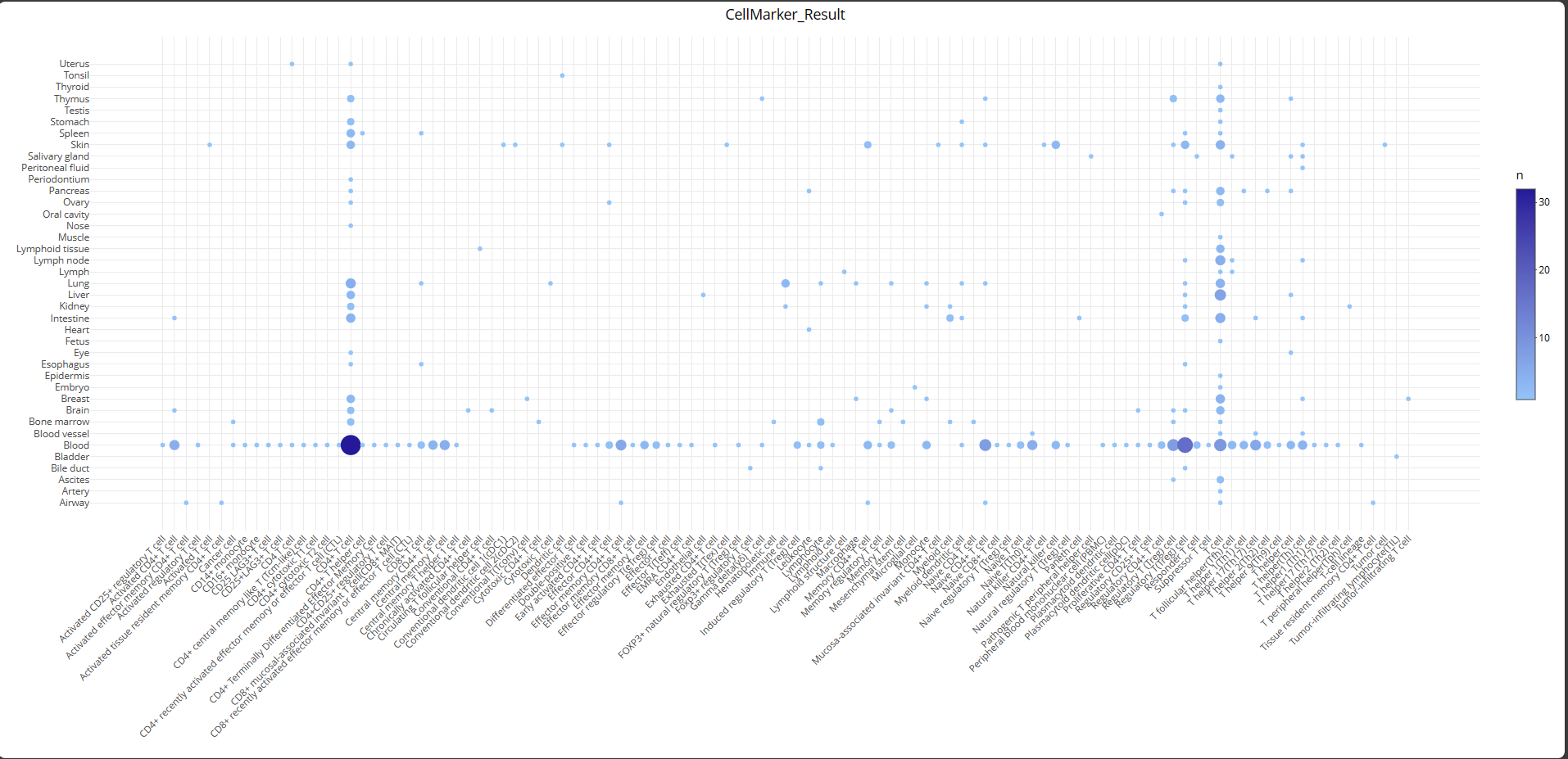
CellMarkerR is a localized tool that includes two core drawing functions. These functions are based on the database that is publicly available on the CellMarker website.(http://bio-bigdata.hrbmu.edu.cn/CellMarker/CellMarkerSearch.jsp)
Recommend R language >= 3.6
(Option A) Use R language code to install dependent packages.
# install.packages()
install.packages(c("plotly","htmlwidgets","wordcloud2","openxlsx"))
# BiocManager::install()
if (!requireNamespace("BiocManager", quietly = TRUE))
install.packages("BiocManager")
BiocManager::install(c("plotly","htmlwidgets","wordcloud2","openxlsx"))
(Option B) Use conda to install dependent packages.
conda install -c conda-forge r-plotly r-htmlwidgets r-wordcloud2 r-openxlsx
(Option A) Install the package directly from local.
install.packages("./CellMarkerR_1.0.0.tar.gz",repos=NULL,type="source")
(Option B) Install the package directly from github using the devtools package.
devtools::install_github("KIRA2ZERO/CellMarkerR")
library(CellMarkerR)
CellMarker = CellMarkerR$new()
# viewMarker
CellMarker$viewMarker(marker="CD4")
CellMarker$getViewMarkerChartResult()
CellMarker$getViewMarkerTableResult()
CellMarker$saveViewMarkerChartResult(filePath)
CellMarker$saveViewMarkerTableResult(filePath)
# viewCell
CellMarker$viewCell()
CellMarker$getViewCellChartResult()
CellMarker$getViewCellTableResult()
CellMarker$getViewCellRankResult()
CellMarker$saveViewCellChartResult(filePath)
CellMarker$saveviewCellTableResult(filePath)
CellMarker$saveviewCellRankResult(filePath)