This repository provides implementations and experiments for the following papers.
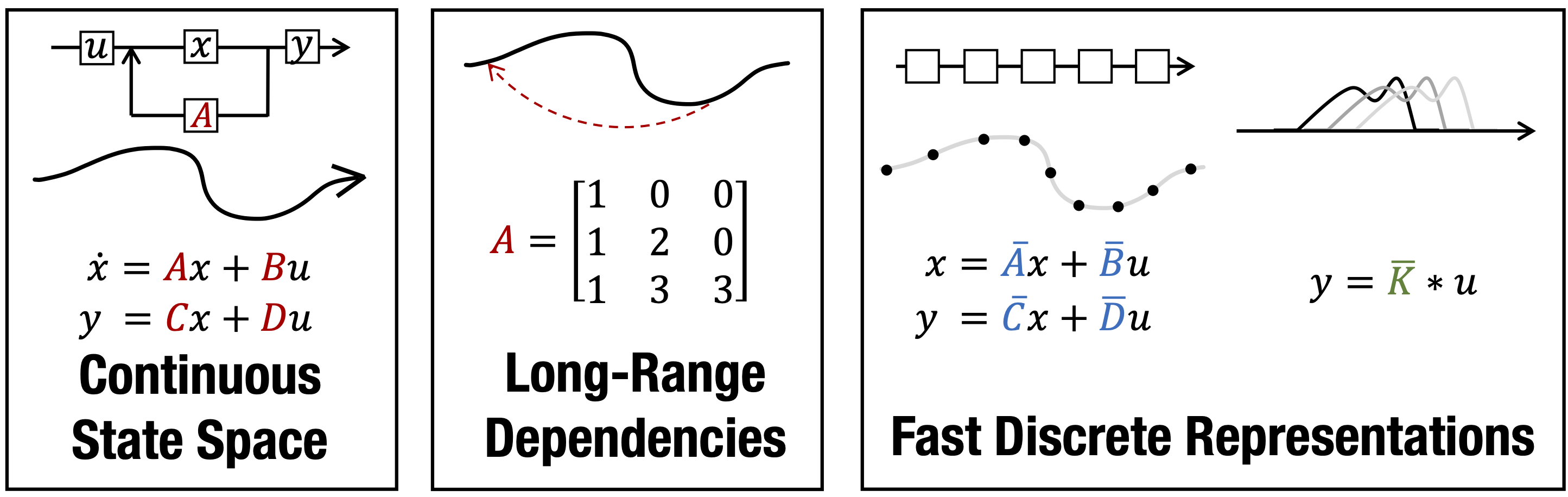
Efficiently Modeling Long Sequences with Structured State Spaces
Albert Gu, Karan Goel, Christopher Ré
Paper: https://arxiv.org/abs/2111.00396
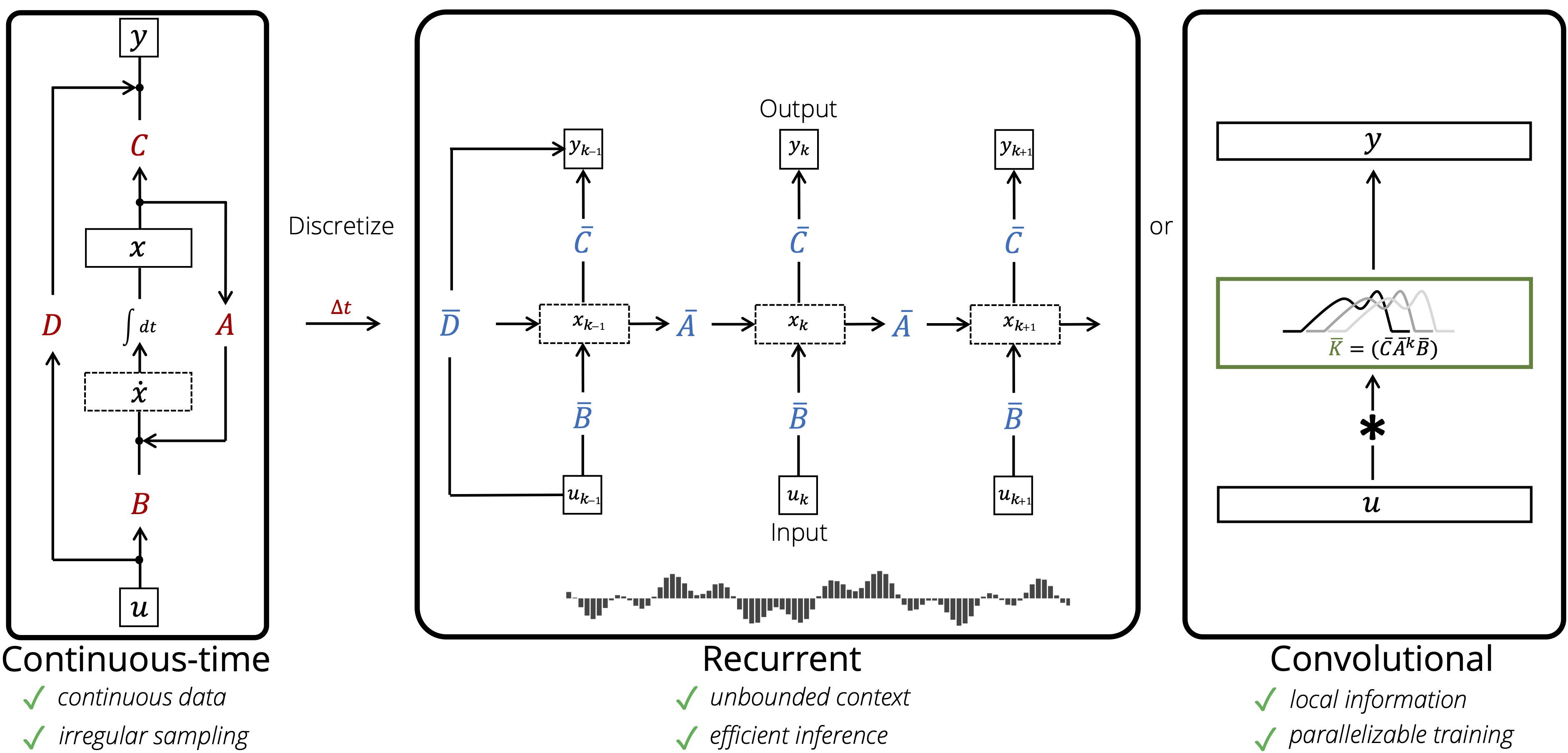
Combining Recurrent, Convolutional, and Continuous-time Models with the Linear State Space Layer
Albert Gu, Isys Johnson, Karan Goel, Khaled Saab, Tri Dao, Atri Rudra, Christopher Ré
Paper: https://arxiv.org/abs/2110.13985
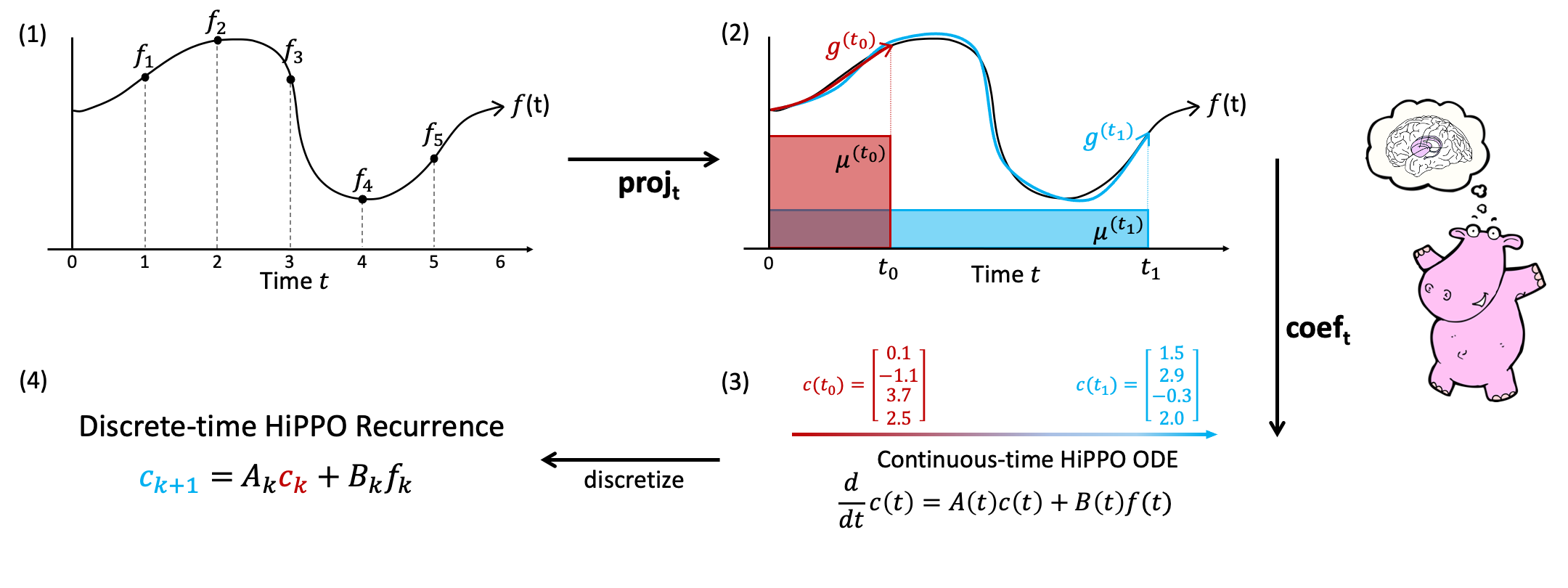
HiPPO: Recurrent Memory with Optimal Polynomial Projections
Albert Gu*, Tri Dao*, Stefano Ermon, Atri Rudra, Christopher Ré
Paper: https://arxiv.org/abs/2008.07669
This repository requires Python 3.8+ and Pytorch 1.9+.
Other packages are listed in requirements.txt.
All logic for creating and loading datasets is in src/dataloaders.
