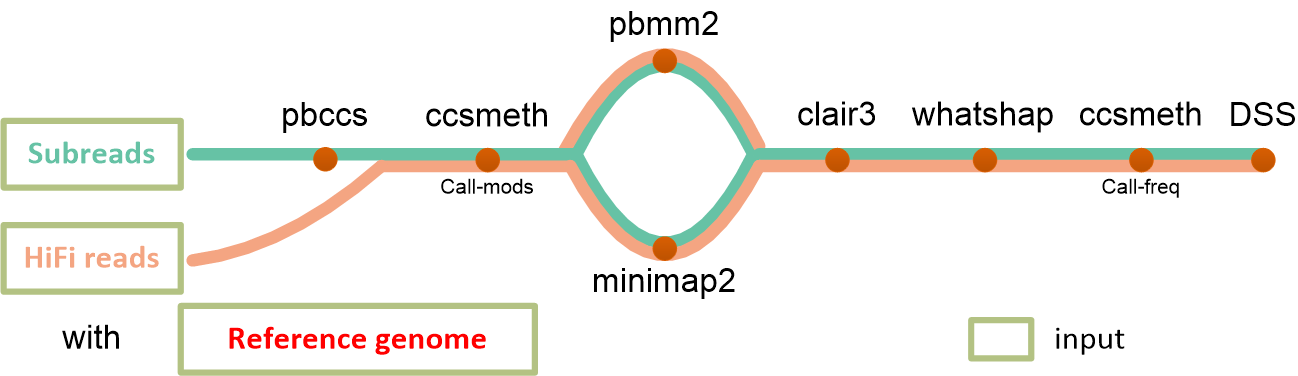
Methylation phasing using PacBio CCS reads
Recommended Hardware requirements: 128 GB RAM, 40 CPU processors, 4 TB disk storage, >=8 GB GPU
Recommended OS: Linux (Ubuntu 16.04, CentOS 7, etc.)
# create a new environment and install nextflow in it
conda create -n nextflow -c conda-forge -c bioconda nextflow
# OR, install nextflow in an existing environment
conda install -c conda-forge -c bioconda nextflow- (3) Download ccsmethphase from github.
git clone https://github.com/PengNi/ccsmethphase.git- (4) Install Docker or Singularity if needed.
# e.g., install singularity using conda
conda install -c conda-forge singularity - (5) [optional] Install graphviz.
conda install -c conda-forge graphvizCheck ccsmethphase/demo for demo data:
- hg002.chr20_demo.hifi.bam: HG002 demo hifi reads which are aligned to human genome chr20:10000000-10100000.
- chr20_demo.fa: reference sequence of human chr20:10000000-10100000.
- hg002_bsseq_chr20_demo.bed: HG002 BS-seq results of region chr20:10000000-10100000.
- input_sheet.tsv: Information of the demo PacBio bam file hg002.chr20_demo.hifi.bam.
ccsmethphase takes files of PacBio reads (subreads.bam or hifi.bam), and a reference genome as input.
The information of PacBio reads files should be organized into a tsv file, like input_sheet.tsv in demo data:
| Group_ID | Sample_ID | Type | Path |
|---|---|---|---|
| G1 | HG002_demo | hifi | ./demo/hg002.chr20_demo.hifi.bam |
| G1 | HG002_demo | hifi | path of another flowcell bam file for HG002_demo |
| G1 | HG003 | hifi | path of a bam file for HG003 |
- Group_ID: For group comparation, values can be like control, case, or anything else.
- Sample_ID: The name of the sample sequenced.
- Type: Data type, should be hifi or subreads.
- Path: Path of a (flowcell) bam file sequenced using the sample. Absolute path recommended.
NOTE: ccsmethphase can be run with conda, docker, and singularity by setting -profile. If you are using -profile conda to run this workflow, ccsmeth models should be set as input too. Check ccsmeth to get ccsmeth 5mCpG models.
--dsname: job name
--input: the tsv file containing information of PacBio reads files
--genome: file path of the reference genome
--include_all_ctgs: "true" or "false". default false, means only [chr][1-22,X,Y] included.
-profile: conda/docker/singularity, test
If it is the first time you run with singularity (e.g. using -profile singularity), the following cmd will cache the dafault singularity image (--singularity_name and/or --clair3_singularity_name) to the --singularity_cache directory (default: local_singularity_cache) first. There will be .img file(s) in the --singularity_cache directory.
NOTE: If you are using relative paths of bam files in input_sheet.tsv, make sure the relative paths are the right relative paths to the directory you launch the workflow.
