Official repo of the 2024 paper: HIT: Estimating Internal Human Implicit Tissues from the Body Surface
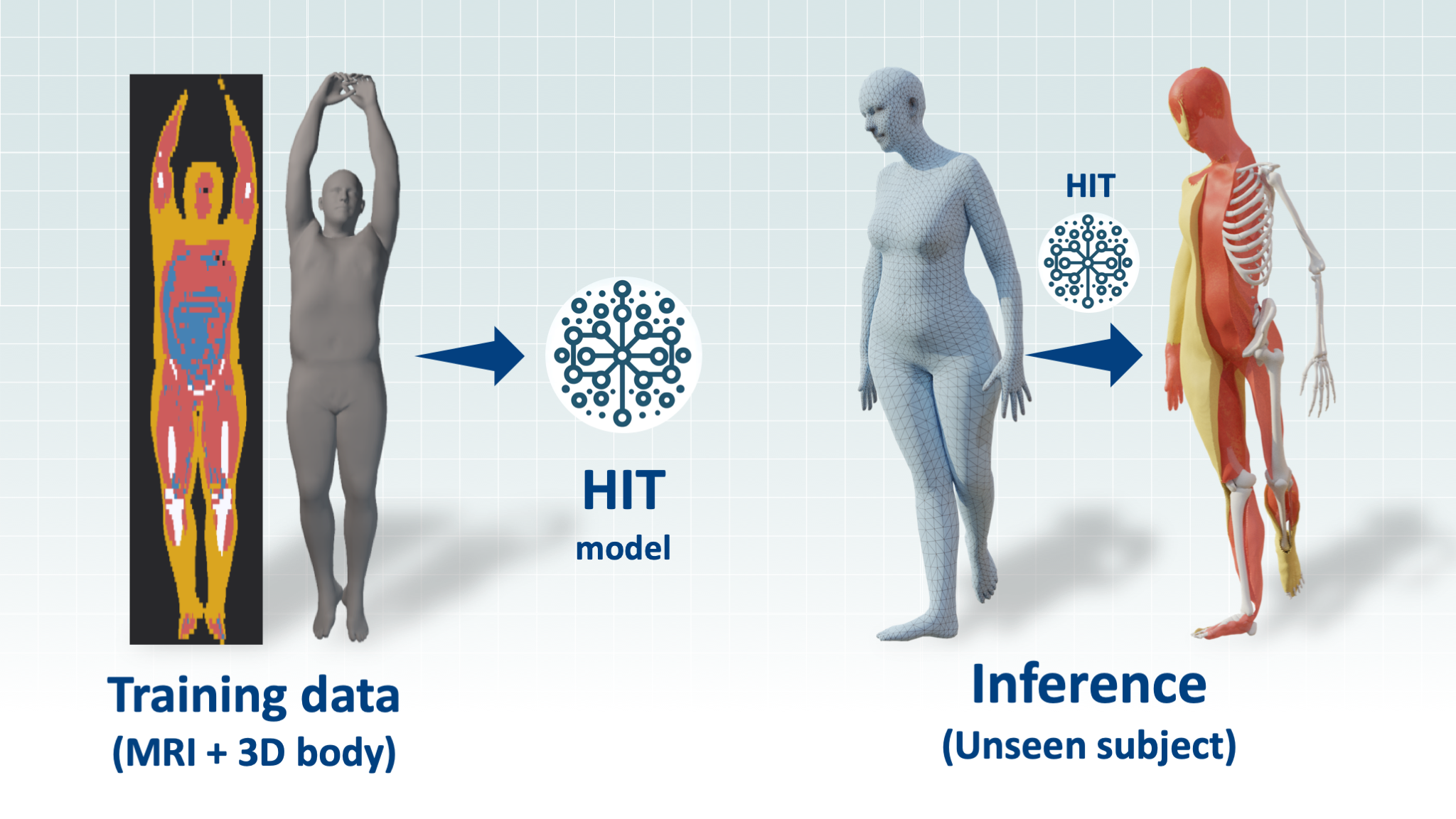
HIT is a neural network that learns to infer the internal tissues of the human body from its surface. The input is a 3D body represented as SMPL parameters (a shape vector β and a pose vector θ) and the output is an implicit function, that given a point in space, returns the probability of this point being part of the following tissue:
- lean tissue (muscles, organs) (LT)
- adipose tissue (subcutaneous) (AT)
- bone (BT)
- empty space (inside the lungs, outside the body) (E)
The implicit HIT function can also be used to generate the 3D mesh of the tissues using the marching cube algorithm.
