Tips and tricks in R & RStudio, gathered from wherever I see them.
Mainly a repo for me to remember cool little R tips I’ve seen around the
place 🤘. I hope to update it regularly – feel free to fork, add
your own, and send a PR. I’ve left the library() calls in every chunk,
in case you want to copy and paste some code. Long code chunks are
hidden for readability. Just click on the little arrow and you can see
the code. This is clearly a work-in-progress-that-might-never-finish,
so any corrections/tips/pull requests/additions are very welcome!
Adding compress = "xz" to your save() function can make things much
smaller. Very useful tip from Ilya
Kasnitsky:
library(congressbr)
data('senate_nominal_votes')
head(senate_nominal_votes)
save(list = ls(), file = "sen.Rda")
save(list = ls(), file = "sen2.Rda", compress = "xz")
file.info("sen.Rda")$size
#> [1] 73485
file.info("sen2.Rda")$size
#> [1] 38416It’s a common task that you have a folder full of files, let’s say
.csv files, and you want to read them all into R and put them in a
single data frame. Here’s a purrr version (you might want to use
data.table’s fread() and rbind.list() if you have lots of files):
library(readr)
library(dplyr)
library(purrr)
library(stringr)
df <- dir() %>%
.[str_detect(., '.csv')] %>%
map_df(read_csv)That’s just beautiful code, isn’t it?
library() or require() only load one package at a time, but…
packages <- c("dplyr", "ggplot2", "rstan", "readr")
lapply(packages, library, character.only = TRUE)This code loads as many packages as you put in packages. They need to
be installed first, of course!
You can of course do something similar with installing packages, for
example lapply(pkgs, install.packages), where pkgs is a character
vector of package names. Or with xfun::pkg_attach().
There are a few ways to do this, but this works. You just need to put in your version of R:
Click to see code
version <- "3.3" #just an example
old.packages <- list.files(paste0("/Library/Frameworks/R.framework/Versions/", version, "/Resources/library"))
# Install packages in the previous version.
# For each package p in previous version...
for (p in old.packages) {
# ... Only if p is not already installed
if (!(p %in% installed.packages()[,"Package"])) {
# Install p
install.packages(p)
}
}I’m not sure this is necessary anymore, the last time I updated R, all my packages were still installed. Anyway.
Nice totals column, from Andrew Heiss:
Click to see code
library(dplyr)
library(stringr)
library(pander)
mtcars %>%
mutate(cars = row.names(.),
cars = str_extract(cars, "[A-Za-z\\b]*")) %>%
count(cars, am) %>%
bind_rows(summarise_at(., vars(n), funs(sum)) %>%
mutate(cars = "**Total**")
) %>%
pandoc.table()
#>
#> ---------------------
#> cars am n
#> ----------- ---- ----
#> AMC 0 1
#>
#> Cadillac 0 1
#>
#> Camaro 0 1
#>
#> Chrysler 0 1
#>
#> Datsun 1 1
#>
#> Dodge 0 1
#>
#> Duster 0 1
#>
#> Ferrari 1 1
#>
#> Fiat 1 2
#>
#> Ford 1 1
#>
#> Honda 1 1
#>
#> Hornet 0 2
#>
#> Lincoln 0 1
#>
#> Lotus 1 1
#>
#> Maserati 1 1
#>
#> Mazda 1 2
#>
#> Merc 0 7
#>
#> Pontiac 0 1
#>
#> Porsche 1 1
#>
#> Toyota 0 1
#>
#> Toyota 1 1
#>
#> Valiant 0 1