This folder may include old and experimental datasets.
The datasets that we consider core are located in src/dataloaders/datasets.py.
The raw data should be organized as follows.
The data path can be configured by the environment variable DATA_PATH, or defaults to ./data by default, where . is the top level directory of this repository (e.g. 'state-spaces').
Most of the dataloaders download their datasets automatically if not found.
External datasets include Long Range Arena (LRA), which can be downloaded from their GitHub page,
and the WikiText-103 language modeling dataset, which can be downloaded by the getdata.sh script from the Transformer-XL codebase.
These external datasets should be organized as follows:
DATA_PATH/
pathfinder/
pathfinder32/
pathfinder64/
pathfinder128/
pathfinder256/
aan/
listops/
wt103/
Fine-grained control over the data directory is allowed, e.g. if the LRA ListOps files are located in /home/lra/listops-1000/, you can pass in +dataset.data_dir=/home/lra/listops-1000 on the command line
A core operation of S4 is the "Cauchy kernel" described in the paper. The implementation of this requires one of two methods:
This version is faster but requires manual compilation on each machine.
Run python setup.py install from the directory extensions/cauchy/.
This version is provided by the pykeops library.
Installation usually works out of the box with pip install pykeops cmake which are provided in the requirements file.
Note that running in a Colab requires installing a different pip package; instructions can be found in the pykeops documentation.
This section describes how to use the latest S4 model and reproduce experiments immediately. More detailed descriptions of the infrastructure are in the subsequent sections.