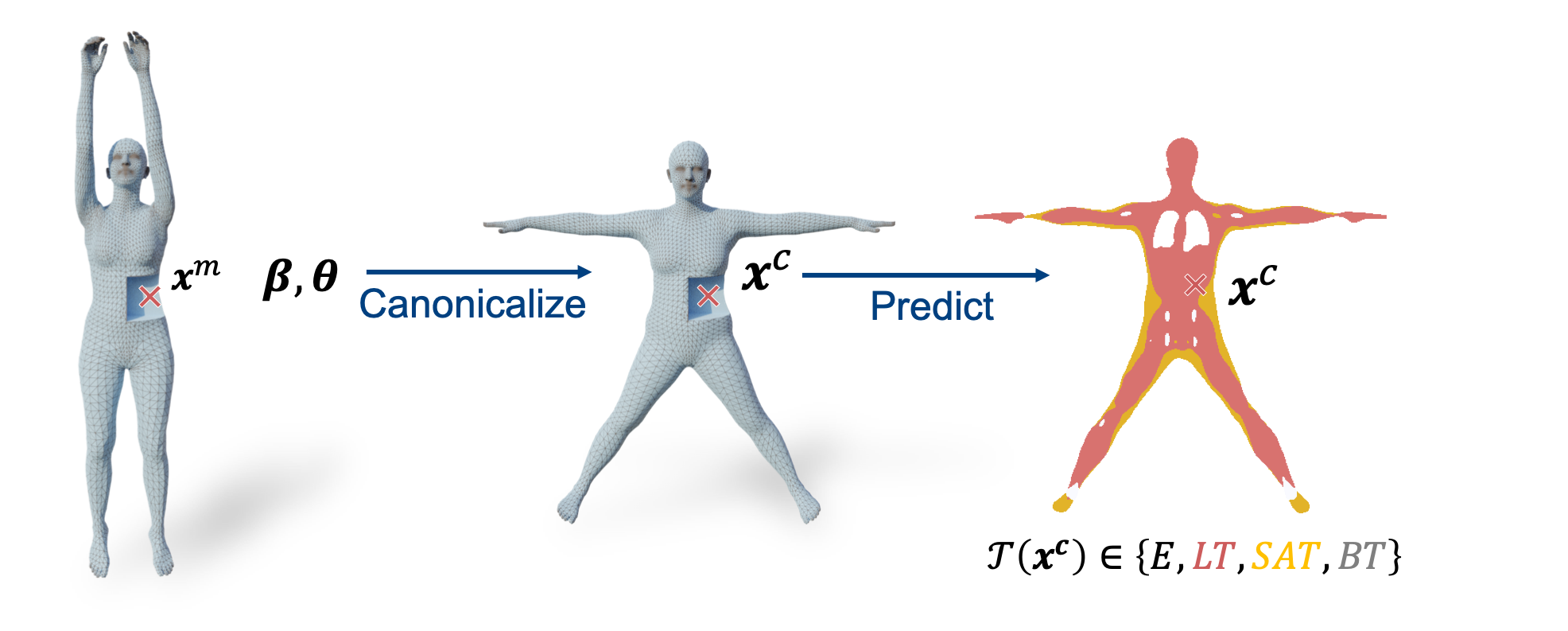
The figure below illustrates our approach. Given a posed body shape (β, θ), and a 3D point xm, HIT first canonicalizes this point to the corresponding location xc inside an average template body, and then predicts the class of the tissue at location xc.
