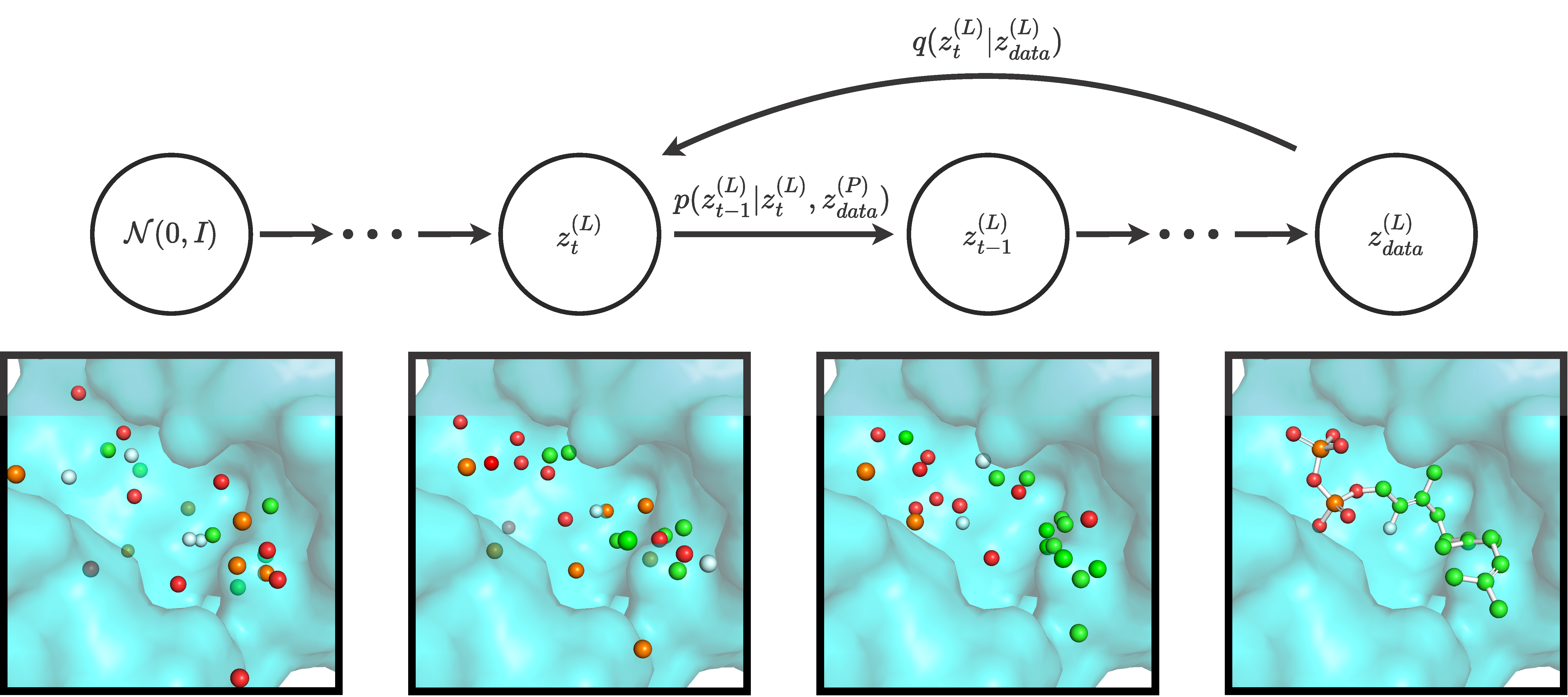
Official implementation of DiffSBDD, an equivariant model for structure-based drug design, by Arne Schneuing*, Yuanqi Du*, Charles Harris, Arian Jamasb, Ilia Igashov, Weitao Du, Tom Blundell, Pietro Lió, Carla Gomes, Max Welling, Michael Bronstein & Bruno Correia.
