
if i not in child1:
child1.append(i)
for i in mom:
if i not in child1:
child1.append(i)
# All the below loops are to prevent the duplication of value by skipping if the child2 already has the value
# if child2 doesn't have the value it inherit it from mom and dad
for j in d1:
if j not in child2:
child2.append(j)
for j in dad:
if j not in child2:
child2.append(j)
for j in mom:
if j not in child2:
child2.append(j)
return child1,child2 # returning both children '''
Random search
'''
def randomSearch(col_list,size):
dist_per_run = []
sol_per_run = [] # Empty lists
all_iterations = []
min_dist = len(col_list)
for i in range(20): # this loop for the no of runs
iterations = []
count = 0
while count < 1000: # this loop is for number of iterations
rand_col = rand(size) # generates a random list of indexes
total = evaluate(col_list,rand_col) # calculates the sum of eucledian distances
if total < min_dist : # we comapre and store the minimum distance with its solution untill the loop exits
min_dist = total
min_sol = rand_col
count +=1 # counter for the loop
iterations.append(min_dist) # appending every itertation into a list
dist_per_run.append(min_dist) # store the minimum distance for every iteration
sol_per_run.append(min_sol) # store the solution corresponding to minimum distance
all_iterations.append(iterations) # storing all the iterations for every run in a list
avg = np.mean(dist_per_run) # calculating the average of the 20 runs
std_dv = np.std(dist_per_run) # calculating the standard deviation of the 20 runs
minn = size
for h in range(0, len(dist_per_run)): # finding the minimum solution from 20 runs
if dist_per_run[h] < minn:
minn = dist_per_run[h] # returning the best distance
vbest_rand = sol_per_run[h] # returning the best solution
best_iteration = all_iterations[h] # returning the corresponding iteration
else:
continue
plot_colours(col_list,vbest_rand) # plotting the colors according to the minimum solution find
return minn,avg,std_dv,best_iteration,dist_per_run'''
Hill climber
'''
def hillClimber(col_list,size):
dist_per_run1 = []
best_soluion = [] # Empty lists
all_iterations1 = []
for _ in range(20): # this loop is for number of runs
iterations1 = []
rand_col = rand(size) # generating a random solution
total = evaluate(col_list,rand_col) # calculating the sum of eucledian distances
count = 0
while count < 1000: # this loop is for the number of iterations
new_rand_col=two_opt_move(rand_col) # mutating the random solution using the two opt move
new_best_dist = evaluate(col_list,new_rand_col) # calculating its sum of distances for mutated solution
if new_best_dist < total: # comapring the evaluation of mutated solution with evaluation of random solution
min_dist = new_best_dist # and saving the evaluation and solution in variables
min_sol = new_rand_col
else:
min_dist = total
min_sol = rand_col
count +=1
total = min_dist # assigning the minimum evaluation as the next starting point
rand_col=min_sol # assigning the corresponding solution as the next starting point
print(count)
iterations1.append(min_dist) # storing the evaluation at every iteration
all_iterations1.append(iterations1) # storing all the iterations according to the runs
dist_per_run1.append(min_dist) # storing the best value that we get at the end of every run
best_soluion.append(min_sol) # storing the corresponding solution
avg1 = np.mean(dist_per_run1) # calculating the average of the best 20 values
std_dv1 = np.std(dist_per_run1) # calculating the standard deviation of the 20 values
minn = len(col_list) # finding the minimum value and solution from the 20 runs
for h in range(0, len(dist_per_run1)):
if dist_per_run1[h] < minn:
minn = dist_per_run1[h] # storing the minimum value
vbest_rand = best_soluion[h] # storing the corresponding solution
best_iteration1 = all_iterations1[h] # storing the corresponding iteration
else:
continue
plot_colours(col_list,vbest_rand) #plotting the colours corresponding to the best found solution
return minn,avg1,std_dv1,best_iteration1,dist_per_run1'''
Iterated local search
'''
def iteratedLocalSearch(col_list,size):
dist_per_run2 = []
best_soluion = [] # Empty lists
all_iterations2 = []
for _ in range(20): # this loop is for number of runs
# we perform hill climbing at first to find the minimum solution
iterations2 = []
rand_col = rand(size) # generating a random solution
total = evaluate(col_list,rand_col) # calculating the sum of eucledian distances
count = 0
while count < 1000: # this loop is for the number of iterations
new_rand_col=two_opt_move(rand_col) # mutating the random solution using the two opt move
new_best_dist = evaluate(col_list,new_rand_col) # calculating the sum of eucledian distances for mutated solution
if new_best_dist < total: # comapring the evaluation of mutated solution with evaluation of random solution
