Copyright (C) 2020-2021
Benjamin Paaßen
The University of Sydney
Daniele Grattarola, Daniele Zambon
Università della Svizzera italiana
This program is free software: you can redistribute it and/or modify it under the terms of the GNU General Public License as published by the Free Software Foundation, either version 3 of the License, or (at your option) any later version.
This program is distributed in the hope that it will be useful, but WITHOUT ANY WARRANTY; without even the implied warranty of MERCHANTABILITY or FITNESS FOR A PARTICULAR PURPOSE. See the GNU General Public License for more details.
You should have received a copy of the GNU General Public License along with this program. If not, see http://www.gnu.org/licenses/.
This repository contains the reference implementation of Graph Edit Networks as described in the paper
- Paaßen, B., Grattarola, D., Zambon, D., Alippi, C., and Hammer, B. (2021). Graph Edit Networks. Proceedings of the Ninth International Conference on Learning Representations (ICLR 2021). Link
@inproceedings{Paassen2021ICLR,
title={Graph Edit Networks},
author={Benjamin Paaßen and Daniele Grattarola and Daniele Zambon and Cesare Alippi and Barbara Hammer},
booktitle={Proceedings of the Ninth International Conference on Learning Representations (ICLR 2021)},
editor={Shakir Mohamed and Katja Hofmann and Alice Oh and Naila Murray and Ivan Titov},
venue={virtual},
year={2021},
url={https://openreview.net/forum?id=dlEJsyHGeaL}
}
In particular, this repository contains all experimental scripts, model implementations, datasets, and auxiliary files necessary to run the experiments.
In the remainder of this README, we provide the required packages to run the modules of this repository, a guide how to train your own graph edit networks on novel data, instructions how to reproduce the experiments reported in the paper, and a detailed list of all enclosed files with explanations.
All software enclosed in this repository is written in Python 3. To train graph edit networks, you additionally require PyTorch (Version >= 1.4.0; torchvision or cuda are not required). To train tree edit networks, you additionally require edist (Version >= 1.1.0), which in turn requires numpy (Version >= 1.17).
To run the kernel time series prediction (Paaßen et al., 2018) baseline, you require edist (Version >= 1.1.0) and scikit-learn (Version >= 0.21.3).
These packages are also sufficient to run the experiments. All dependencies are available on pip.
To train your own graph edit networks you require two ingredients. First,
training data in the form of time series of graphs. Each time series should
be a list of graphs, and each graph should be a tuple (A, X) of a numeric
node attribute matrix
As a second ingredient, you require a teaching protocol, i.e. a function that determines for each graph pair (A, X) and (B, Y) the graph edit scores that the graph edit network should return to get from (A, X) to (B, Y). While the paper provides a general-purpose construction based on graph edit distance approximators, we recommend to define teaching protocols specific for each dataset, which is expected to be more efficient. As an example, consider the following two graphs
import torch
X = torch.tensor([[1., 0.], [0., 1.]])
A = torch.tensor([[0., 1.], [1., 0.]])
Y = torch.tensor([[1., 0.]])
B = torch.tensor([[0.]])In other words, the first graph has two connected nodes, labelled with the vectors (1, 0) and (0, 1) respectively, and the second graph has one node, labelled with the vector (1, 0). Accordingly, the quickest way to get from the first to the second graph is to delete the second node. In a graph edit network, we can realize this operation with the following edit scores.
delta = torch.tensor([0., -1.])
Epsilon = torch.zeros((2, 2))which tells the network to delete the second node, but make no edge changes.
Now, let training_data be a list of graph time series and let
training_protocol be a function which implements our training protocol.