- RDKit
- PyTorch
- BioPython
- imageio
- Scipy
- wandb
- torch-scatter
- PyTorch Lightning
- openbabel
- QuickVina 2
- MGLTools
conda create -n sbdd-env
conda activate sbdd-env
conda install pytorch cudatoolkit=10.2 -c pytorch
conda install -c conda-forge pytorch-lightning
conda install -c conda-forge wandb
conda install -c conda-forge rdkit
conda install -c conda-forge biopython
conda install -c conda-forge imageio
conda install -c anaconda scipy
conda install -c pyg pytorch-scatter
conda install -c conda-forge openbabelThe code was tested with the following versions
| Software | Version |
|---|---|
| Python | 3.10.4 |
| CUDA | 10.2.89 |
| PyTorch | 1.12.1 |
| PyTorch Lightning | 1.7.4 |
| WandB | 0.13.1 |
| RDKit | 2022.03.2 |
| BioPython | 1.79 |
| imageio | 2.21.2 |
| SciPy | 1.7.3 |
| PyTorch Scatter | 2.0.9 |
| OpenBabel | 3.1.1 |
For docking, install QuickVina 2:
wget https://github.com/QVina/qvina/raw/master/bin/qvina2.1
chmod +x qvina2.1 We need MGLTools for preparing the receptor for docking (pdb -> pdbqt) but it can mess up your conda environment, so I recommend to make a new one:
conda create -n mgltools -c bioconda mgltoolsDownload and extract the dataset as described by the authors of Pocket2Mol: https://github.com/pengxingang/Pocket2Mol/tree/main/data
Process the raw data using
python process_crossdock.py <crossdocked_dir> --no_HDownload the dataset
wget http://www.bindingmoad.org/files/biou/every_part_a.zip
wget http://www.bindingmoad.org/files/biou/every_part_b.zip
wget http://www.bindingmoad.org/files/csv/every.csv
unzip every_part_a.zip
unzip every_part_b.zipProcess the raw data using
python process_bindingmoad.py <bindingmoad_dir>or, to suppress warnings,
python -W ignore process_bindingmoad.py <bindingmoad_dir>Starting a new training run:
python -u train.py --config <config>.ymlResuming a previous run:
python -u train.py --config <config>.yml --resume <checkpoint>.ckptTo sample small molecules for a given pocket with a trained model use the following command:
python generate_ligands.py <checkpoint>.ckpt --pdbfile <pdb_file>.pdb --outdir <output_dir> --resi_list <list_of_pocket_residue_ids>For example:
python generate_ligands.py last.ckpt --pdbfile 1abc.pdb --outdir results/ --resi_list A:1 A:2 A:3 A:4 A:5 A:6 A:7 Alternatively, the binding pocket can also be specified based on a reference ligand in the same PDB file:
python generate_ligands.py <checkpoint>.ckpt --pdbfile <pdb_file>.pdb --outdir <output_dir> --ref_ligand <chain>:<resi>Optional flags:
| Flag | Description |
|---|---|
--n_samples |
Number of sampled molecules |
--all_frags |
Keep all disconnected fragments |
--sanitize |
Sanitize molecules (invalid molecules will be removed if this flag is present) |
--relax |
Relax generated structure in force field |
--resamplings |
Inpainting parameter (doesn't apply if conditional model is used) |
--jump_length |
Inpainting parameter (doesn't apply if conditional model is used) |
test.py can be used to sample molecules for the entire testing set:
python test.py <checkpoint>.ckpt --test_dir <bindingmoad_dir>/processed_noH/test/ --outdir <output_dir> --fix_n_nodesUsing the optional --fix_n_nodes flag lets the model produce ligands with the same number of nodes as the original molecule. Other optional flags are identical to generate_ligands.py.
For assessing basic molecular properties create an instance of the MoleculeProperties class and run its evaluate method:
from analysis.metrics import MoleculeProperties
mol_metrics = MoleculeProperties()
all_qed, all_sa, all_logp, all_lipinski, per_pocket_diversity = \
mol_metrics.evaluate(pocket_mols)evaluate() expects a list of lists where the inner list contains all RDKit molecules generated for one pocket.
For computing docking scores, run QuickVina as described below.
First, convert all protein PDB files to PDBQT files using MGLTools
conda activate mgltools
cd analysis
python docking_py27.py <bindingmoad_dir>/processed_noH/test/ <output_dir> bindingmoad
cd ..
conda deactivateThen, compute QuickVina scores:
conda activate sbdd-env
python analysis/docking.py --pdbqt_dir <docking_py27_outdir> --sdf_dir <test_outdir> --out_dir <qvina_outdir> --write_csv --write_dict@article{schneuing2022structure,
title={Structure-based Drug Design with Equivariant Diffusion Models},
author={Schneuing, Arne and Du, Yuanqi and Harris, Charles and Jamasb, Arian and Igashov, Ilia and Du, Weitao and Blundell, Tom and Li{\'o}, Pietro and Gomes, Carla and Welling, Max and others},
journal={arXiv preprint arXiv:2210.13695},
year={2022}
}