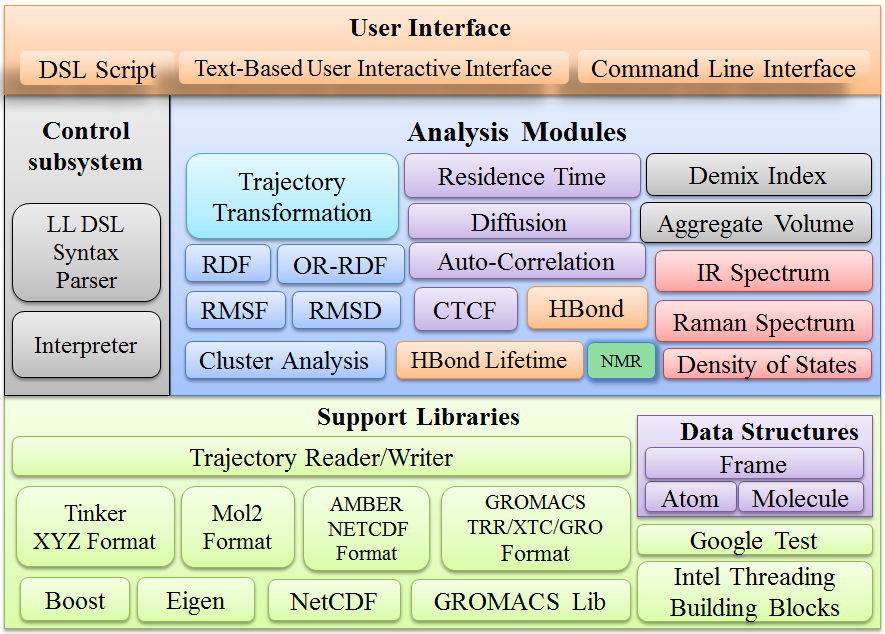
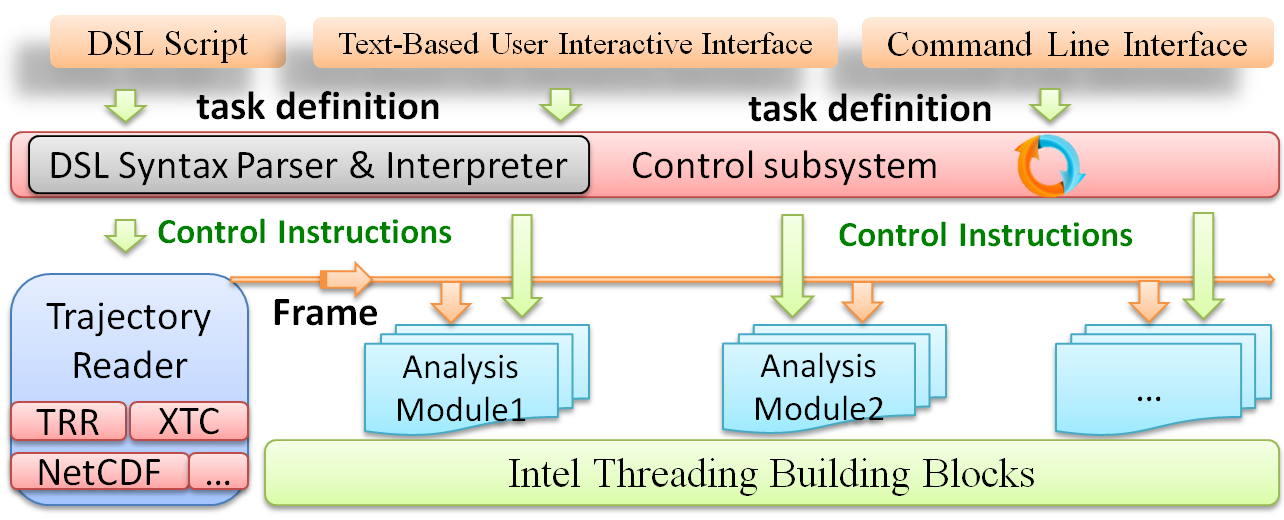
- trajectory format conversion between Tinker ARC, Gromacs Trr, Gromacs Xtc, Gromacs Gro and Amber Netcdf
- Diffusion coefficient based on Einstein and Green-Kubo equation
- Hydrogen bond analysis
- Rotational autocorrelation function
- NMR NOE calculation
- Water residence time
- Cluster analysis based on linkage method
- Aggregation volume calculation for specific molecule
- IR & Raman Spectrum calculation
- Density of State (DOS) Spectrum
- Hydrogen bond lifetime analysis
- Various angle related analysis and can convert probability distribution to Gibbs free energy plot
- Orientation-Resolved Radial Distribution Functions (JCTC 2019, 15, 803−812)
- Conditional Time Correlation Function (JCTC 2019, 15, 803−812)
- Radius of gyration (mass-weighted)
- Biphase Mix Index
- Co-plane Index (CPI)
- RMSD & RMSF with outputting superposed structures
- Use AmberMask for selecting Residues, Atoms in topology structure
- Use Domain Specific Language(DSL) script to drive analysis process
- Use SQL like language to view topology file content
- Other practical utilities for using Tinker
- NBO spin summary
- NAO orbital contribution by driving Multiwfn
- Delocalization Index (DI) by driving Multiwfn
- QTAIM analysis by driving Multiwfn
- ADCH Charge
- Language : C++20 ( GCC 9.x or above )
- Build System : CMake 3.15
- Third-party libraries : Boost 1.72, Intel Threading Building Blocks(TBB) 2020.1 ( for Multi-Core Parallelism ), NetCDF, FFTW3, GROMACS library 5.1.4 ( for reading and writting Gromacs topology and trajectory file ), Google Test 1.10 ( for unit test ), Eigen 3.3 ( C++ template library for linear algebra ), Pugixml 1.10 ( XML format process ), nlohmann/json 3.7.3, GNU readline