TransRegLipMet_CamelinaSeed
This repository contains a description of primary data, processed data, and codes used to obtain the main results/figures of this project.
Graphical abstract
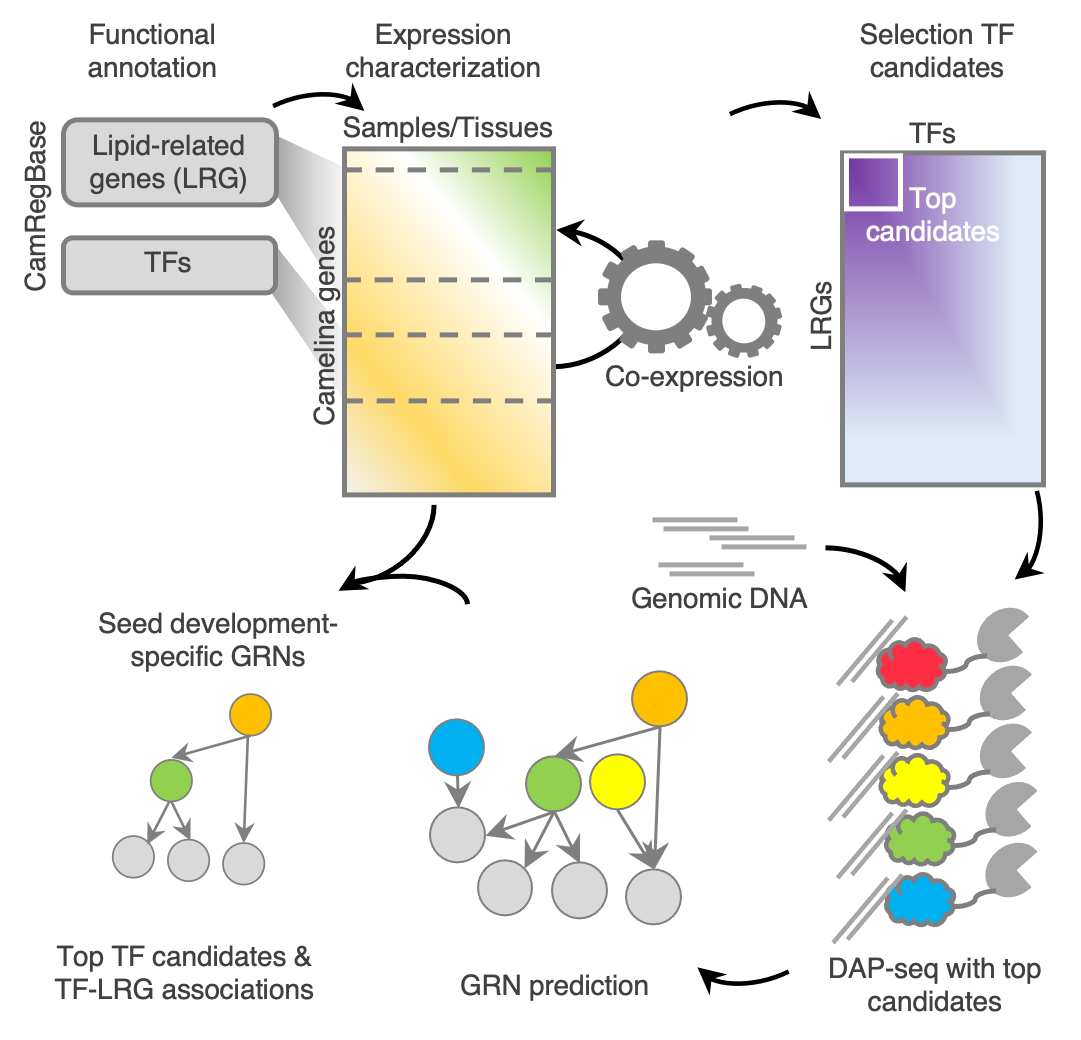
- Analysis of Seed expression data: RNA-seq
- Co-expression (MI) calcualations
- Identification of DNA-binding site (peaks)based on DAP-seq
Expression analysis of genes involved in lipid accumulation during Camelina seed development
Identification of candidate lipid transcriptional regulators by co-expression analysis
Establishing the DNA-binding landscape of the candidate transcription factors
Predicting gene targets for the selected TFs
Identified TFs associate with distinct aspects of lipid metabolism
Dynamic behavior of the predicted networks during seed development