The S4 module is found at
src/models/sequence/ss/s4.py.
For users who would like to import a single file that has the self-contained S4 layer,
a standalone version can be found at src/models/sequence/ss/standalone/s4.py.
For testing, we frequently use synthetic datasets or the Permuted MNIST dataset.
This can be run with python -m train wandb=null pipeline=mnist model=s4, which should get to around 90% after 1 epoch which takes 1-3 minutes depending on GPU.
python -m train wandb=null experiment=s4-lra-listops
python -m train wandb=null experiment=s4-lra-imdb
python -m train wandb=null experiment=s4-lra-cifar
python -m train wandb=null experiment=s4-lra-aan
python -m train wandb=null experiment=s4-lra-pathfinder
python -m train wandb=null experiment=s4-lra-pathx
Note that these experiments may take different amounts of time to train. IMDB should take 1-2 hours, while Path-X will take several epochs to take off and take over a day to train to completion.
python -m train wandb=null experiment=s4-cifar
The above command line reproduces our best sequential CIFAR model. Decreasing the model size should yield close results, e.g. decreasing the hidden dimension and number of layers with model.d_model=512 model.n_layers=4.
The Speech Commands dataset that our baselines use is a modified smaller 10-way classification task.
python -m train wandb=null experiment=s4-sc
To use the original version with the full 35 classes, pass in dataset.all_classes=true
python -m train wandb=null experiment=s4-wt103
The default settings require 8 GPUs with 32GB memory. Modifications can be made by decreasing batch size and accumulating gradients, e.g. loader.batch_size=4 trainer.accumulate_grad_batches=2
One notable difference in this codebase is that some S4 parameters use different optimizer hyperparameters. In particular, the SSM kernel is particularly sensitive to the A, B, and dt parameters, so the optimizer settings for these parameters are usually fixed to learning rate 0.001 and weight decay 0.
Our logic for setting these parameters can be found in the OptimModule class under src/models/sequence/ss/kernel.py and the corresponding optimizer hook in SequenceLightningModule.configure_optimizers under train.py.
The core training infrastructure of this repository is based on Pytorch-Lightning with a configuration scheme based on Hydra. The structure of this integration largely follows the Lightning+Hydra integration template described in https://github.com/ashleve/lightning-hydra-template.
The main experiment entrypoint is train.py and configs are found in configs/.
In brief, the main config is found at configs/config.yaml, which is combined with other sets of configs that can be passed on the command line, to define an overall YAML config.
Most config groups define one single Python object (e.g. a PyTorch nn.Module).
The end-to-end training pipeline can broken down into the following rough groups, where group XX is found under configs/XX/:
model: the sequence-to-sequence model backbone (e.g. a src.models.sequence.SequenceModel)
dataset: the raw dataset (data/target pairs) (e.g. a pytorch Dataset)
loader: how the data is loaded (e.g. a pytorch DataLoader)
encoder: defines a Module that interfaces between data and model backbone
decoder: defines a Module that interfaces between model backbone and targets
task: specifies loss and metrics
Default combinations of dataset+loader+encoder+decoder+task are further consolidated into groups called pipelines.
A run can be performed by passing in a pipeline config, model config, and any additional arguments modifying the default configurations. A simple example experiment is
python -m train pipeline=mnist dataset.permute=True model=s4 model.n_layers=3 model.d_model=128 model.norm=batch model.prenorm=True wandb=null
This uses the permuted sequential MNIST task and uses an s4 model with a specified number of layers, backbone dimension, and normalization type.
It is recommended to read the Hydra documentation to fully understand the configuration framework. For help launching specific experiments, please file an Issue.
This codebase uses a modification of the hydra instantiate utility that provides shorthand names of different classes, for convenience in configuration and logging.
The mapping from shorthand to full path can be found in src/utils/registry.py.
Logging with WandB is built into this repository.