For the example data:
# activate nextflow environment
conda activate nextflow
cd /path/to/ccsmethphase
nextflow run main.nf \
--dsname test \
-profile singularity,testThe above command is equal to the command following:
nextflow run /path/to/ccsmethphase \
--dsname test \
--genome /path/to/ccsmethphase/demo/chr20_demo.fa \
--input /path/to/ccsmethphase/demo/input_sheet.tsv \
--include_all_ctgs true \
--max_cpus 8 \
--max_memory "12.GB" \
--max_time "6.h" \
-profile singularityTry the following command to enable GPU:
CUDA_VISIBLE_DEVICES=0 nextflow run /path/to/ccsmethphase \
--dsname test \
--genome /path/to/ccsmethphase/demo/chr20_demo.fa \
--input /path/to/ccsmethphase/demo/input_sheet.tsv \
--include_all_ctgs true \
-profile singularityThe downloaded singularity .img file(s) can be re-used, without being downloaded again:
nextflow run /path/to/ccsmethphase \
--dsname test2 \
--genome /path/to/some/other/genome/fa \
--input /path/to/some/other/input_sheet.tsv \
-profile singularity \
--singularity_cache /path/to/local_singularity_cacheTry -resume to re-run a modified/failed job to save time:
nextflow run /path/to/ccsmethphase \
--dsname test \
--genome /path/to/ccsmethphase/demo/chr20_demo.fa \
--input /path/to/ccsmethphase/demo/input_sheet.tsv \
--include_all_ctgs true \
-profile singularity \
-resumePossible issues:
no space left on deviceERROR
This may be caused by limited space of the SINGULARITY_TMPDIR (default /tmp) dir. Try setting SINGULARITY_TMPDIR to a disk that have enough space in current environment:
export SINGULARITY_TMPDIR=/path/to/another/dirThe output directory should look like the following:
ccsmethphase_results/
├── pipeline_info
└── test
├── bam
│ ├── G1.HG002_demo.hifi.ccsmeth.modbam.pbmm2.merged_size1.SNV_PASS_whatshap.bam
│ └── G1.HG002_demo.hifi.ccsmeth.modbam.pbmm2.merged_size1.SNV_PASS_whatshap.bam.bai
├── diff_methyl
│ ├── G1.HG002_demo.hifi.ccsmeth.modbam.pbmm2.merged_size1.SNV_PASS_whatshap.freq.aggregate.hp_callDML.txt
│ ├── G1.HG002_demo.hifi.ccsmeth.modbam.pbmm2.merged_size1.SNV_PASS_whatshap.freq.aggregate.hp_callDMR.autosomes_cf0.2.bed
│ └── G1.HG002_demo.hifi.ccsmeth.modbam.pbmm2.merged_size1.SNV_PASS_whatshap.freq.aggregate.hp_callDMR.txt
├── mods_freq
│ ├── G1.HG002_demo.hifi.ccsmeth.modbam.pbmm2.merged_size1.SNV_PASS_whatshap.freq.aggregate.all.bed
│ ├── G1.HG002_demo.hifi.ccsmeth.modbam.pbmm2.merged_size1.SNV_PASS_whatshap.freq.aggregate.hp1.bed
│ └── G1.HG002_demo.hifi.ccsmeth.modbam.pbmm2.merged_size1.SNV_PASS_whatshap.freq.aggregate.hp2.bed
└── vcf
├── clair3_called
│ ├── G1.HG002_demo.hifi.ccsmeth.modbam.pbmm2.merged_size1.clair3_merge.vcf.gz
│ └── G1.HG002_demo.hifi.ccsmeth.modbam.pbmm2.merged_size1.clair3_merge.vcf.gz.tbi
└── whatshap_phased
├── G1.HG002_demo.hifi.ccsmeth.modbam.pbmm2.merged_size1.clair3_merge.SNV_PASS_whatshap.vcf.gz
└── G1.HG002_demo.hifi.ccsmeth.modbam.pbmm2.merged_size1.clair3_merge.SNV_PASS_whatshap.vcf.gz.tbi
- pipeline_info: Information of the workflow execution
- test: directory to save the results of ccsmethphase, name set by
--dsname- bam: modbam files
- diff_methyl: DMLs and DMRs generated by CCS
- mods_freq: bedmethyl files, site-level methylation frequencies
- vcf: VCF files for SNVs
- clair3_called: SNVs generated by clair3
- whatshap_phsed: Phased SNVs by whatshap
- Some code were referenced from nanome and nf-core.
- Code for ASM detection using DSS were referenced from NanoMethPhase of Akbari et al.
input format -> group_id type(hifi/subreads) file_abs_pathhave tested docker on cpu, singularity on cpu/gpu/cpu-in-gpu-machine;did not test docker on gpu/cpu-in-gpu-machine yet- complete --help/-h option
default model in docker container- add quality-control/statistics process for ccs data?
add DSS-2.44.0, update docker/env.yml/readme/image- report-summary (Rmarkdown->html?)
addtestprofile?