min_dist = new_best_dist # and saving the evaluation and solution in variables
min_sol = new_rand_col
else:
min_dist = total # return the random evaluation and solution if it is less
min_sol = rand_col
count +=1
total = min_dist # assigning the minimum evaluation as the next starting point
rand_col = min_sol # assigning the corresponding solution as the next starting point
#print(min_dist)
# after we get the min sol then we pertubate it using the pertubated two opt move inorder to escape the local optima
per_min_sol = pertub_two_opt_move(min_sol)
# After pertubating we perform hill climbing again inorder to escape local optima
count1 = 0
while count1 < 1000: # this is the second loop which will look in the neighbourhood of the pertubated solution inorder to escape local optima
per_min_sol1=two_opt_move(per_min_sol) # mutating the pertubated solution with two opt move
per_best_dist1 = evaluate(col_list,per_min_sol1) # calculate the sum of the eucledian distances
if per_best_dist1 < min_dist: # compare the evaluation of mutated pertubated solution with the local optima
min_dist1 = per_best_dist1 # if the evaluation and solution are better then assign it to the variables
min_sol1 = per_min_sol1
else:
min_dist1 = min_dist # if not then assign the local optima as the best solution
min_sol1 = min_sol
count1 += 1
min_dist = min_dist1 # assign the minimum evaluation as the next starting point
min_sol = min_sol1 # assigning the corresponding solution as the next starting point
iterations2.append(min_dist1) # storing every evaluation at every iteration
all_iterations2.append(iterations2) # storng all the iterations according to the runs
dist_per_run2.append(min_dist1) # storing the best value at the end of every run
best_soluion.append(min_sol1) # storing the corresponding solution to that run
avg2 = np.mean(dist_per_run2) # calculating the average of all the 20 best values
std_dv2 = np.std(dist_per_run2) # calculating the standard deviation of the 20 best values
minn1 = len(col_list) # Finding the minimum values from the 20 best values
for h in range(0, len(dist_per_run2)):
if dist_per_run2[h] < minn1:
minn1 = dist_per_run2[h] # storing the minimum value
vbest_rand = best_soluion[h] # storing the corresponding solution
best_iteration2 = all_iterations2[h] # storing the corresoinding iteration
else:
continue
plot_colours(col_list,vbest_rand) # plotting the colours with the best permutation
return minn1,avg2,std_dv2,best_iteration2,dist_per_run2'''
Evolutionary algorithm
'''
def evolutionaryAlgorithm(col_list,size):
best_dist_list=[]
best_sol_list = [] # Empty lists
all_iterations3 = []
for i in range(20): # this loop is to perform number of runs
iterations3 = []
population = []
for i in range(100): # Here we generate a new population at random for every run an the number represents the length of population
new_col_list = rand(size) # generating a random solution
new_dist = evaluate(col_list,new_col_list) # evaluating the random solution
population.append((new_col_list,new_dist)) # appending the solution and evaluation in the form of tuple
for o in range(1000): # this loop is for the number of iterations
rand_sol = random.sample(population,4) # we take four random solutions from the population
dad1 = rand_sol[0]
dad2 = rand_sol[1] # Assign every solution to different variables which represent as parents
mom1 = rand_sol[2]
mom2 = rand_sol[3]
# here the tournament selection happens
if dad1[1]<dad2[1]:
best_dad = dad1
else: # here we choose the best dad on the basis of evaluation
best_dad = dad2
if mom1[1]<mom2[1]:
best_mom = mom1
else: # here we chose the best mom on the basis of evaluation
best_mom = mom2
child1,child2 = crossover(best_dad[0],best_mom[0]) # This step we create two new children from mom and dad using the one point crossover function
child1 = two_opt_move(child1) # mutate the first child using the two opt move
child1_dist = evaluate(col_list,child1) # calculate the sum of eucledian distances
child2 = two_opt_move(child2) # mutate the second child
child2_dist = evaluate(col_list,child2) # calculate the sum of eucledian distances
value = 0
for j in range(len(population)): # here we find the worst value in the population and replace it with child 1
if population[j][1] > value:
value = population[j][1]
worst1 = population[j][0] # worst value
index = j # saving the index of the worst value
population[index]=(child1,child1_dist) # replacing the child1 on the particular index in the form of tuple(solution,evaluation)
min_val = len(col_list)
for v in population: # here we find the first minimum value in the population inorder to store it for every iteration
if v[1] < min_val:
min_val = v[1]
else:
continue
value1 = 0
for k in range(len(population)): # here we find the worst value in the population and replace it with child 2
