A Retrieval Augmented Generation based chatbot that uses langchain, embeddings and LLMs to answer any query related to the GE Documentation.
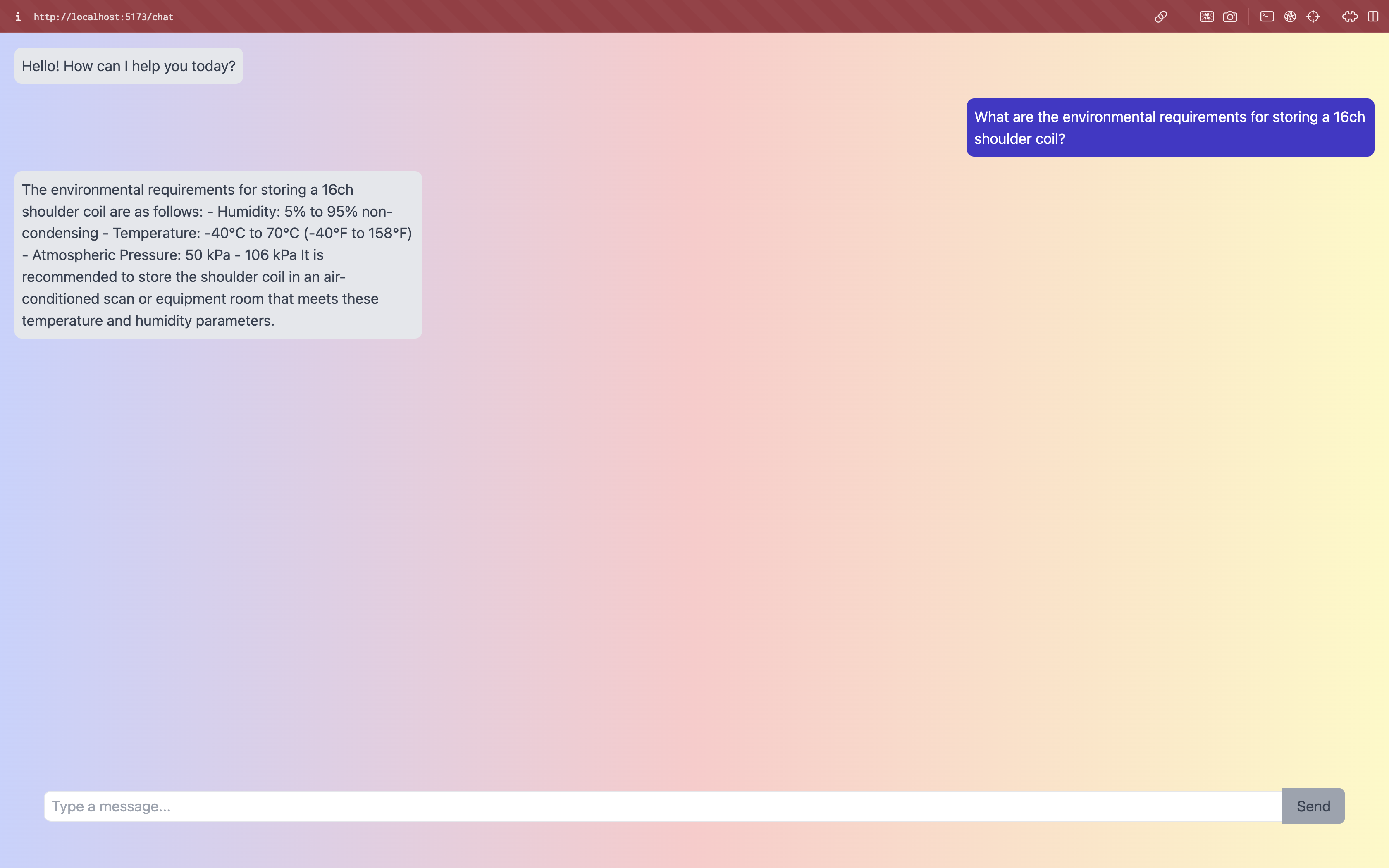

- GE HealthCare (GEHC) manages a vast volume of medical documentation across various languages and modalities. Customers and GE Field engineers face challenges in efficiently accessing and retrieving specific information, potentially causing operational delays and affecting patient care.
-
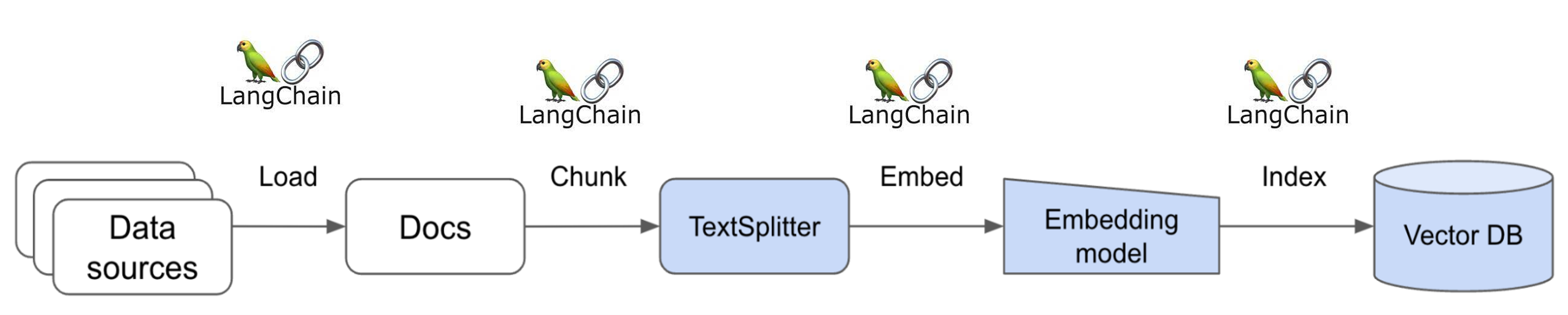
The Data Retrieval Engine interfaces with the documentation portal's database, converts them into embeddings and stores them into a vector database, retrieves relevant information, and provides context.