Then, we can train a graph edit network with the following template.
import random
import torch
import pytorch_graph_edit_networks as gen
model = gen.GEN(num_layers = 2, dim_in = 2, dim_hid = 64)
optimizer = torch.optim.Adam(model.parameters(), lr=1E-3, weight_decay=1E-5)
loss_fun = gen.GEN_loss()
for epoch in range(10000):
optimizer.zero_grad()
# sample a random time series from the training data
i = random.randrange(len(training_data))
time_series = training_data[i]
# iterate over the time series
for t in range(len(time_series)-1):
# retrieve the current graph
A, X = time_series[t]
# call the graph edit network to predict edit scores
delta, Epsilon = model(A, X)
# retrieve the actual next graph
B, Y = time_series[t+1]
# call the teaching protocol to infer the scores the network
# should have returned
delta_hat, Epsilon_hat = teaching_protocol(A, X, B, Y)
# compute the loss
loss = loss_fun(delta, Epsilon, delta_hat, Epsilon_hat, A)
# compute the gradient. Pytorch will automatically accumulate
# gradients across the time series
loss.backward()
# after the time series has been processed, perform an optimizer step
optimizer.step()Note that the hyper-parameters given here are used in all our experiments on
synthetic data. For other data, though, they may need to be adjusted.
Further, it may be prudent to use different conditions to end training (like
getting below a certain loss threshold) and to track the training loss over
time. Finally, note that we do not include node attributes as output here,
because all our non-tree experiments concern unlabeled graphs. For labeled
graphs, pytorch_graph_edit_networks needs to be adjusted to also output a
predicted attribute for replacements and insertions.
After training, we can predict changes on a graph (A, X) via the following
template.
import pytorch_graph_edit_networks as gen
# predict the edit scores
delta, Epsilon = model(A, X)
# transform into graph edits
edits = gen.to_edits(A, delta, Epsilon)
# apply the edits to the graph
for edit in edits:
B, Y = edit.apply(A, X)Trees are a an interesting special case because their edit distance is
efficiently computeable (Zhang and Shasha, 1989). Accodingly,
we can derive a general-purpose teaching protocol which always predicts a
shortest edit script between two trees. Our implementation of this sheme
is the pytorch_tree_edit_networks module.
To train your own tree edit network, you again require training data in
form of time series of trees, where each tree is given in terms of a list
of node labels and an adjacency list. For example, the tree and(x, not y)
would have the node list ['and', 'x', 'not_y'] and the adjacency list
[[1, 2], [], []] because node 0 ('and') has nodes 1 and 2 ('x' and 'not y')
as children, whereas the other nodes have no children.
Now, let training_data be a list of tree time series and let alphabet
be a list of possible labels. Then we can train a tree edit network using
the following template.
import random
import torch
import pytorch_tree_edit_networks as ten
model = ten.TEN(num_layers = 2, alphabet = alphabet, dim_hid = 64)
optimizer = torch.optim.Adam(model.parameters(), lr=1E-3, weight_decay=1E-5)
for epoch in range(10000):
optimizer.zero_grad()
# sample a random time series from the training data
i = random.randrange(len(training_data))
time_series = training_data[i]
# compute the loss over the entire time series
loss = model.loss_over_time_series(time_series)
# compute the gradient
loss.backward()
# perform an optimizer step
optimizer.step()After training, we can predict the edits on a new tree via the following template.
_, nodes_next, adj_next = model.predict_macrostep(nodes, adj)Note that this scheme has caveats. In order to be general-purpose, we need to permit arbitrarily many insertions, which in turn may require several prediction steps on intermediate trees ('microsteps') before arriving at the final destination. To avoid this, it can be more effective to use a dedicated teaching protocol for each dataset, which is what we do in the Boolean and Peano dataset.
Reproducing our experiments is possible by running the four included ipython notebooks. All notebooks should run without any additional preparation. Installing the dependencies listed above should suffice. Note that slight deviations may occur due to different sampling. In the remainder of this section, we list the different experiments and their notebooks with the expected results in each case.
graph_dynamical_systems.ipynb contains the experiments on the three graph
dynamical systems (edit cycles, degree rules, and game of life). The results
should be as follows.
