Refed (Rib Extaction and Fracture Detection Model) by Youyou Jiang and Shiye Lei. Our model has a great performance on extracting ribs and detecting fracture from CT images. In Rib extraction, we can get 23.2 ribs (96.7%) in average for every patients; In Fraction Detection,recall: 71%, precision: 100%.
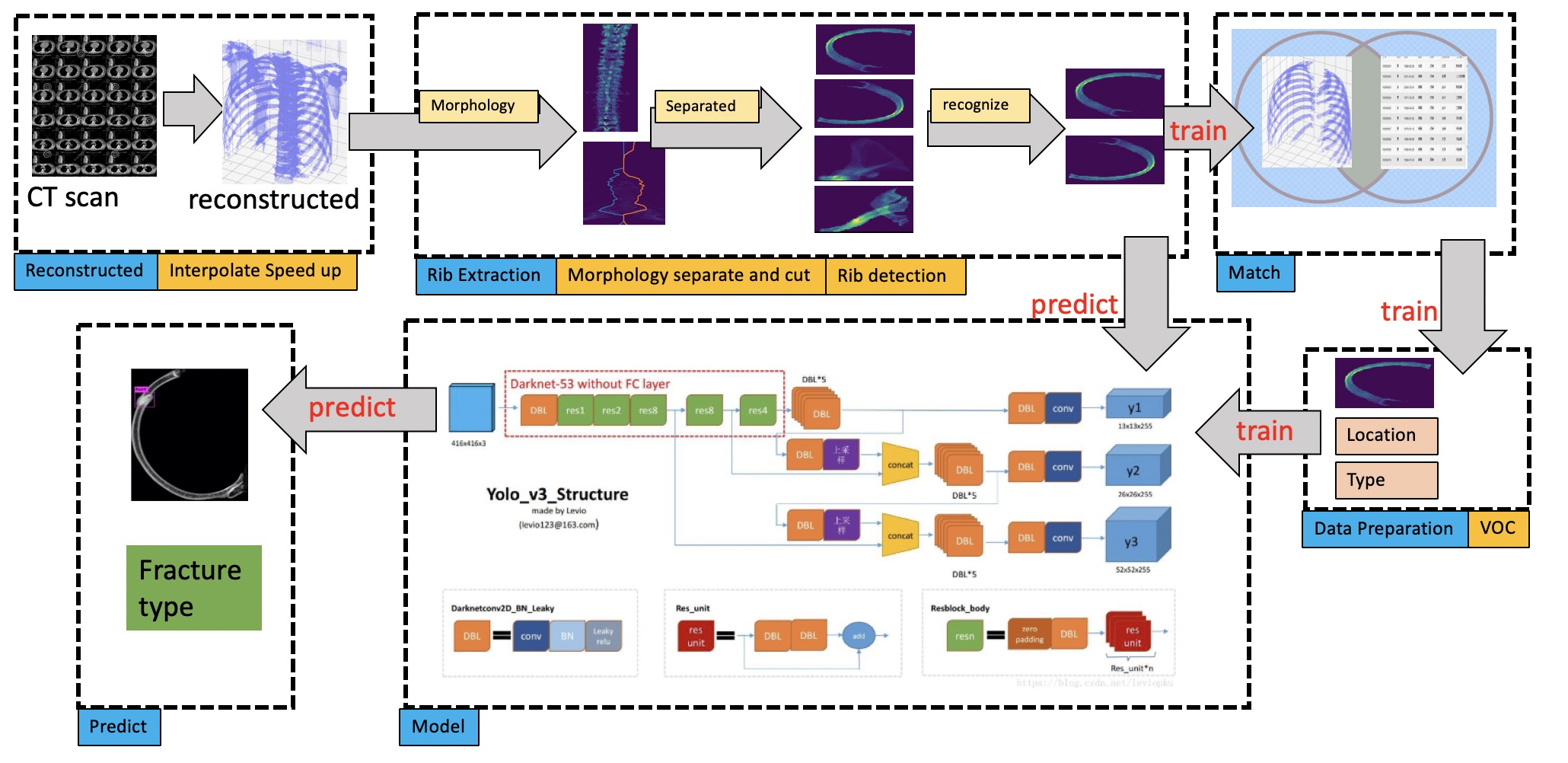
Refed is consist by two modules: rib extraction module and fracture detection module.
