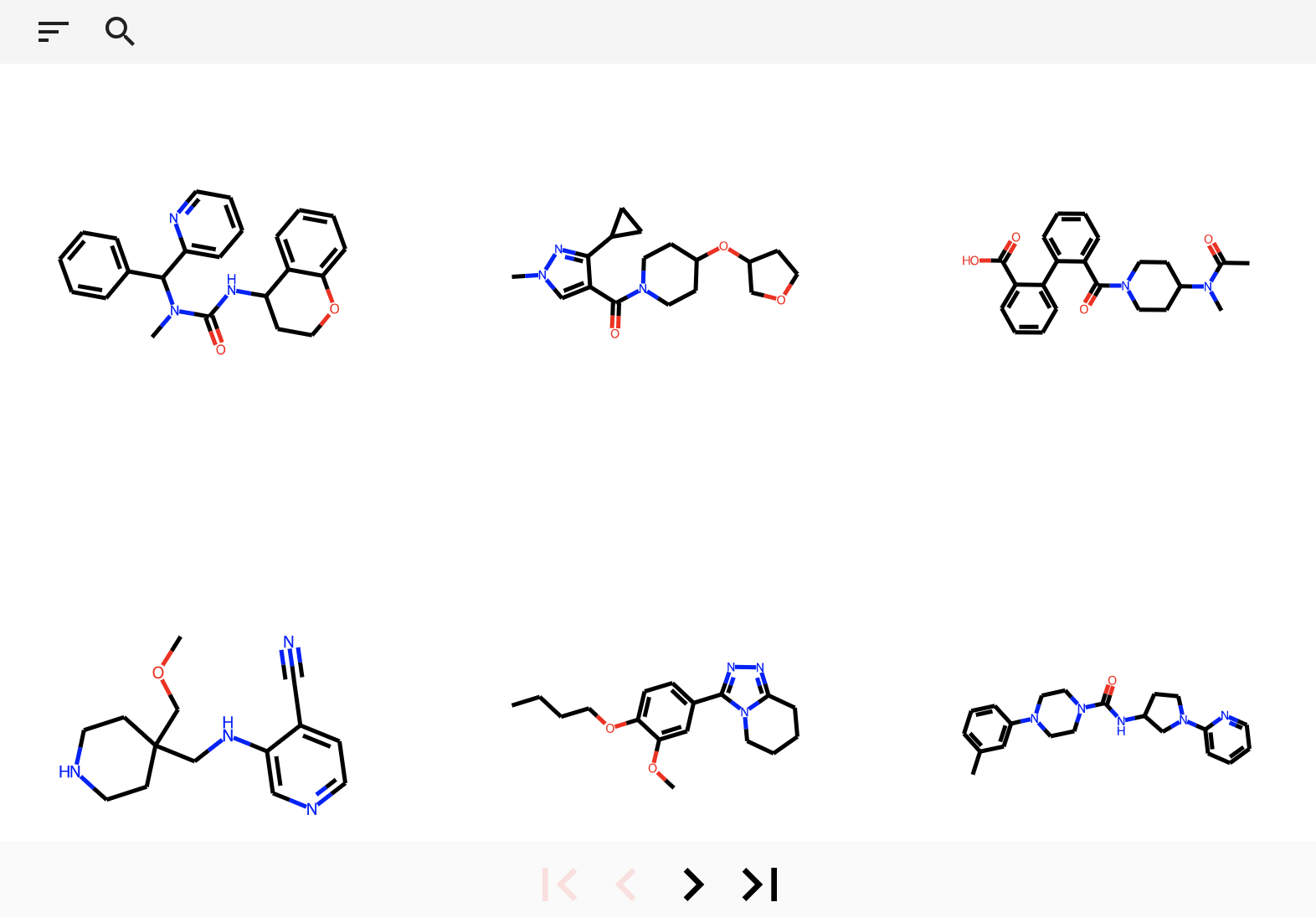
The splore framework aims to offer a simple graphical interface for scrolling through and exploring data sets of
molecules.
The GUI can easily be launched from the command line using the splore command:
# Load molecules from a local file
splore --file path-to-molecules.sdf
# Load molecules from a public QCArchive dataset
splore --qcf-dataset "OpenFF BCC Refit Study COH v1.0" --qcf-datatype basic
splore --qcf-dataset "OpenFF Rowley Biaryl v1.0" --qcf-datatype tdA full list of options can be printed using the --help flag:
splore --help
Usage: splore [OPTIONS]
Options:
--file FILE The path to the file of molecules (.smi,
.sdf, .sdf.gz) to display.
--qcf-dataset TEXT The name of a QC dataset stored in the public
QCArchive to extract the molecules to
visualize from.
--qcf-datatype [basic|opt|td] The type of dataset referenced by the
`--qcf-dataset` input.
--port INTEGER The port to run the GUI on. [default: 8000;
required]
--help Show this message and exit.The framework and its required dependencies can be installed using conda:
conda install -c conda-forge -c simonboothroyd sploreThe required dependencies for this framework can be installed using conda:
conda env create --name splore --file devtools/conda-envs/test-env.yamlafter which the GUI can be built by running:
python setup.py build_guiand the package installed in the normal ways, e.g.:
python setup.py developThe main package is release under the MIT license.
Copyright (c) 2021, Simon Boothroyd