--- data set edit_cycles --- --- model VGAE --- node_ins_recall: 0.616033 +- 0.0137438 node_ins_precision: 1 +- 0 node_del_recall: 1 +- 0 node_del_precision: 0.690418 +- 0.0622833 edge_ins_recall: 1 +- 0 edge_ins_precision: 1 +- 0 edge_del_recall: 1 +- 0 edge_del_precision: 1 +- 0 --- model GEN_crossent --- node_ins_recall: 1 +- 0 node_ins_precision: 1 +- 0 node_del_recall: 1 +- 0 node_del_precision: 1 +- 0 edge_ins_recall: 1 +- 0 edge_ins_precision: 1 +- 0 edge_del_recall: 1 +- 0 edge_del_precision: 1 +- 0 --- model GEN --- node_ins_recall: 1 +- 0 node_ins_precision: 1 +- 0 node_del_recall: 1 +- 0 node_del_precision: 1 +- 0 edge_ins_recall: 1 +- 0 edge_ins_precision: 1 +- 0 edge_del_recall: 1 +- 0 edge_del_precision: 1 +- 0 --- data set degree_rules --- --- model VGAE --- node_ins_recall: 0.146334 +- 0.033497 node_ins_precision: 1 +- 0 node_del_recall: 1 +- 0 node_del_precision: 0.95537 +- 0.0244676 edge_ins_recall: 0.881462 +- 0.0254464 edge_ins_precision: 0.973631 +- 0.0516198 edge_del_recall: 1 +- 0 edge_del_precision: 0.965385 +- 0.0692308 --- model GEN_crossent --- node_ins_recall: 1 +- 0 node_ins_precision: 1 +- 0 node_del_recall: 1 +- 0 node_del_precision: 1 +- 0 edge_ins_recall: 0.967017 +- 0.0453511 edge_ins_precision: 0.9869 +- 0.0167524 edge_del_recall: 1 +- 0 edge_del_precision: 1 +- 0 --- model GEN --- node_ins_recall: 1 +- 0 node_ins_precision: 1 +- 0 node_del_recall: 1 +- 0 node_del_precision: 1 +- 0 edge_ins_recall: 0.970246 +- 0.0569909 edge_ins_precision: 0.985629 +- 0.0287424 edge_del_recall: 1 +- 0 edge_del_precision: 1 +- 0 --- data set game_of_life --- --- model VGAE --- node_ins_recall: 0.268 +- 0.0808455 node_ins_precision: 1 +- 0 node_del_recall: 1 +- 0 node_del_precision: 0.0341 +- 0.00352307 edge_ins_recall: 1 +- 0 edge_ins_precision: 1 +- 0 edge_del_recall: 1 +- 0 edge_del_precision: 1 +- 0 --- model GEN_crossent --- node_ins_recall: 1 +- 0 node_ins_precision: 1 +- 0 node_del_recall: 1 +- 0 node_del_precision: 0.979732 +- 0.0405352 edge_ins_recall: 1 +- 0 edge_ins_precision: 1 +- 0 edge_del_recall: 1 +- 0 edge_del_precision: 1 +- 0 --- model GEN --- node_ins_recall: 1 +- 0 node_ins_precision: 0.997893 +- 0.0042144 node_del_recall: 1 +- 0 node_del_precision: 0.9991 +- 0.00111355 edge_ins_recall: 1 +- 0 edge_ins_precision: 1 +- 0 edge_del_recall: 1 +- 0 edge_del_precision: 1 +- 0
pytorch_ten.ipynb runs a tree edit network on the two tree dynamical systems
(boolean and peano addition). Here, we expect the following results.
--- data set boolean --- Accuracy: 1 +- 0 Epochs: 8320.2 +- 716.397 --- data set peano --- Accuracy: 1 +- 0 Epochs: 12768.4 +- 933.092
kernel_tree_time_series_prediction.ipynb runs the kernel regression baseline
on the two tree dynamical systems. Here, we expect the following results.