This repo contains the code to train and test HIT on the HIT dataset (available on our project page). It also contains some demo code to infer the tissue meshes for a given SMPL body.
This code is tested on Ubuntu 20.04 with Python 3.8. We do not guarantee that it will work on other systems.
git clone https://github.com/MarilynKeller/HIT
cd hitpython3.8 -m venv hit_venv
source hit_venv/bin/activate
pip install -r requirements.txtCheck your CUDA toolkit version
nvcc --versionInstall torch depending on CUDA toolkit version. See: https://pytorch.org/get-started/previous-versions/ .
For 11.8:
pip install torch==2.2.0 torchvision==0.17.0 torchaudio==2.2.0 --index-url https://download.pytorch.org/whl/cu118Install relevant packages
pip install -r requirements.txt
pip install git+https://github.com/MPI-IS/mesh.git
pip install git+https://github.com/mattloper/chumpy
pip install -e .The LEAP package is used for its marching cube implementation and creating ground truth occupancy. Install it with:
cd hit
mkdir external
cd external
git clone https://github.com/neuralbodies/leap.git
cd leap
python setup.py build_ext --inplace
pip install -e .Download the SMPL model from https://smpl.is.tue.mpg.de/ and update the path smplx_models_path in hit_config.py to the proper path.
The folder hierarchy should be the following:
${MODELS}
├── smpl
│ ├── SMPL_FEMALE.pkl
│ └── SMPL_MALE.pkl
│ └── SMPL_NEUTRAL.pkl
├── smplh
│ ├── SMPLH_FEMALE.pkl
│ └── SMPLH_MALE.pkl
└── smplx
├── SMPLX_FEMALE.npz
├── SMPLX_FEMALE.pkl
├── SMPLX_MALE.npz
├── SMPLX_MALE.pkl
├── SMPLX_NEUTRAL.npz
└── SMPLX_NEUTRAL.pkl
You can download the pretrained model checkpoints for male and female from the Download tab at [https://hit.is.tue.mpg.de/].
Create a folder HIT/pretrained and place the pretrained models inside, or edit the path trained_models_folder in hit_config.py.
You should have the following hierarchy:
${HIT}
├── pretrained
│ ├── hit_female
│ ├── ckpt
│ └── config.yaml
│ └── hit_male
│ └── ...
├── hit
└── ...
python demos/infer_smpl.py --exp_name=hit_male --to_infer smpl_templateYou can also evaluate the different values (occupancy, skinning weights, beta displacement) on 2D slices inside the body for a given body shape (betas):
python demos/infer_smpl.py --exp_name=hit_male --to_infer smpl_template --ckpt_choice=best --output='slices' --betas -2.0 0.0This will generate the per slice prediction for a body shape of shape beta=[-2.0, 0.0, 0.0, ...].