if population[k][1] > value1:
value1 = population[k][1]
worst2 = population[k][0] # worst value
index = k # saving the index of the worst value
population[index]=(child2,child2_dist) # replacing the child2 on the particular index in the form of tuple(solution,evaluation)
min_val1 = len(col_list)
for u in population: # here we find the second minimum value in the population inorder to store it for every iteration
if u[1] < min_val1:
min_val1 = u[1]
else:
continue
iterations3.append(min_val) # storing the first minimum value for every iteration
iterations3.append(min_val1) # storing the second minimum value for every iteration
value2 = len(col_list)
for i in population: # here we find the minimum value inside the population for every run
if i[1] < value2: # as we are doing 20 runs, so we get 20 values in the end
value2 = i[1]
sol = i[0]
else:
continue
all_iterations3.append(iterations3) # storing all the iterations for every
best_dist_list.append(value2) # storing the minimum value at every run
best_sol_list.append(sol) # storing the corresponding solution
minn = len(col_list)
for h in range(0, len(best_dist_list)): # here we find the best minimum value from the 20 values
if best_dist_list[h] < minn:
minn = best_dist_list[h] # store the minimum value in a variable
best_sol = best_sol_list[h] # store the corresponding solution
best_iteration3 = all_iterations3[h] # store the corresponing iteration
else:
continue
avg3 = np.mean(best_dist_list) # Calculate the average of the best 20 values
std_dv3 = np.std(best_dist_list) # calculate the standard deviation of the best 20 values
plot_colours(col_list,best_sol) # plot the colours with the best found solution
return minn,avg3,std_dv3,best_iteration3,best_dist_list'''
Box plots, line plots and plotting the permutations
'''
# The below function is used for plotting the colours according to the permutation
def plot_colours(colours, permutation):
assert len(colours) == len(permutation)
ratio = 90 # ratio of line height/width, e.g. colour lines will have height 10 and width 1
img = np.zeros((ratio, len(colours), 3))
for i in range(0, len(colours)):
img[:, i, :] = colours[permutation[i]]
fig, axes = plt.subplots(1, figsize=(8,4)) # figsize=(width,height) handles window dimensions
axes.imshow(img, interpolation='nearest')
axes.axis('off') # removing all the axis
plt.show()
# The below function is used for plotting the line plot
def linePlot(itr1,itr2,itr3,itr4): # take the input from all the algorithms
lst = [itr1,itr2,itr3,itr4]
algo = ['Random Search','Hill Climber','Iterated Local Search','Evolutionary Algorithm']
plot_list = [list(zip(*[(z+1,y) for z,y in enumerate(x)])) for x in lst] # using list comprehensions for nested list
for p in plot_list: # iterating through the list and plotting it
plt.plot(*p)
plt.ylabel("Best fitness")
plt.xlabel("Number of Iterations")
plt.legend(algo, fontsize=7, loc = 'upper right')
return plt.show()
# The below function is used for plotting the box plots
def boxplot(lst1,lst2,lst3,lst4):
mylist = [lst1,lst2,lst3,lst4] # collecting all the inputs in a list
plt.boxplot(mylist) # plotting the list
plt.xticks([1,2,3,4],['Random Search','Hill Climber','Iterated Local Search','Evolutionary Algorithm']) # changing the x-axis to the algorithm's name
plt.xticks(rotation=30) # rotating the values on x-axis
return plt.show()'''
Main function
'''
def main():
Inp = int(input('Enter the no of colours : ')) # here we take the input from the user
colours,size = readFile('colours.txt',Inp) # here we provide the data file and get the output in the form of list ans size of the file
RS = randomSearch(colours,size) # here we call our first algorithm and all the returned values are stored in the form of a tuple
HC = hillClimber(colours,size) # here we call our second algorithm and all the returned values are stored in the form of a tuple
ILS = iteratedLocalSearch(colours,size) # here we call our third algorithm and all the returned values are stored in the form of a tuple
EA = evolutionaryAlgorithm(colours,size) # here we call our fourth algorithm and all the returned values are stored in the form of a tuple
print(' No of Colours:',Inp,' Average Standard Deviation Best Fitness ')
print('-----------------------------------------------------------------------------------')
print(" Random Search Algorithm : ",RS[1],' ',RS[2],' ',RS[0])
print(" Hill Climber Algorithm : ",HC[1],' ',HC[2],' ',HC[0]) # here we print the results returned from the algorithms
print(" Iterated Local Search : ",ILS[1],' ',ILS[2],' ',ILS[0])
print(" Evolutionary Algorithm : ",EA[1],' ',EA[2],' ',EA[0])
boxplot(RS[4],HC[4],ILS[4],EA[4]) # here we call he boxplot function for plotting the boxes
linePlot(RS[3],HC[3],ILS[3],EA[3]) # here we call the lineplot function for plotting the line plots
return