-
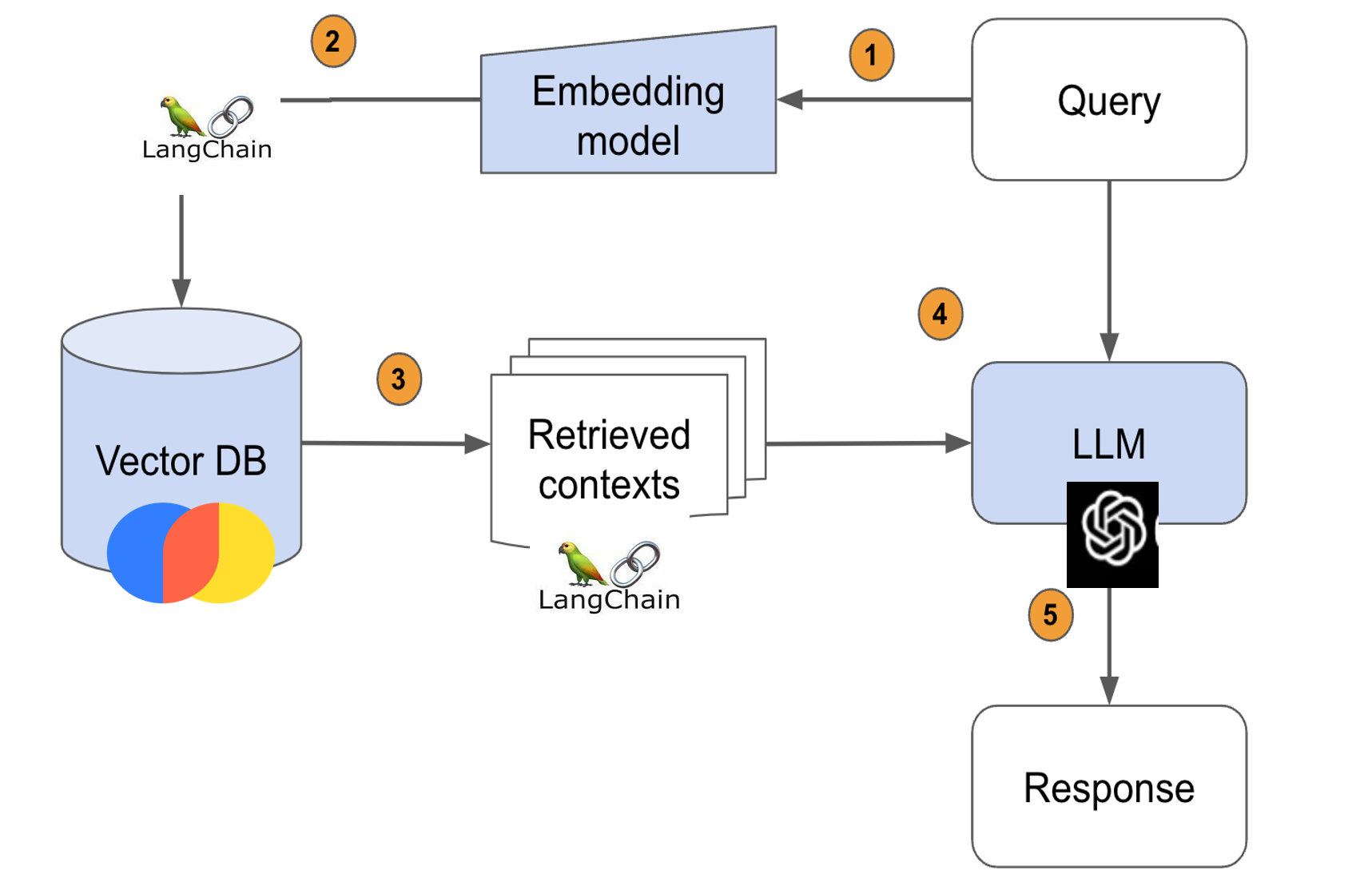
RAG pipeline
- Passes the query to the embedding model to semantically represent it as an embedded query vector.
- Passes the embedded query vector to our vector DB.
- Retrieves the top-k relevant contexts – measured by distance between the query embedding and all the embedded chunks in our knowledge base.
- Passes the query text and retrieved context text to our LLM.
- The LLM will generate a response using the provided content.
- Experimented with Databases like Pinecone, FAISS, Vectara and ChromaDB.
- Experimented with open source large language models like flanT5, Llama2 , gpt2
- Experimented with library of lightweight LLMs LaMini
- Learnt about the SOTA model for Document QA chains , RAG(Retrieval Augmented Generation)
- Learnt about various data formats like parquet which help in data compression for large databases.
- Learnt about flask APIs to manage requests to LLMs.
- Learnt the implementation of open sourced Large language models locally.