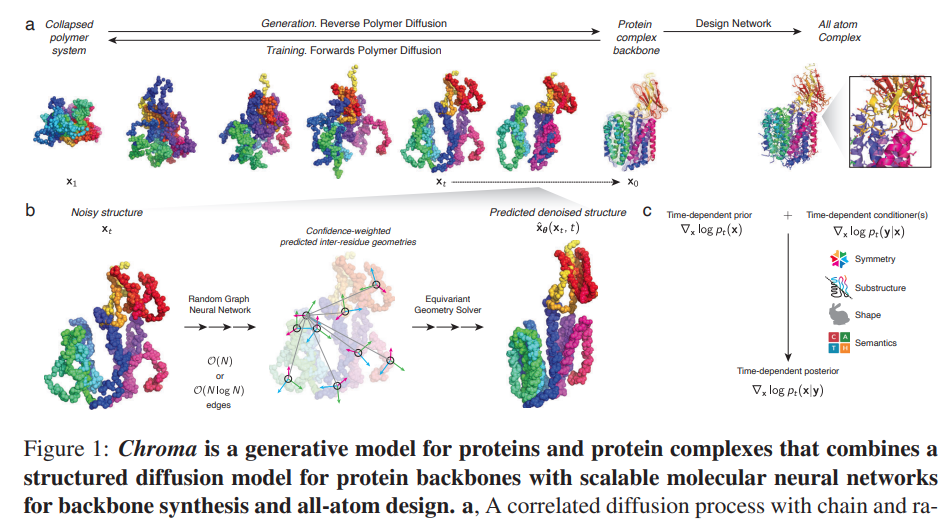
figure 1 in paper
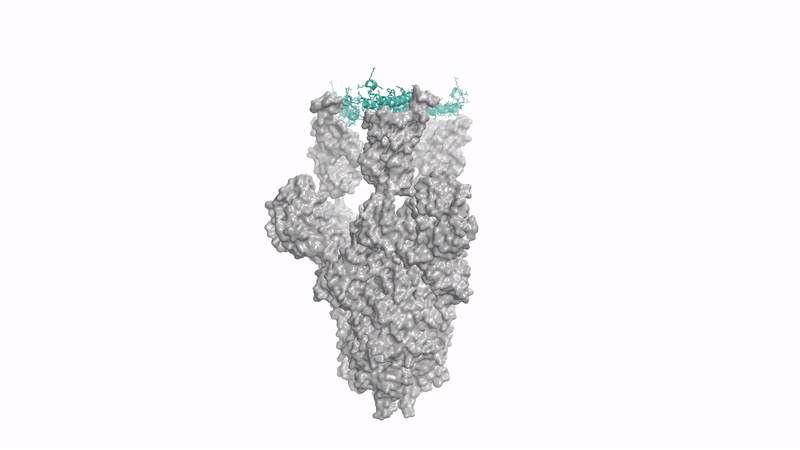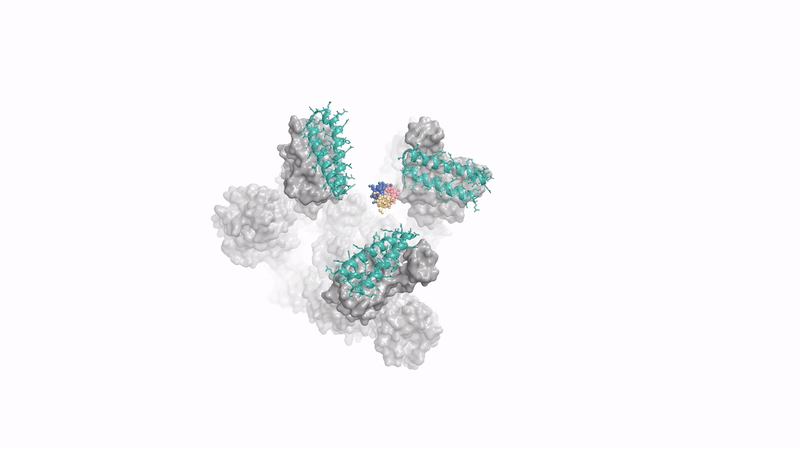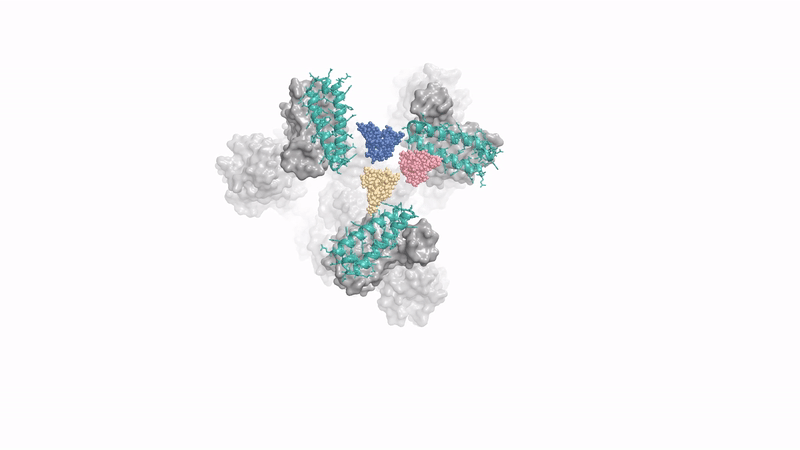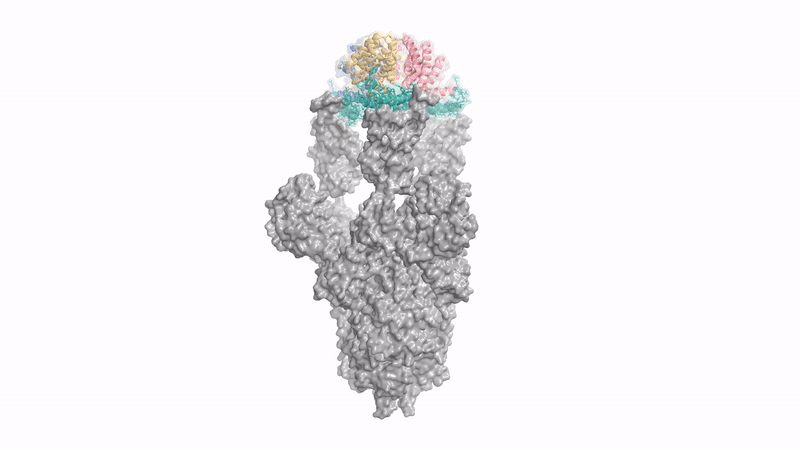
generating a protein that binds to spike protein of coronavirus - Baker lab's concurrent RFDiffusion work
Implementation of Chroma, generative model of proteins using DDPM and GNNs, in Pytorch. Concurrent work seems to suggest we have a slight lift-off applying denoising diffusion probabilistic models to protein design. Will also incorporate self-conditioning, applied successfully by Baker lab in RFDiffusion.
If you are interested in open sourcing works like these out in the wild, please consider joining OpenBioML
- use galactica
@misc{
title = {Illuminating protein space with a programmable generative model},
author = {John Ingraham, Max Baranov, Zak Costello, Vincent Frappier, Ahmed Ismail, Shan Tie, Wujie Wang, Vincent Xue, Fritz Obermeyer, Andrew Beam, Gevorg Grigoryan},
year = {2022},
url = {https://cdn.generatebiomedicines.com/assets/ingraham2022.pdf}
}