An R package for lightweight cost-effectiveness analysis using decision trees.
Requests and comments welcome; please use Issues.
To install the development version from github:
library(devtools)
install_github("Health-Economics-in-R/CEdecisiontree")Then, to load the package, use:
library(CEdecisiontree)Decisions trees can be modelled as special cases of more general models using available packages in R e.g. heemod, mstate or msm. Further, full probabilty models could be fit using a Bayesian model with e.g. jags or WinBUGS. However, simple decision tree models are often built in Excel, using statistics from literature or expert knowledge. This package is a analogue to these, such that models can be specified in a very similar and simple way.
A decision tree is defined by parent-child pairs, i.e. from-to
connections, and the probability and associated value (e.g. cost) of
traversing each of the connections. Denote the probability of
transitioning from node
to
as
and
the cost attributable to node
as
. Where no
connection exists between two nodes we shall say that the parent’s set
of children is the empty set
. Denote the set of children by
. Clearly, there are no
or
in this case but
for computational purposes we will assume that
and
.
The expected value at each node is
calculated by ‘folding back’ using the recursive formula
with boundary values at the terminal nodes
Quietly load libraries.
suppressPackageStartupMessages(library(CEdecisiontree))
suppressPackageStartupMessages(library(readr))
#> Warning: package 'readr' was built under R version 3.4.4
suppressPackageStartupMessages(library(dplyr))
#> Warning: package 'dplyr' was built under R version 3.4.4
suppressPackageStartupMessages(library(reshape2))
#> Warning: package 'reshape2' was built under R version 3.4.4
suppressPackageStartupMessages(library(tidyr))
#> Warning: package 'tidyr' was built under R version 3.4.4
suppressPackageStartupMessages(library(assertthat))
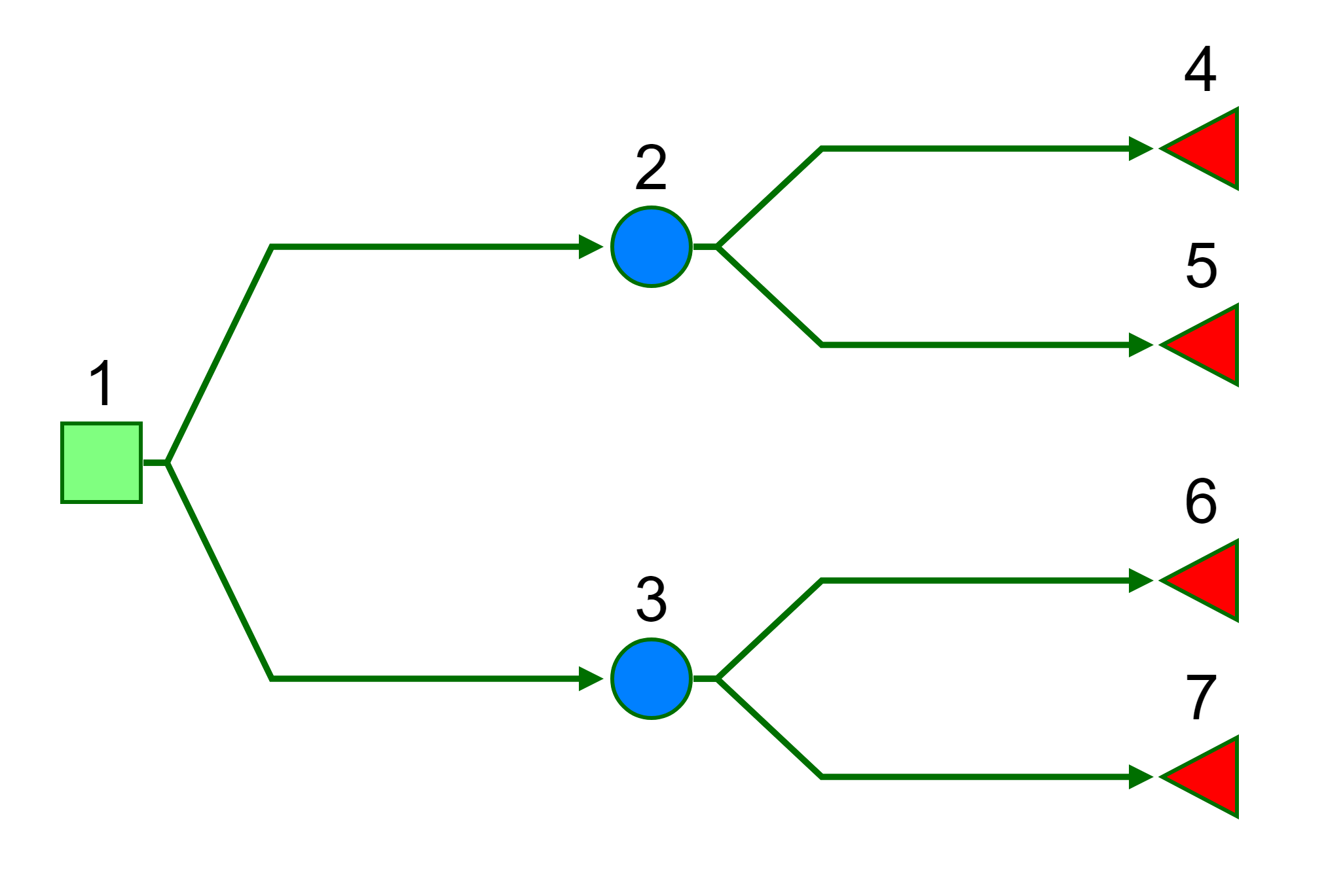
#> Warning: package 'assertthat' was built under R version 3.4.4We will consider a simple 7 node binary tree.
Load example data from the package.
data("cost")
data("probs")The cost and probability matrices we will use in this example are sparse
arrays indicating the edge values (rows=from node, columns=to node).
