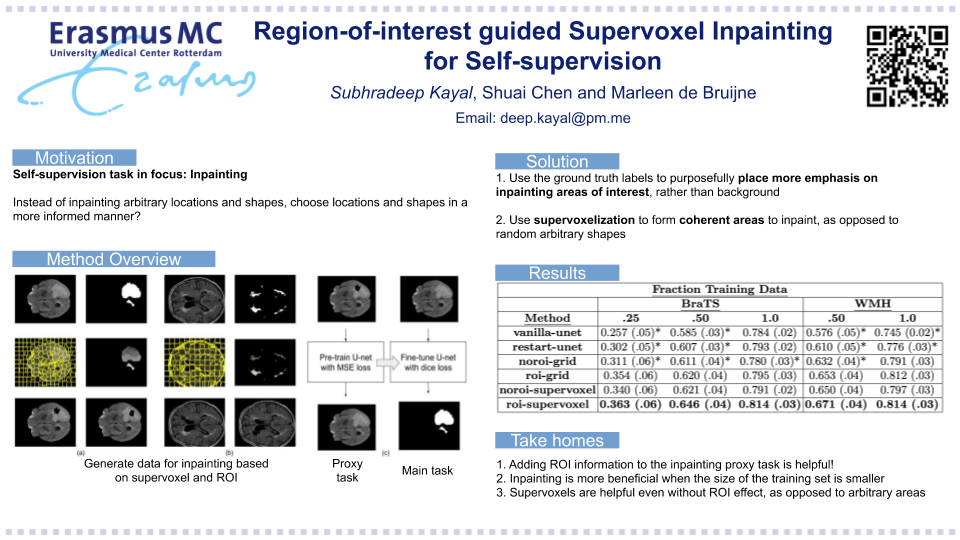
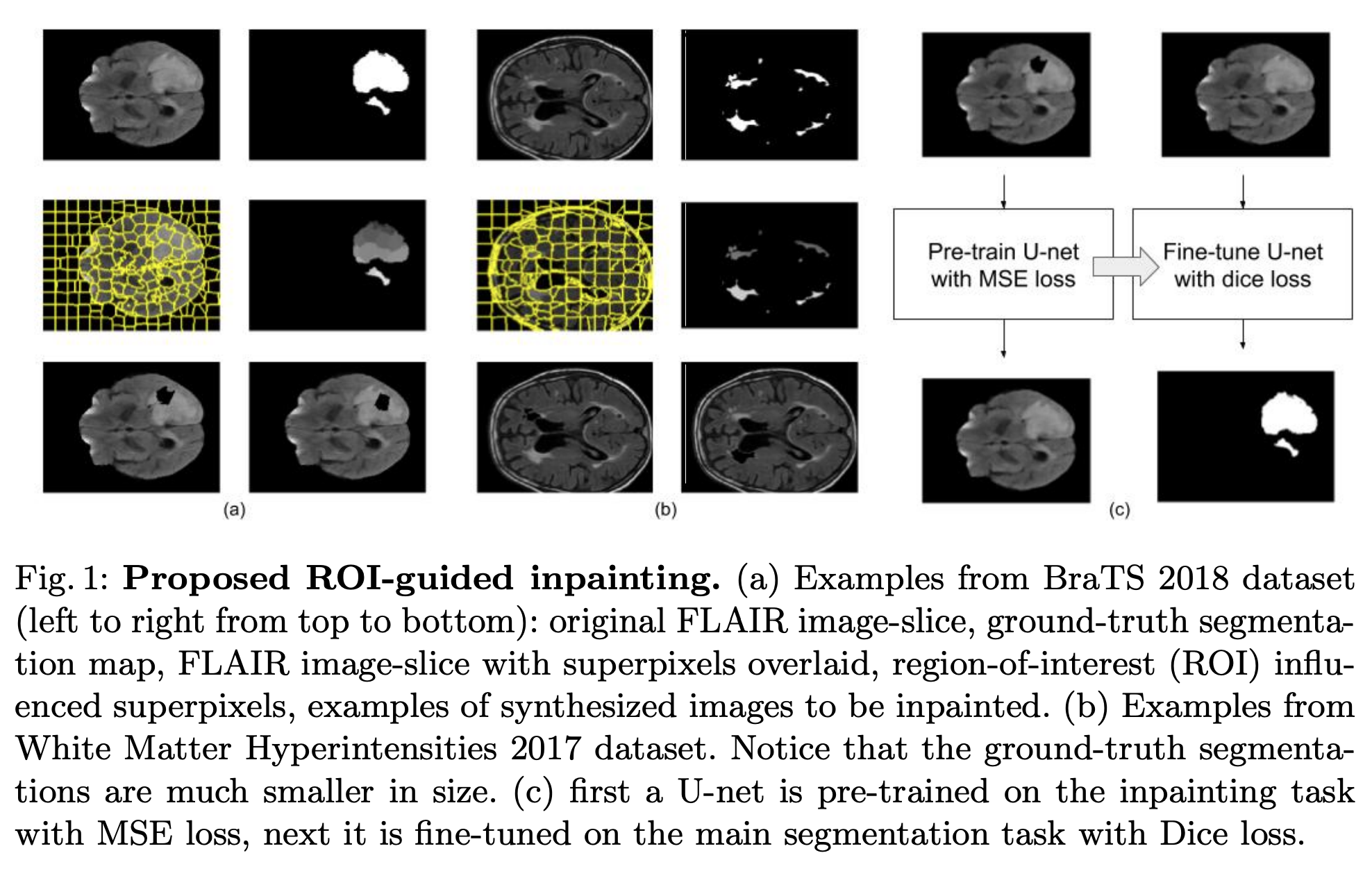
Code the paper titled Region-of-interest guided Supervoxel Inpainting for Self-supervision published at MICCAI 2020 (https://arxiv.org/pdf/2006.15186.pdf) by Subhradeep Kayal, et al.
Citation
If you find this paper useful for your research, please consider citing the paper:
@inproceedings{kayal2020region,
title={Region-of-interest guided Supervoxel Inpainting for Self-supervision},
author={Kayal, Subhradeep and Chen, Shuai and de Bruijne, Marleen},
booktitle={International Conference on Medical Image Computing and Computer-Assisted Intervention},
pages={500--509},
year={2020},
organization={Springer}
}
Requirements
- nibabel
- keras
- tensorflow
- numpy
- sklearn
- skimage
- tqdm
- cv2
Steps (to repeat experiments for BraTS 2018)
- Get BraTS data from https://www.med.upenn.edu/sbia/brats2018/data.html
- Create a folder within
inpaintingcalleddataand put the BraTS data in there, with the folder nameMICCAI_BraTS_2018_Data_Training. Make sure there is a folderHGGin there with the relevant files. We only use the images concerning high grade glioma (HGG) in these experiments. - Run
train.sh - Run
evaluate.shwhen the above is finished