For example, this generates the occupancy for the frontal (x,0,z) slice:
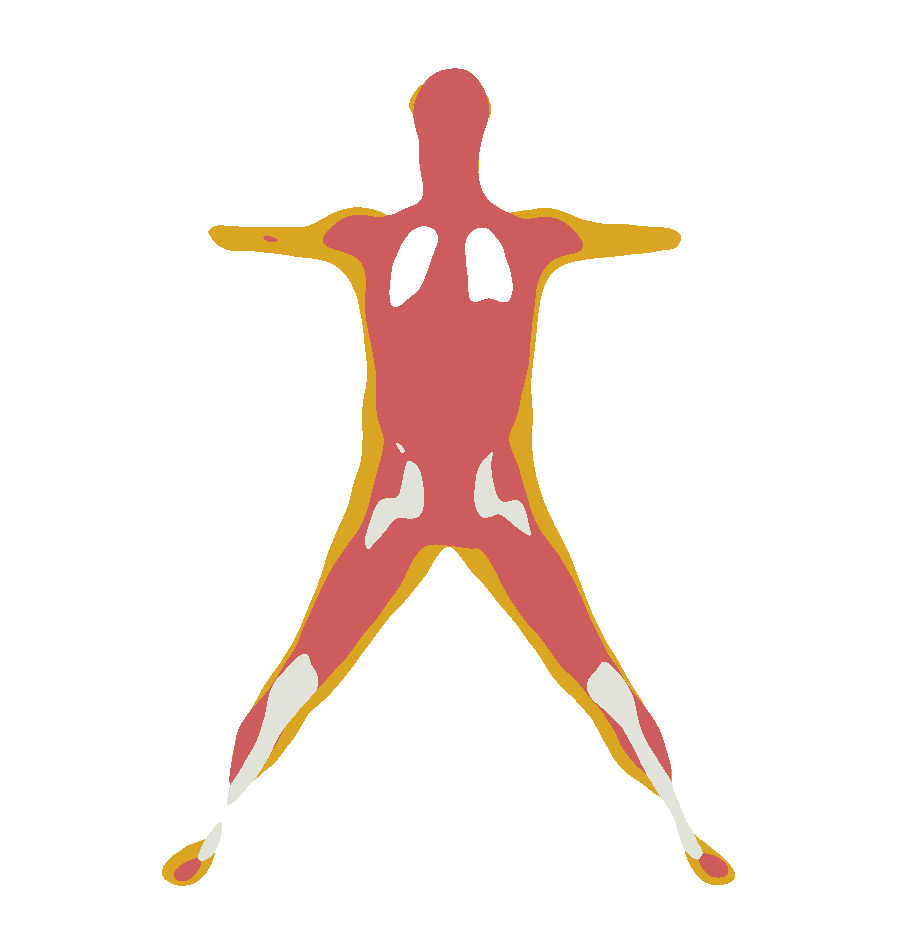
python demos/infer_smpl.py --exp_name=hit_male --to_infer smpl_file --target_body assets/standing.pklYou can download the HIT dataset from HIT project page [https://hit.is.tue.mpg.de/], in the Download tab. Then add the path to the dataset folder in hit_config.py.
We also provide a huggingface version of the HIT dataset. To use it instead, just set data_cfg.huggingface = False in your training command line and the dataset will be downloaded automatically from the huggingface hub.
This dataset contains data for 127 males and 191 females. For each subject, it contains:
{
'gender': "gender of the subject",
'mri_seg': "annotated array with the labels 0,1,2,3",
'mri_labels': "dictionary of mapping between label integer and name",
'mri_seg_dict': "a dictionary of individual masks of the different tissues (LT, AT, BT, ...)",
'resolution': "per slice resolution in meters",
'center': "per slice center, in pixels",
'smpl_dict': "dictionary containing all the relevant SMPL parameters of the subject along with mesh faces
and vertices ('verts': original fit, 'verts_free': compressed fit")
}In the hit_config.py file, set the path to the dataset folder in packaged_data_folder.
