Enhancing Cardiac Magnetic Resonance Segmentation using Transfer Learning and Stochastic Weight Averaging Ensembling
This repository is the official implementation of Enhancing Cardiac Magnetic Resonance Segmentation using Transfer Learning and Stochastic Weight Averaging Ensembling.
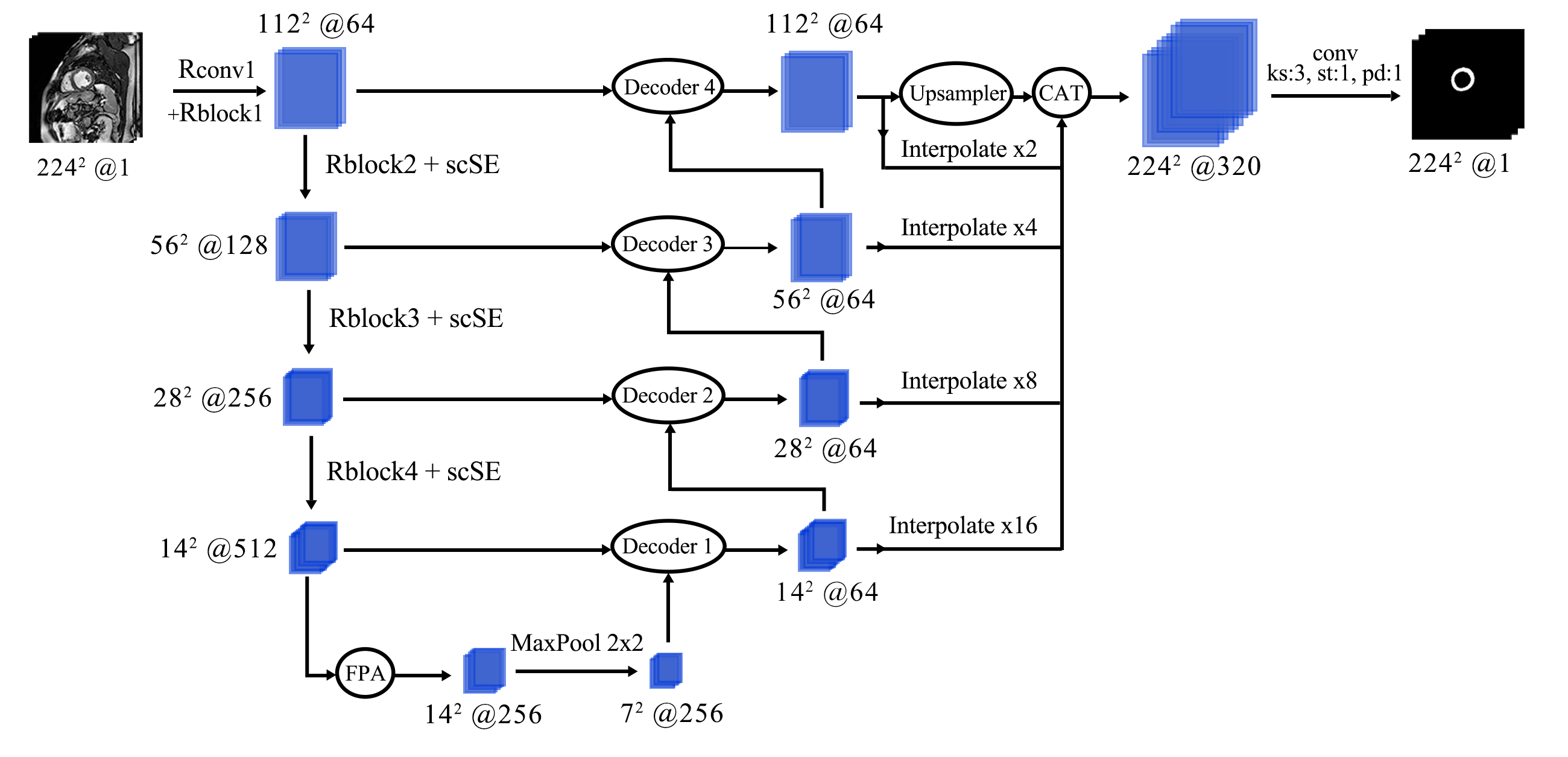
Network architecture of the segmentator. We use scSE and FPA modules for pixel-precise attention. Pretrained Resnet modules are denoted as Rblocks. Batch dimension is omitted. Decoder blocks use transposed convolutions to enlarge previous decoder steps, concatenate them with corresponding encoder blocks, and then merge all features through a convolution.
The network was implemented using the Pytorch framework and trained in parallel with two NVIDIA RTX 2080 GPU. For data augmentation, we used the albumentations library.