There are therefore the same dimensions and have the same entry pattern.
Empty cells have NA.
cost
#> # A tibble: 3 x 7
#> `1` `2` `3` `4` `5` `6` `7`
#> <dbl> <int> <int> <int> <int> <int> <int>
#> 1 NA 10 1 NA NA NA NA
#> 2 NA NA NA 10 1 NA NA
#> 3 NA NA NA NA NA 10 1probs
#> # A tibble: 3 x 7
#> `1` `2` `3` `4` `5` `6` `7`
#> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl>
#> 1 NA 0.2 0.8 NA NA NA NA
#> 2 NA NA NA 0.2 0.8 NA NA
#> 3 NA NA NA NA NA 0.2 0.8probs is a probability transition matrix. This is like pmatrix.msm
in the msm package, or define_transition in the heemod package.
The transMat() function in the mstate package creates a closely
related multi-state model transition matrix. Copying this package, we
can create a decision tree transition matrix to use with this.
CEdecisiontree:::trans_binarytree(depth = 3)
#> to
#> from 1 2 3 4 5 6 7
#> 1 NA 1 2 NA NA NA NA
#> 2 NA NA NA 3 4 NA NA
#> 3 NA NA NA NA NA 5 6The expected value at each node is calculate as follows.
dectree_expected_values(vals = cost,
p = probs)
#> 1 2 3 4 5 6 7
#> 5.6 12.8 3.8 10.0 1.0 10.0 1.0There is also an Rcpp version of this function.
Cdectree_expected_values(vals = as.matrix(cost),
p = as.matrix(probs))For additional information, inclusing for the purposes of model checking we can calculate other tree statistics. We can obtain the contributing cost as weighted by the chance of occurrence. This can be thought of as a trade-off between the raw, original cost and branch position.
wcost <- branch_joint_probs(probs) * cost
wcost
#> 1 2 3 4 5 6 7
#> 1 NA 2 0.8 NA NA NA NA
#> 2 NA NA NA 0.4 0.16 NA NA
#> 3 NA NA NA NA NA 1.6 0.64We can check that this sums to the same total expected cost.
sum(wcost, na.rm = TRUE)
#> [1] 5.6We can also calculate the joint probabilities of traversing to each
terminal state using branch_joint_probs. Here we assume node labelling
order from root such that terminal nodes are last.
n_from_nodes <- nrow(probs)
n_to_nodes <- ncol(probs)
terminal_states <- (n_from_nodes + 1):n_to_nodes
p_terminal_state <-
branch_joint_probs(probs)[ ,terminal_states] %>%
colSums(na.rm = TRUE)
p_terminal_state
#> 4 5 6 7
#> 0.04 0.16 0.16 0.64
sum(p_terminal_state)
#> [1] 1See package vignette for more details and examples.
GPL-3 ©