Use Deep Learning models to segment cells in microscopy image.
The dataset is downloaded from: https://www.kaggle.com/competitions/sartorius-cell-instance-segmentation/data
Sample image in this dataset:
Models:
- Mask RCNN
- Cellpose
- Training Size Model (Cellpose's assistant model)
Data:
- Mosaic
- Add extra data for the
SH-SY5Ycell line from LIVECell dataset which is the predecessor of this dataset - Data Augmentation (Flip left/right, Flip up/down, Crop, Add noise, Rotation
Loss:
- L2 Regularization
Installing all packages in this repository.
pip install -r requirement.txt
./browser/ folder stores all demo files which use Mask RCNN or/and Cellpose to detect cells in microscopy image on web.
Using command streamlit run main.py to start server.
.
├── browser
│ ├── cellpose_utils.py
│ ├── main.py
│ ├── mrcnn_utils.py
│ └── utils.py
Mask RCNN and Cellpose packages are stored in ./models/ folder.
.
├── models
│ ├── Mask_RCNN
│ ├── cellpose
./technique/ folder stores all files demo of those techniques used in this project
.
├── technique
│ ├── augmentation
│ ├── helper_package
│ ├── livecell prepare.ipynb
│ ├── mini mask.ipynb
│ └── mosaic
All .ipynb files which used to train and test model are stored in ./train-infer-model/ folder.
.
├── train-infer-model
│ ├── cellpose
│ ├── data
│ ├── mask_rcnn
│ └── performance
Using streamlit framework to demo on website
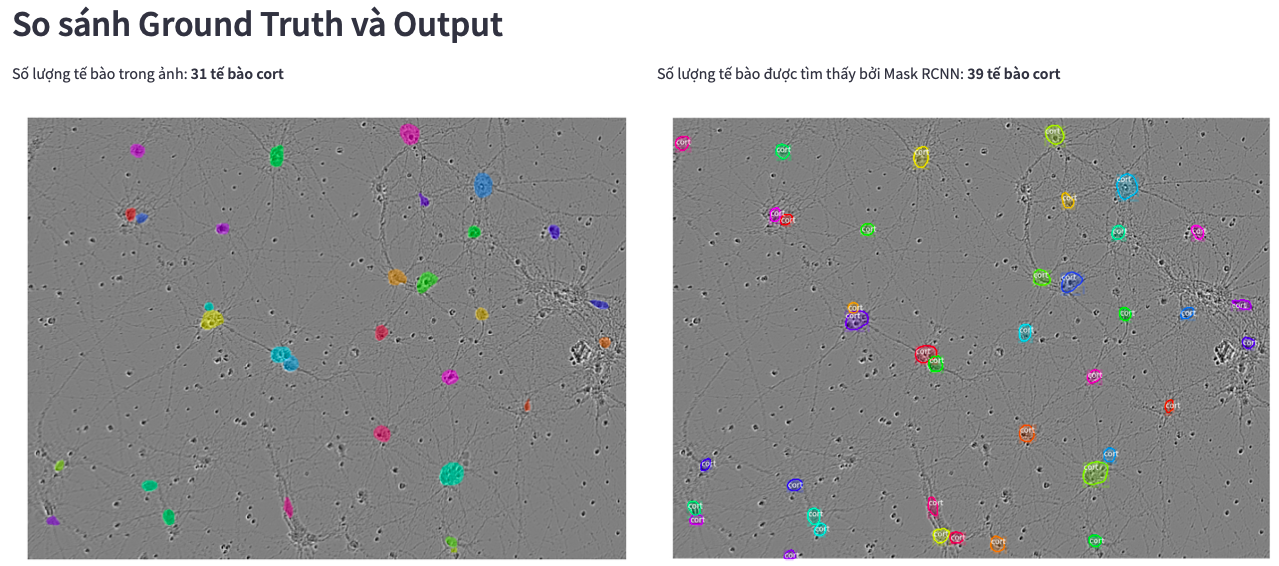
Mask RCNN also detects the class label for each instance
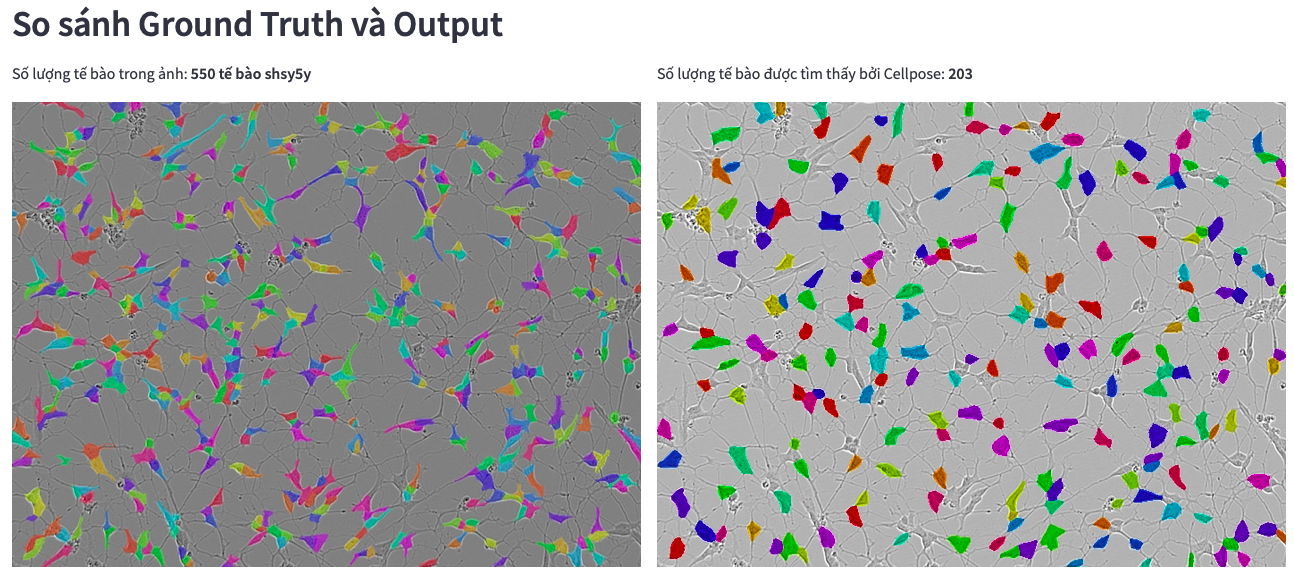
Cellpose does not detect label for instance
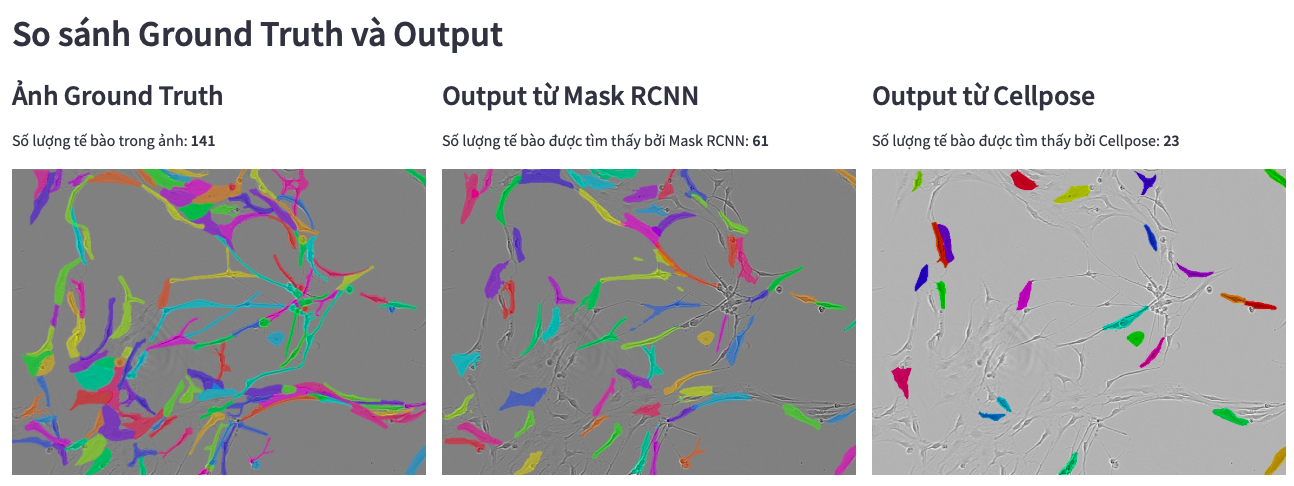
Comparing Mask RCNN and Cellpose

Mask RCNN: https://github.com/leekunhee/Mask_RCNN
Cellpose: https://github.com/MouseLand/cellpose
Streamlit: https://streamlit.io/