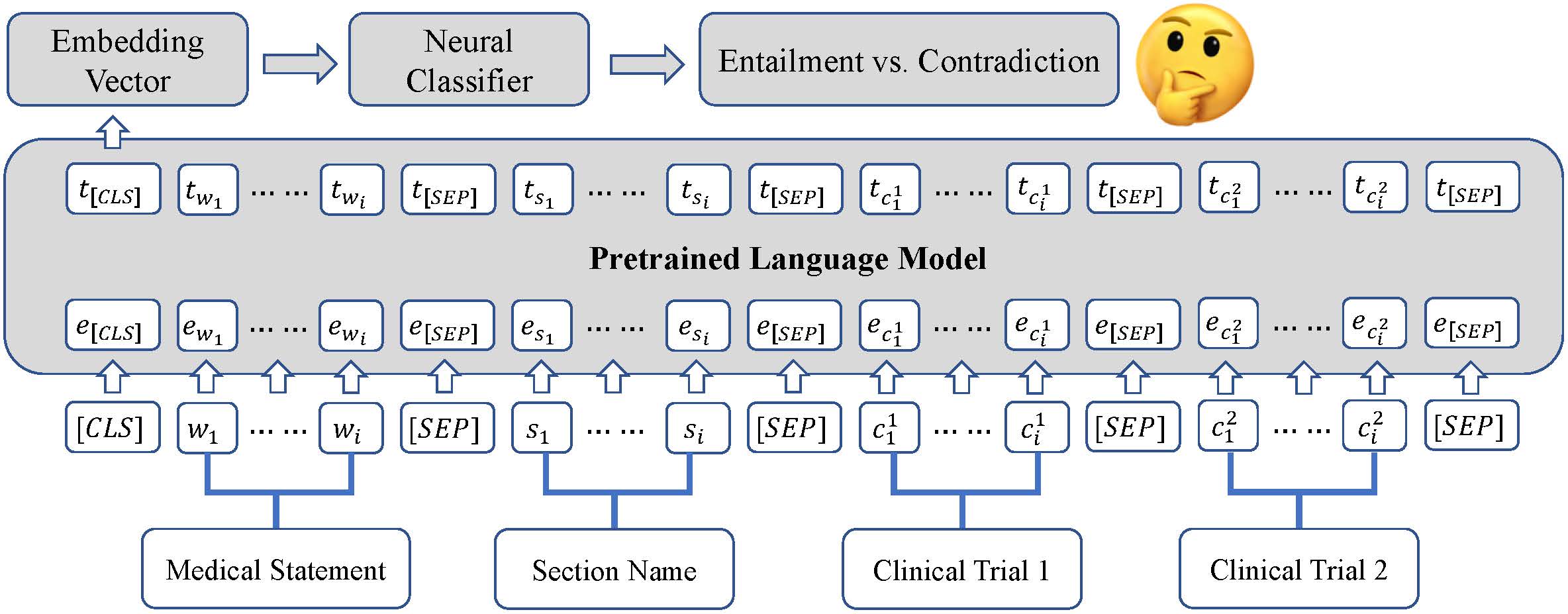
This is the official code repository for the SemEval2023 paper: KnowComp at SemEval-2023 Task 7: Fine-tuning Pretrained Language Models for Clinical Trial Entailment Identification. Our system officially ranks No.5th on the leaderboard and achieves an F1 score of 0.764.
Please download the full CT records dataset
from here.
Then, put the Complete_dataset folder under the data/ directory.
Required packages are listed in requirements.txt. Install them by running:
pip install -r requirements.txtTrain the model with the following command:
python train.py \
--ptlm microsoft/deberta-v3-large \
--lmn deberta \
--task_name atomic \
--data ./data/Complete_dataset \
--epoch 20 \
--eval_every 10 \
--prompt 2 \
--mode trn \
--gpu 0 \
--seed 621Our system's best model, which is based on a pretrained DeBERTa-v3-Large model, is
available here.
The from_check_point, model_dir, and tokenizer_dir arguments can be used to load the pretrained weight.
Please use the bibtex below for citing our paper:
@inproceedings{KnowCompNLI4CT,
author = {Weiqi Wang and
Baixuan Xu and
Tianqing Fang and
Lirong Zhang and
Yangqiu Song},
title = {KnowComp at SemEval-2023 Task 7: Fine-tuning Pre-Trained Language Models for Clinical Trial Entailment Identification},
year = {2023},
booktitle = {Proceedings of the 17th International Workshop on Semantic Evaluation, {SemEval} 2023}
}