--- task boolean --- repeat 1 of 5 took 0.137753 seconds to train took 0.470501 seconds to predict RMSE: 2.41091 versus baseline RMSE 1.45774 repeat 2 of 5 took 0.129635 seconds to train took 0.469569 seconds to predict RMSE: 2.43812 versus baseline RMSE 1.61589 repeat 3 of 5 took 0.135248 seconds to train took 0.419243 seconds to predict RMSE: 2.46306 versus baseline RMSE 1.52753 repeat 4 of 5 took 0.130834 seconds to train took 0.215008 seconds to predict RMSE: 2 versus baseline RMSE 1.7097 repeat 5 of 5 took 0.131168 seconds to train took 0.382734 seconds to predict RMSE: 2.65684 versus baseline RMSE 1.59041 --- task peano --- repeat 1 of 5 took 3.70749 seconds to train took 58.3135 seconds to predict RMSE: 5.38069 versus baseline RMSE 3 repeat 2 of 5 took 4.74174 seconds to train took 65.4441 seconds to predict RMSE: 3.96863 versus baseline RMSE 2.79322 repeat 3 of 5 took 3.83915 seconds to train took 61.2994 seconds to predict RMSE: 4.89415 versus baseline RMSE 3.44944 repeat 4 of 5 took 4.83539 seconds to train took 60.4223 seconds to predict RMSE: 3.52332 versus baseline RMSE 2.77302 repeat 5 of 5 took 5.58519 seconds to train took 88.7766 seconds to predict RMSE: 4.94673 versus baseline RMSE 3.19398
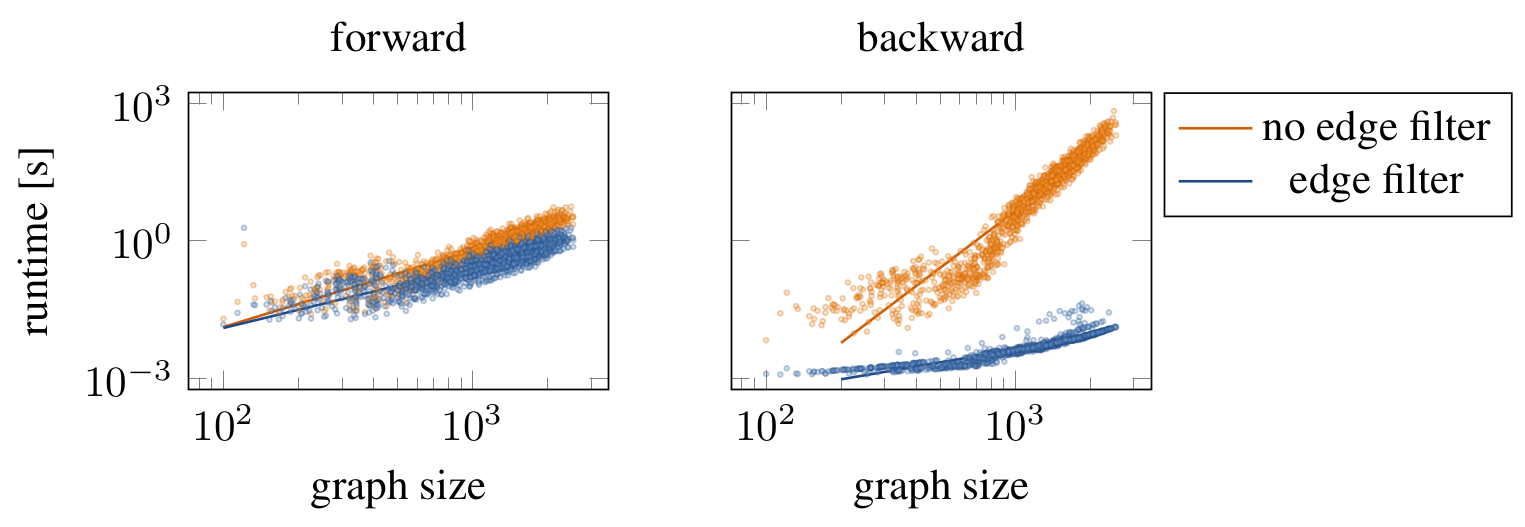
hep_th_runtimes.ipynb runs graph edit networks on realistic graphs from the
HEP-Th dataset and reports the runtime needed. Then, we compute a log-log fit
of runtime versus graph size. We expect the following results.
