A lightweight and super fast C/C++ library for sequence alignment using edit distance.
Calculating edit distance of two strings is as simple as:
edlibAlign("hello", 5, "world!", 6, edlibDefaultAlignConfig()).editDistance;Edlib is also available for Python 
There are third-party bindings to edlib in other languages as well:
- Edlib.jl, a Julia package created and supported by Christopher Rowley (@cjdoris)
- edlibR, an R package created and supported by Evan Biederstedt (@evanbiederstedt)
- ruby-edlib, a Ruby package created and supported by @kojix2
- Text::Levenshtein::Edlib, a Perl wrapper created and supported by Marius Gavrilescu
- Calculates edit distance (Levenshtein distance).
- It can find optimal alignment path (instructions how to transform first sequence into the second sequence).
- It can find just the start and/or end locations of alignment path - can be useful when speed is more important than having exact alignment path.
- Supports multiple alignment methods: global(NW), prefix(SHW) and infix(HW), each of them useful for different scenarios.
- You can extend character equality definition, enabling you to e.g. have wildcard characters, to have case insensitive alignment or to work with degenerate nucleotides.
- It can easily handle small or very large sequences, even when finding alignment path, while consuming very little memory.
- Super fast thanks to Myers's bit-vector algorithm.
- Features
- Building
- Using Edlib in your project
- Usage and examples
- API documentation
- Alignment methods
- Aligner
- Running tests
- Time and space complexity
- Test data
- Development and contributing
- Publication
- Acknowledgements
You can use Edlib in you project by either directly copying header and source files from edlib/, or by linking Edlib library (see Building for instructions how to build Edlib libraries).
In any case, only thing that you have to do in your source files is to include edlib.h.
To get you started quickly, let's take a look at a few ways to get simple Hello World project working.
Our Hello World project has just one source file, helloWorld.cpp file, and it looks like this:
#include <cstdio>
#include "edlib.h"
int main() {
EdlibAlignResult result = edlibAlign("hello", 5, "world!", 6, edlibDefaultAlignConfig());
if (result.status == EDLIB_STATUS_OK) {
printf("edit_distance('hello', 'world!') = %d\n", result.editDistance);
}
edlibFreeAlignResult(result);
}Running it should output edit_distance('hello', 'world!') = 5.
Here we directly copied edlib/ directory to our project, to get following project structure:
edlib/ -> copied from edlib/
include/
edlib.h
src/
edlib.cpp
helloWorld.cpp -> your program
Since helloWorld is a c++ program, we can compile it with just one line: c++ helloWorld.cpp edlib/src/edlib.cpp -o helloWorld -I edlib/include.
If hello world was a C program, we would compile it like this:
c++ -c edlib/src/edlib.cpp -o edlib.o -I edlib/include
cc -c helloWorld.c -o helloWorld.o -I edlib/include
c++ helloWorld.o edlib.o -o helloWorld
Instead of copying edlib source files, you could copy static library (check Building on how to create static library). We also need to copy edlib header files. We get following project structure:
edlib/ -> copied from edlib
include/
edlib.h
edlib.a
helloWorld.cpp -> your program
Now you can compile it with c++ helloWorld.cpp -o helloWorld -I edlib/include -L edlib -ledlib.
Alternatively, you could avoid copying any Edlib files and instead install libraries by running sudo make install (check Building for exact instructions depending on approach you used for building). Now, all you have to do to compile your project is c++ helloWorld.cpp -o helloWorld -ledlib.
If you get error message like cannot open shared object file: No such file or directory, make sure that your linker includes path where edlib was installed.
If you are using CMake for compilation, we suggest adding edlib as a git submodule with the command git submodule add https://github.com/martinsos/edlib vendor/edlib. Afterwards, modify your top level CMakeLists.txt file accordingly:
add_subdirectory(vendor/edlib EXCLUDE_FROM_ALL)
target_link_libraries(your_exe edlib) # or target_link_libraries(your_exe edlib)
The add_subdirectory command adds a folder to the build tree, meaning it will run CMakeLists.txt from the included folder as well. Flag EXCLUDE_FROM_ALL disables building (and instalment) of targets in the added folder which are not needed in your project. In the above example only the (static) library edlib will be build, while edlib-aligner, hello_world and the rest won't. In order to access the edlib API, add #include "edlib.h" in your source file (CMake will automatically update your include path).
For more example projects take a look at applications in apps/.