You can visualize a subject of the dataset using:
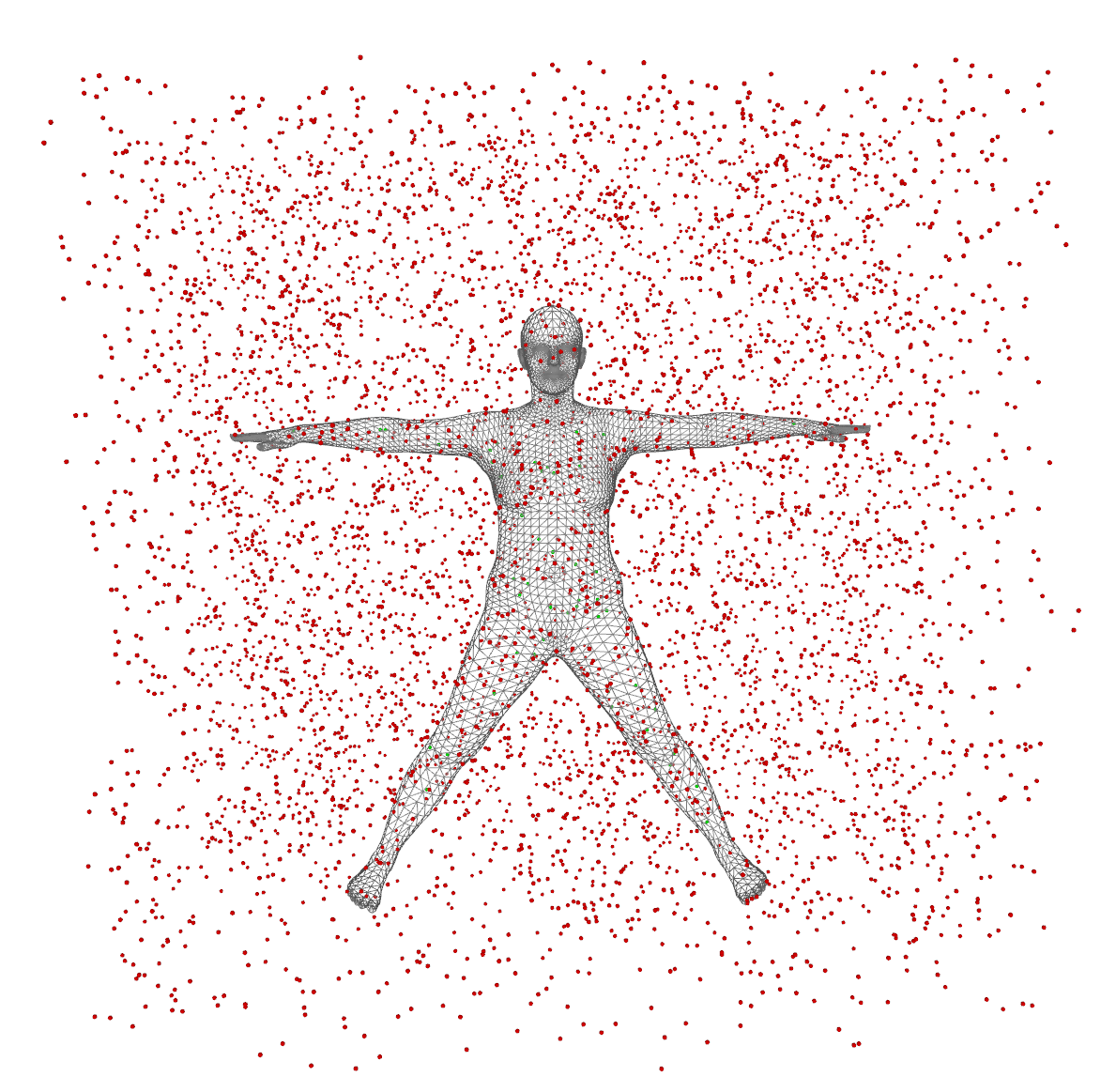
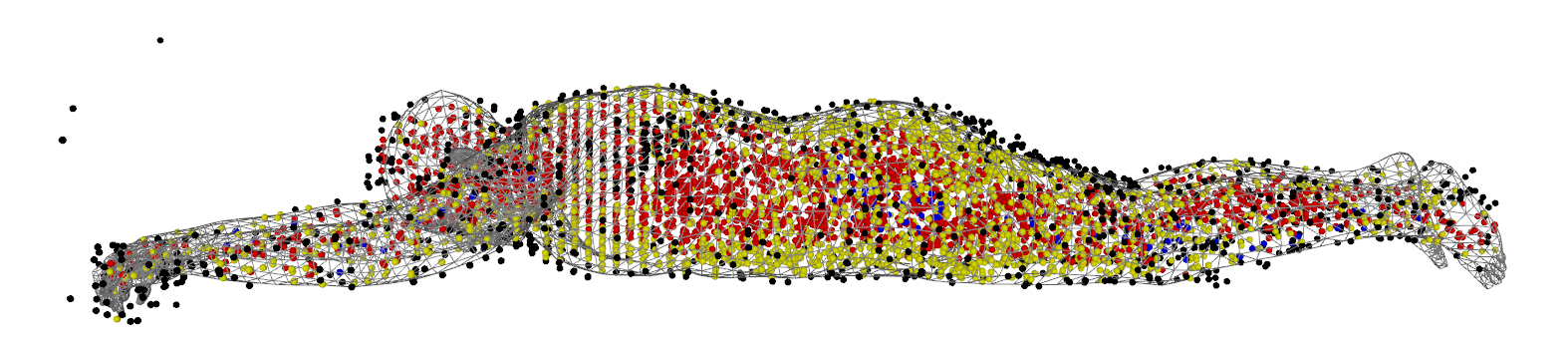
python demos/load_data.py -s train -i 0 --gender female -DThis shows the tight SMPL fit to the subject and the MRI points sampled for one iteration of the training, colored according to their ground truth label:
- black: E (empty)
- red: LT (lean tissue)
- yellow: AT (adipose tissue)
- blue: BT (bone tissue)
After closing the window, this will show the points sampled in the SMPL canonical space for one iteration of the training. In red are the points outside the SMPL template mesh, in green the points inside.
The first loading of the dataset requires sampling which takes time. We do this once and then cache the result for further fast loading. To force the recaching of the dataset for a gender, run:
python demos/load_data.py -s train -i 0 --gender male --all --recache To load the cached training dataset in memory then, you will need at least 25 GB of RAM for males and 36 GB for females.
The default training parameters are in config.yaml. The project uses hydra to load this config file.
Each parameter of this file can be overwritten through command line arguments.
The training is logged on Weight and Biasis (https://wandb.ai/). You can set the wandb entity and project name in hit_config.py.
To retrain HIT, first you need to pretrain, for each gender, the submodules on generated SMPL meshes to learn the LBS and inverse beta fields. Here is the command for females.
python hit/train.py exp_name=pretrained_female smpl_cfg.gender=female train_cfg=train_smpl data_cfg.synt_style=random data_cfg.synthetic=True trainer.max_epochs=200 train_cfg.networks.lbs.dropout=0.001 Once this trained for a gender, edit the path to the pretrained network pretrained_female_smpl in hit/hit_config.py.