log-fit for forward computation of no_filter model: log(y) = 1.66529 * log(x) + -12.0214 log-fit for forward computation of flex_filter model: log(y) = 1.32117 * log(x) + -10.4584 log-fit for forward computation of const_filter model: log(y) = 1.33302 * log(x) + -10.4088 log-fit for backward computation of no_filter model: log(y) = 4.10825 * log(x) + -26.9001 log-fit for backward computation of flex_filter model: log(y) = 0.93353 * log(x) + -11.883 log-fit for backward computation of const_filter model: log(y) = 1.28752 * log(x) + -12.707 log-fit for inference computation of no_filter model: log(y) = 1.69191 * log(x) + -12.1958 log-fit for inference computation of flex_filter model: log(y) = 1.32336 * log(x) + -10.4626 log-fit for inference computation of const_filter model: log(y) = 1.31487 * log(x) + -10.2706
The visualization should look roughly like this.
In more detail, the following files are contained in this repository (in alphabetical order):
baseline_models.py: An implementation of variational graph autoencoders (VGAE; Kipf and Welling, 2016) for time series prediction.boolean_formulae.py: A Python script generating the Boolean dataset and its teaching protocol.boolean_formulae_test.py: A unit test forboolean_formulae.py.degree_rules.py: A Python script generating the degree rules dataset and its teaching protocol.degree_rules_test.py: A unit test fordegree_rules.py.game_of_life.py: A Python script generating the game of life dataset and its teaching protocol.game_of_life_test.py: A unit test forgame_of_life.py.graph_dynamical_systems.ipynb: An ipython notebook containing the graph edit cycles, degree rules, and game of life experiments.graph_edit_cycles.py: A Python script generating the graph edit cycles dataset and its teaching protocol.graph_edits.py: A Python implementation of graph edits.hep-th: A directory containing the HEP-Th dataset as used in this paper, including the preprocessing script used.hep_th.py: A Python script to load the HEP-Th dataset and its teaching protocol.hep_th_runtimes.csv: A table of runtime results obtained on the HEP-Th dataset.hep_th_runtimes.ipynb: An ipython notebook to generate the runtime results.hep_th_runtimes.png: An image file displaying the runtime results obtained on the HEP-Th dataset.hep_th_test.py: A unit test forhep_th.py.kernel_time_series_prediction.py: An implementation of kernel time series prediction for trees as proposed by Paaßen et al., 2018.kernel_time_series_prediction_test.py: A unit test forkernel_time_series_prediction.py.kernel_tree_time_series_prediction.ipynb: An ipython notebook to run kernel time series prediction on the Boolean and Peano datasets.peano_addition.py: A Python script generating the Peano dataset and its teaching protocol.peano_addition_test.py: A unit test forpeano_addition.py.pygraphviz_interface.py: An auxiliary file to draw graphs using graphviz.pytorch_graph_edit_networks.py: An implementation of graph edit networks and the according loss function as reported in the paper.pytorch_graph_edit_networks_test.py: A unit test forpytorch_graph_edit_networks.py.pytorch_ten.ipynb: An ipython notebook containing the Boolean and the Peano experiments.pytorch_tree_edit_networks.py: An implementation of tree edit networks including a general-purpose teaching protocol (with the caveats described above).pytorch_tree_edit_networks_test.py: A unit test forpytorch_tree_edit_networks.py.README.md: this file.
- Bougleux, Brun, Carletti, Foggia, Gaüzère, and Vento (2017). Graph edit distance as a quadratic assignment problem. Pattern Recognition Letters, 87, 38-46. doi:10.1016/j.patrec.2016.10.001. Link
- Kipf, and Welling (2016). Variational Graph Auto-Encoders. Proceedings of the NIPS 2016 Workshop on Bayesian Deep Learning. Link
- Paaßen, Göpfert, and Hammer (2018). Time Series Prediction for Graphs in Kernel and Dissimilarity Spaces. Neural Processing Letters, 48, 669-689. doi:10.1007/s11063-017-9684-5. Link
- Zhang, and Shasha (1989). Simple Fast Algorithms for the Editing Distance between Trees and Related Problems. SIAM Journal on Computing, 18(6), 1245-1262. doi:10.1137/0218082