Edlib is available on VCPKG package manager. With VCPKG on your system, Edlib can be downloaded using the VCPKG install command vcpkg install edlib. Once the library has been downloaded, add the following instructions to your CMakeLists.txt file:
find_package(edlib CONFIG REQUIRED)
target_link_libraries(MyProject PRIVATE edlib::edlib)
then you should be able to include the library header in your project (#include "edlib.h)
Primary way of building Edlib is via Meson build tool.
Requirements: make sure that you have meson installed on your system.
Execute
make
to build static library and binaries (apps and tests) and also run tests.
To build shared library and binaries, do make LIBRARY_TYPE=shared.
Library and binaries will be created in meson-build directory.
You can choose alternate build directory like this: make BUILD_DIR=some-other-dir.
Optionally, you can run
sudo make install
to install edlib library on your machine (on Linux, this will usually install it to usr/local/lib and usr/local/include).
Check Makefile if you want to run individual steps on your own (building, tests, ...).
NOTE: If you need more control, use meson command directly, Makefile is here only to help with common commands.
Edlib can alternatively be built with CMake.
Execute following command to build Edlib using CMAKE:
cd build && cmake -D CMAKE_BUILD_TYPE=Release .. && make
This will create binaries in bin/ directory and libraries (static and shared) in lib/ directory.
./bin/runTests
to run tests.
Optionally, you can run
sudo make install
to install edlib library on your machine.
Edlib can also be installed via Conda: :
conda install edlib.
Main function in edlib is edlibAlign. Given two sequences (and their lengths), it will find edit distance, alignment path or its end and start locations.
char* query = "ACCTCTG";
char* target = "ACTCTGAAA"
EdlibAlignResult result = edlibAlign(query, 7, target, 9, edlibDefaultAlignConfig());
if (result.status == EDLIB_STATUS_OK) {
printf("%d", result.editDistance);
}
edlibFreeAlignResult(result);NOTE: One character is expected to occupy one char/byte, meaning that characters spanning multiple chars/bytes are not supported. As long as your alphabet size is <= 256 you can manually map it to numbers/chars from 0 to 255 and solve this that way, but if its size is > 256 then you will not be able to use Edlib.
edlibAlign takes configuration object (it is a struct EdlibAlignConfig), which allows you to further customize how alignment will be done. You can choose alignment method, tell edlib what to calculate (just edit distance or also path and locations) and set upper limit for edit distance.
For example, if you want to use infix(HW) alignment method, want to find alignment path (and edit distance), are interested in result only if edit distance is not larger than 42 and do not want to extend character equality definition, you would call it like this:
edlibAlign(seq1, seq1Length, seq2, seq2Length,
edlibNewAlignConfig(42, EDLIB_MODE_HW, EDLIB_TASK_PATH, NULL, 0));Or, if you want to use suffix(SHW) alignment method, want to find only edit distance, do not have any limits on edit distance and want character '?' to match both itself and characters 'X' and 'Y', you would call it like this:
EdlibEqualityPair additionalEqualities[2] = {{'?', 'X'}, {'?', 'Y'}};
edlibAlign(seq1, seq1Length, seq2, seq2Length,
edlibNewAlignConfig(-1, EDLIB_MODE_SHW, EDLIB_TASK_DISTANCE, additionalEqualities, 2));We used edlibNewAlignConfig helper function to easily create config, however we could have also just created an instance of it and set its members accordingly.
edlibAlign function returns a result object (EdlibAlignResult), which will contain results of alignment (corresponding to the task that you passed in config).
EdlibAlignResult result = edlibAlign(seq1, seq1Length, seq2, seq2Length,
edlibNewAlignConfig(-1, EDLIB_MODE_HW, EDLIB_TASK_PATH, NULL, 0));
if (result.status == EDLIB_STATUS_OK) {
printf("%d\n", result.editDistance);
printf("%d\n", result.alignmentLength);
printf("%d\n", result.endLocations[0]);
}
edlibFreeAlignResult(result);It is important to remember to free the result object using edlibFreeAlignResult function, since Edlib allocates memory on heap for certain members. If you decide to do the cleaning manually and not use edlibFreeAlignResult, do not forget to manually free() required members.
Cigar is a standard way to represent alignment path. Edlib has helper function that transforms alignment path into cigar.
char* cigar = edlibAlignmentToCigar(result.alignment, result.alignmentLength, EDLIB_CIGAR_STANDARD);
printf("%s", cigar);
free(cigar);For complete documentation of Edlib library API, visit http://martinsos.github.io/edlib (should be updated to the latest release).
To generate the latest API documentation yourself from the source, you need to have doxygen installed.