You can train HIT for this gender:
python hit/train.py exp_name=hit_female smpl_cfg.gender=female Here hit_female is the name of the experiment. This training will be logged and saved in a folder with this name. Note that if you launch a new training with the same name, the last checkpoint with this name will be loaded.
Debugging
To debug you might want to turn off wandb and use a single worker so that breakpoints are catched:
python train_mri.py exp_name=hit_female smpl_cfg.gender=female wdboff=True train_cfg.num_workers=0
Generate the test metrics (Accuracy, IOU, Dice score)
python hit/train.py exp_name=hit_female smpl_cfg.gender=female run_eval=True wdboff=TrueWe thank the authors of the COAP and gDNA for their codebase. HIT is built on top of these two projects. We also thank Soubhik Sanyal for his help on the project.
If you use this code, please cite the following paper:
@inproceedings{keller2024hit,
title = {{HIT}: Estimating Internal Human Implicit Tissues from the Body Surface},
author = {Keller, Marilyn and Arora, Vaibhav and Dakri, Abdelmouttaleb and Chandhok, Shivam and Machann, J{\"u}rgen and Fritsche, Andreas and Black, Michael J. and Pujades, Sergi},
booktitle = {IEEE/CVF Conf.~on Computer Vision and Pattern Recognition (CVPR)},
pages = {3480--3490},
month = jun,
year = {2024},
month_numeric = {6}
}
For more questions, please contact hit@tue.mpg.de
This code repository in the provided License. For the licensing of the retrained models and the dataset, please refer to the HIT project page [https://hit.is.tue.mpg.de/].