First we will install the packages and virtualenv if we do not have it:
# Step 1: We update the system repositories
sudo apt-get update
# Step 2: Install pip for Python 3
sudo apt-get install build-essential libssl-dev libffi-dev python-dev
sudo apt install python3-pip
# Step 3: Use pip to install virtualenv
sudo pip3 install virtualenv Next, we create our virtual environment and activate it:
# Create the virtual environment 'cmr_env'
python3 -m venv cmr_env
# Activate the virtual environment
source cmr_env/bin/activateFinally, to install requirements:
pip install -r requirements.txt
To train and evaluate the model(s) in the paper, run this command for LVSC:
./scripts/lvsc2d.sh
Or this command for Open M&Ms:
./scripts/mms2d.sh
Our model achieves the following performance on :
| Method | Jaccard | Dice |
|---|---|---|
| Our approach | 0.744 | 0.845 |
| + Transfer Learning | 0.756 | 0.854 |
| + SWA | 0.765 | 0.860 |
| + Transfer Learning & SWA | 0.771 | 0.865 |
| Method | Jaccard | Dice | Hausdorff | ASSD |
|---|---|---|---|---|
| Our approach | 0.722 | 0.827 | 21.359 | 1.592 |
| + Transfer Learning | 0.738 | 0.840 | 19.540 | 1.589 |
| + SWA | 0.771 | 0.865 | 15.821 | 1.302 |
| + Transfer Learning & SWA | 0.779 | 0.870 | 12.897 | 1.137 |
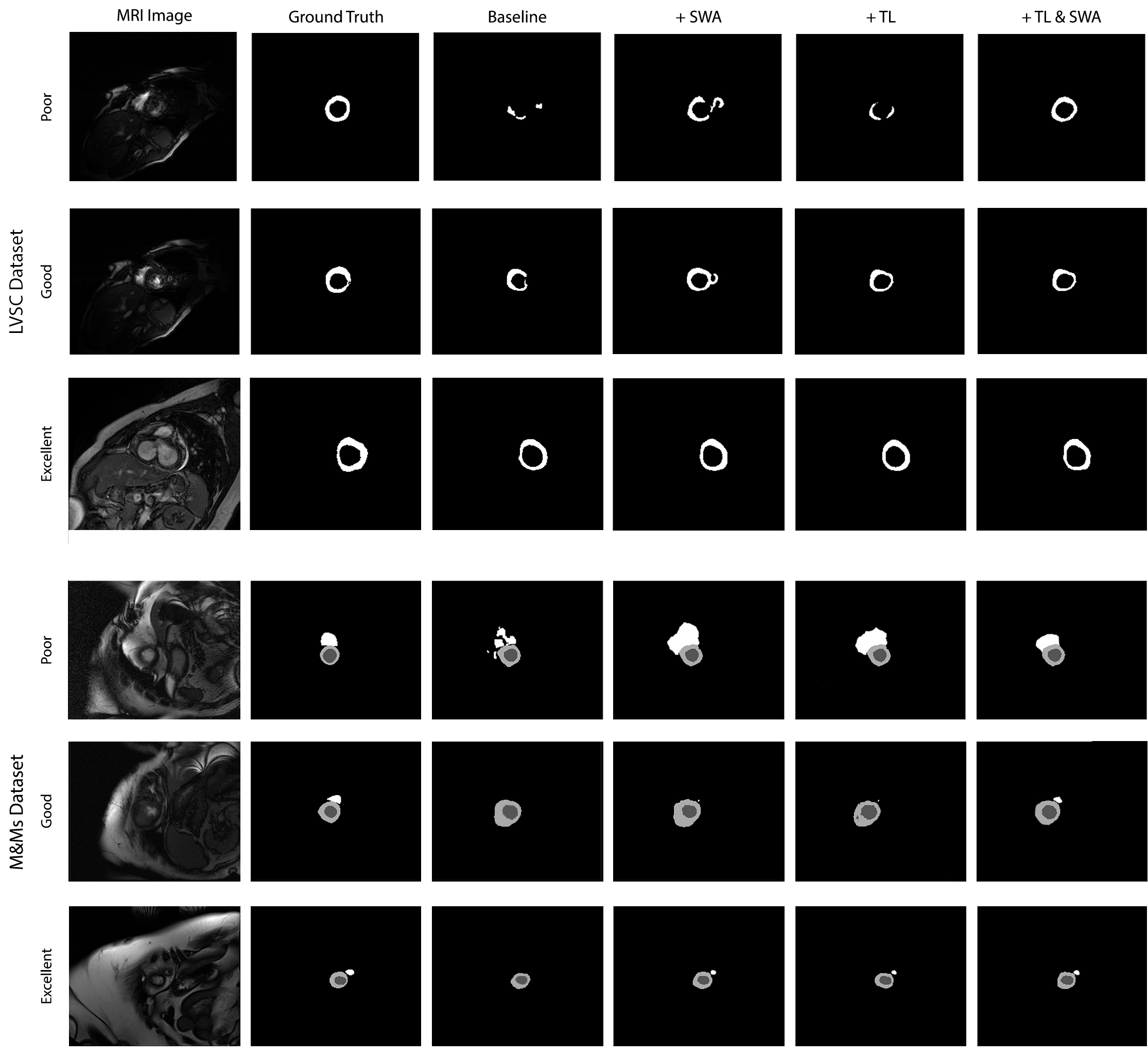
Qualitative results comparing the baseline approach, adding Transfer Learning (TL) and SWA techniques. Samples are grouped as poor (< 0.5 Jaccard), good (between 0.5 and 0.8 Jaccard), and excellent (> 0.8 Jaccard) when using the baseline approach.
The authors thank EU-FEDER Comunitat Valenciana 2014-2020 grant IDIFEDER/2018/025.