Position yourself in the root directory and run doxygen, this will generate docs/ directory. Then open docs/html/index.html file with you favorite browser.
Alternatively, you can directly check edlib.h.
Edlib supports 3 alignment methods:
- global (NW) - This is the standard method, when we say "edit distance" this is the method that is assumed. It tells us the smallest number of operations needed to transform first sequence into second sequence. This method is appropriate when you want to find out how similar is first sequence to second sequence.
- prefix (SHW) - Similar to global method, but with a small twist - gap at query end is not penalized. What that means is that deleting elements from the end of second sequence is "free"!
For example, if we had
AACTandAACTGGC, edit distance would be 0, because removingGGCfrom the end of second sequence is "free" and does not count into total edit distance. This method is appropriate when you want to find out how well first sequence fits at the beginning of second sequence. - infix (HW): Similar as prefix method, but with one more twist - gaps at query end and start are not penalized. What that means is that deleting elements from the start and end of second sequence is "free"!
For example, if we had
ACTandCGACTGAC, edit distance would be 0, because removingCGfrom the start andGACfrom the end of second sequence is "free" and does not count into total edit distance. This method is appropriate when you want to find out how well first sequence fits at any part of second sequence. For example, if your second sequence was a long text and your first sequence was a sentence from that text, but slightly scrambled, you could use this method to discover how scrambled it is and where it fits in that text. In bioinformatics, this method is appropriate for aligning read to a sequence.
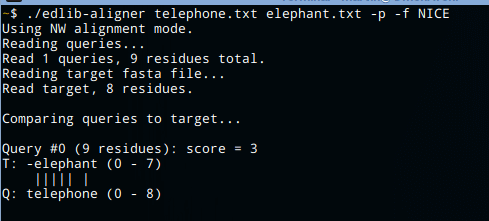
Edlib comes with a standalone aligner cli app, which can be found at apps/aligner/.
Aligner reads sequences from fasta files, and it can display alignment path in graphical manner or as a cigar. It also measures calculation time, so it can be useful for testing speed and comparing Edlib with other tools.
Check Building to see how to build binaries (including edlib-aligner).
Run ./build/bin/edlib-aligner with no params for help and detailed instructions.
Example of usage:
./build/bin/edlib-aligner -p apps/aligner/test_data/query.fasta apps/aligner/test_data/target.fasta
NOTE: Aligner currently does not work on Windows, because it uses getopt to parse command line arguments, which is not supported on Windows.
Check Building to see how to build binaries (including binary runTests).
To run tests, just run ./runTests. This will run random tests for each alignment method, and also some specific unit tests.
Edlib is based on Myers's bit-vector algorithm and extends from it.
It calculates a dynamic programming matrix of dimensions Q x T, where Q is the length of the first sequence (query), and T is the length of the second sequence (target). It uses Ukkonen's banded algorithm to reduce the space of search, and there is also parallelization from Myers's algorithm, however time complexity is still quadratic.
Edlib uses Hirschberg's algorithm to find alignment path, therefore space complexity is linear.
Time complexity: O(T * Q).
Space complexity: O(T + Q).
It is worth noting that Edlib works best for large, similar sequences, since such sequences get the highest speedup from banded approach and bit-vector parallelization.
In test_data/ directory there are different genome sequences, ranging from 10 kbp to 5 Mbp in length. They are ranging in length and similarity, so they can be useful for testing and measuring speed in different scenarios.
Feel free to send pull requests and raise issues.
When developing, you may want to use -D CMAKE_BUILD_TYPE=Debug flag when calling cmake in order to get debugging flags passed to compiler. This should also happen if you just run cmake .. with no flags, but I think I have noticed it does not always works as expected (probably has something to do with cmake cache). To check which flags is compiler using, run make with VERBOSE=1: make VERBOSE=1.
Martin Šošić, Mile Šikić; Edlib: a C/C ++ library for fast, exact sequence alignment using edit distance. Bioinformatics 2017 btw753. doi: 10.1093/bioinformatics/btw753
Mile Šikić (@msikic) - Mentoring and guidance through whole project.
Ivan Sović (@isovic) - Help with testing and prioritizing features, valuable comments on the manuscript.
NW stands for Needleman-Wunsch, HW for Hybrid Wunsch, and SHW for Semi Hybrid Wunsch. While NW is a common abbreviation, HW and SHW abbreviations were made up at the very start of this project to describe additional modes of alignment. Later we started using terms "global", "infix" and "prefix" more, as they describe the modes better, but terms NW, HW and SHW are still very present in the project.