In order to use this, simply set your WANDB_API_KEY environment variable, and change the wandb.project attribute of configs/config.yaml (or pass it on the command line python -m train .... wandb.project=s4).
Set wandb=null to turn off WandB logging.
This repository provides a modular and flexible implementation of sequence models at large.
SequenceModule src/models/sequence/base.py is the abstract interface that all sequence models adhere to.
In this codebase, sequence models are defined as a sequence-to-sequence map of shape (batch size, sequence length, input dimension) to (batch size, sequence length, output dimension).
The SequenceModule comes with other methods such as step which is meant for autoregressive settings, and logic to carry optional hidden states (for stateful models such as RNNs or S4).
SequenceModel src/models/sequence/model.py is the main backbone with configurable options for residual function, normalization placement and type, etc.
SequenceModel accepts a black box config for a layer. Compatible layers are SequenceModules (i.e. composable sequence transformations) found under src/models/sequence/.
This is the main model of this repository. See instructions in Getting Started.
The LSSL is the predecessor of S4. It is currently not recommended for use, but the model can be found at src/models/sequence/ss/lssl.py.
It can be run with model/layer=lssl or model/layer=lssl model.layer.learn=0 for the LSSL-fixed model which does not train A, B, or dt.
HiPPO is the mathematical framework upon which the papers HiPPO, LSSL, and S4 are built on.
The logic for HiPPO operators is found under src/models/hippo/.
HiPPO-RNN cells from the original paper can be found under the RNN cells
This codebase contains a flexible and modular implementation of many RNN cells.
Some examples include model=rnn/hippo-legs and model=rnn/hippo-legt for HiPPO variants from the original paper, or model=rnn/gru for a GRU reimplementation, etc.
An exception is model=lstm to use the PyTorch LSTM.
Example command (reproducing the Permuted MNIST number from the HiPPO paper, which was SotA at the time):
python train.py pipeline=mnist model=rnn/hippo-legs model.cell_args.hidden_size=512 train.epochs=50 train.batch_size=100 train.lr=0.001
Other sequence models are easily incorporated into this repository, and several other baselines have been ported.
These include CNNs such as the WaveGAN Discriminator and CKConv and continuous-time/RNN models such as UnICORNN and LipschitzRNN.
python -m train dataset=mnist model={ckconv,unicornn}
configs/ config files for model, data pipeline, training loop, etc.
data/ default location of raw data
extensions/ CUDA extension for Cauchy kernel
src/ main source code for models, datasets, etc.
callbacks/ training loop utilities (e.g. checkpointing)
dataloaders/ data loading logic
models/ model backbones
baselines/ misc. baseline models
functional/ mathematical utilities
nn/ standalone modules and components
hippo/ core HiPPO logic
sequence/ sequence model backbones and layers including RNNs and S4/LSSL
tasks/ encoder/decoder modules to interface between data and model backbone
utils/
train.py training loop entrypoint
If you use this codebase, or otherwise found our work valuable, please cite:
@article{gu2021efficiently,
title={Efficiently Modeling Long Sequences with Structured State Spaces},
author={Gu, Albert and Goel, Karan and R{\'e}, Christopher},
journal={arXiv preprint arXiv:2111.00396},
year={2021}
}
@article{gu2021combining,
title={Combining Recurrent, Convolutional, and Continuous-time Models with Linear State-Space Layers},
author={Gu, Albert and Johnson, Isys and Goel, Karan and Saab, Khaled and Dao, Tri and Rudra, Atri and R{\'e}, Christopher},
journal={Advances in neural information processing systems},
volume={34},
year={2021}
}
@article{gu2020hippo,
title={HiPPO: Recurrent Memory with Optimal Polynomial Projections},
author={Gu, Albert and Dao, Tri and Ermon, Stefano and Rudra, Atri and Re, Christopher},
journal={Advances in neural information processing systems},
volume={33},
year={2020}
}