- read slices of CT data and reconstruct, see dcm read
- separate bones using morphology and recognize all ribs, see separated, code see ribs_obtain
- match labels to the ribs, see prepare_data, code see join
- (Optional,only for train) data preparation for train data, voc2007, see prepare_data, code seevoc2007
- yolo-v3 predict for demo/test or train for train data, see darknet/yolo-v3
- (Optional,only for test)predict scores.
-
Install tensorflow. It is required that you have access to GPUs, The code is tested with Ubuntu 16.04 Python 3.6+, CUDA 9.0.176 and cudnn 7.4.2.
-
Clone the repository
git clone https://github.com/jiangyy5318/medical-rib.git- Python dependencies (with
pip3 install) orpip3 install -r requirements.txt:
Deprecated==1.2.4
image==1.5.27
imageio==2.4.1
interval==1.0.0
lxml==4.2.5
matplotlib==3.0.0
numpy==1.15.2
opencv-python==3.4.3.18
pydicom==1.2.0
pyparsing==2.2.2
scikit-image==0.14.1
scikit-learn==0.20.0
scipy==1.1.0
six==1.11.0
nibabel==2.3.1
pandas==0.23.4
- Config darknet models
cd ${projects}/models
git clone https://github.com/pjreddie/darknet
cd darknet
vim Makefile
# update these three values.
GPU=1 # 0 if use cpu
CUDNN=1 # 0 if use cpu or cudnn not available
OPENCV=1 # 1 if use opencv else 0
NVCC=/path/to/nvcc
# build
make
cp darknet_cfg/cfg/* darknet/cfg/
cp darknet_cfg/data/hurt_voc.names darknet/data/
You can download pre-trained models, including GBDT model HERE,
GBDT features HERE and yolo-v3 models HERE. Put feature.pkl, gbdt.pkl under the project root path (${project}/experiments/cfgs) and
Put yolov3-voc_last.weights under the project root path (${projects}/models/darknet/backup)
mkdir [demo_dir]
mkdir [demo_dir]/pkl_cache/ # save desensitization data, pkl
mkdir [demo_dir]/ribs_df_cache # save collected ribs (sparsely saved)
mkdir [demo_dir]/voc_test_data # ribs projected on xoy
mkdir [demo_dir]/voc_test_predict # predict
cd ${projects}
# use raw data, slices of CT data
./experiments/scripts/demo.sh [DCM_PATH] [demo_dir]
# use desensitization data, pkl data, CT data
./experiments/scripts/demo.sh [PKL_PATH] [demo_dir]
./experiments/scripts/demo.sh [demo_dir]/pkl_cache/patient_id.pkl [demo_dir]
# DCM_PATH is folder path where CT slices existed.For dataSet structure and more information, follow the README under the preprocessing folder.
./experiments/scripts/nii_read.sh [DATA] [SLICING]
./experiments/scripts/dcm_read.sh [DATA]
./experiments/scripts/ribs_obtain.sh [LOGS_DIR] [FORMAT] [SLICING]
./experiments/scripts/prepare_data.sh
# [DATA] in {updated48labeled_1.31, all_labeled} originated from different batches of data, has been defined in dcm_read.sh and nii_read.sh
# [SLICING] in {0,1} is defined for interpolation interval, if 0, interval=1mm, else interval=slicing thickness
# [LOGS_DIR] used for debug
# [FORMAT] '.dcm'step ./experiments/scripts/ribs_obtain.sh will generate many separated bones and GBDT model will recognize all the ribs,
all the features for every bone will be saved in the path ./data/bone_info_merges, you can added the misclassified bone in the
path ./data/csv_files/update_err_bone_info.csv and run the below script, details see gbdt model
./experiments/scripts/generate_gbdt.sh cd ${projects}/models/darknet
# only download once
wget https://pjreddie.com/media/files/darknet53.conv.74
./darknet detector train ./cfg/hurt_voc.data ./cfg/yolov3-voc.cfg ./darknet53.conv.74 -gpus 0,1,2,3- refactoring