#>
#> Volvo 1 1
#>
#> **Total** NA 32
#> ---------------------Also, from the comments to the above tweet, by Sam Firke, the author the janitor package (I prefer this actually):
library(dplyr)
library(janitor)
mtcars %>%
mutate(cars = row.names(.),
cars = str_extract(cars, "[A-Za-z\\b]*")) %>%
count(cars, am) %>%
adorn_totals()
#> cars am n
#> AMC 0 1
#> Cadillac 0 1
#> Camaro 0 1
#> Chrysler 0 1
#> Datsun 1 1
#> Dodge 0 1
#> Duster 0 1
#> Ferrari 1 1
#> Fiat 1 2
#> Ford 1 1
#> Honda 1 1
#> Hornet 0 2
#> Lincoln 0 1
#> Lotus 1 1
#> Maserati 1 1
#> Mazda 1 2
#> Merc 0 7
#> Pontiac 0 1
#> Porsche 1 1
#> Toyota 0 1
#> Toyota 1 1
#> Valiant 0 1
#> Volvo 1 1
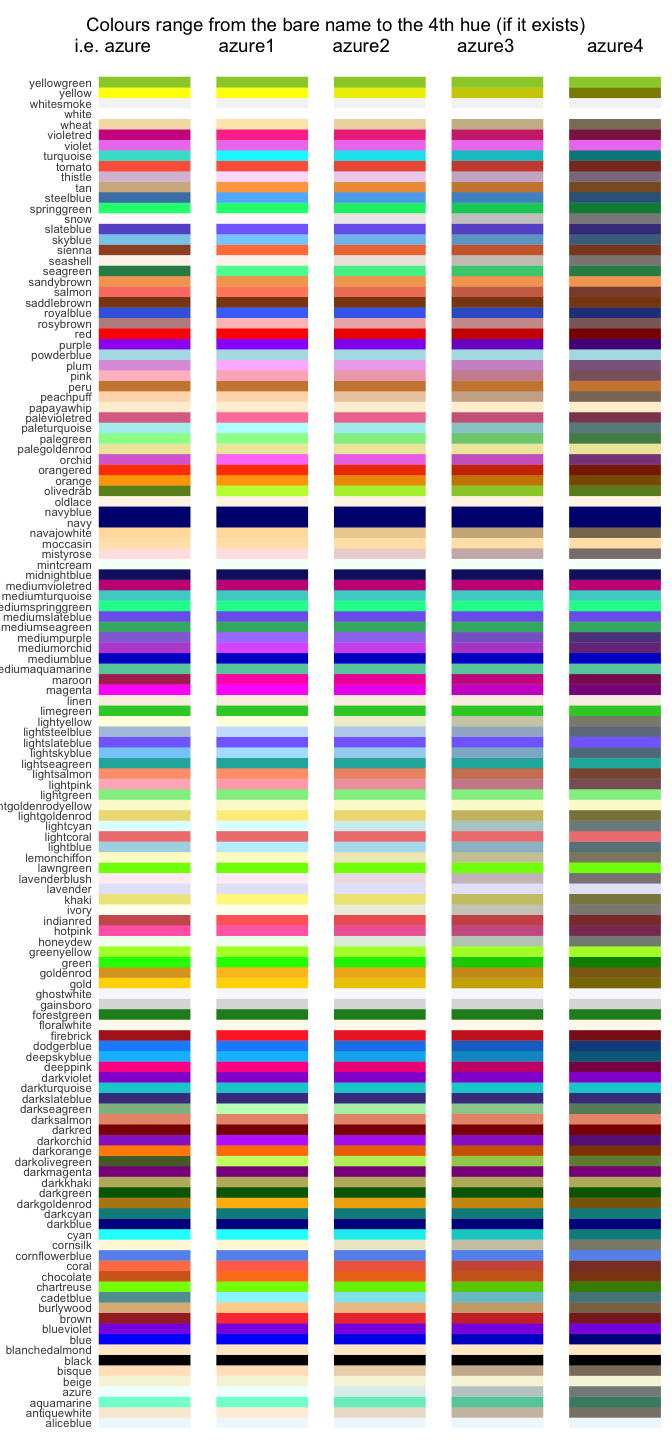
#> Total 11 32Want to see all the colours available in R? Here’s a ggplot2 version of this great gist:
Click to see code
# R colors minus 100 shades of grey
library(dplyr)
library(stringr)
library(ggplot2)
library(tibble)
library(cowplot)
# get 'data':
colour <- tibble(colours = colors()) %>%
filter(!grepl("gray", colours),
!grepl("grey", colours)) %>%
mutate(general_colour = gsub("[0-9]", "", colours),
c1 = ifelse(grepl("1", colours), 1, 0),
c2 = ifelse(grepl("2", colours), 1, 0),
c3 = ifelse(grepl("3", colours), 1, 0),
c4 = ifelse(grepl("4", colours), 1, 0)) %>%
select(-1) %>%
group_by(general_colour) %>%
summarise_all(funs(sum)) %>%
ungroup() %>%
mutate(c1 = ifelse(grepl(1, c1), paste0(general_colour, c1), NA),
c2 = ifelse(grepl(1, c2), paste0(general_colour, "2"), NA),
c3 = ifelse(grepl(1, c3), paste0(general_colour, "3"), NA),
c4 = ifelse(grepl(1, c4), paste0(general_colour, "4"), NA),
c1 = ifelse(is.na(c1), general_colour, c1),
c2 = ifelse(is.na(c2), general_colour, c2),
c3 = ifelse(is.na(c3), general_colour, c3),
c4 = ifelse(is.na(c4), general_colour, c4))
## create six plots:
# Just the names, by setting alpha to 0:
g0 <- ggplot(colour, aes(x = general_colour)) +
geom_bar(position = "stack", alpha = 0) +
coord_flip() +
theme_minimal() +
theme(legend.position = "none") +
theme(axis.text.x = element_blank(), axis.ticks.x = element_blank(),
axis.title.x = element_blank(), panel.grid = element_blank(),
axis.title.y = element_blank())
g <- ggplot(colour, aes(x = general_colour, color = general_colour,
fill = general_colour)) +
geom_bar(position = "stack") +
coord_flip() +
scale_color_manual(values = colour$general_colour) +
scale_fill_manual(values = colour$general_colour) +
theme_minimal() +
theme(legend.position = "none") +
theme(axis.text = element_blank(), axis.ticks = element_blank(),
axis.title = element_blank(), panel.grid = element_blank())
g_1 <- ggplot(colour, aes(x = c1, color = c1,
fill = c1)) +
geom_bar(position = "stack") +
coord_flip() +
scale_color_manual(values = colour$c1) +
scale_fill_manual(values = colour$c1) +
theme_minimal() +
theme(legend.position = "none") +
theme(axis.text = element_blank(), axis.ticks = element_blank(),
axis.title = element_blank(), panel.grid = element_blank())
g_2 <- ggplot(colour, aes(x = c2, color = c2,
fill = c2)) +
geom_bar(position = "stack") +
coord_flip() +
scale_color_manual(values = colour$c2) +
scale_fill_manual(values = colour$c2) +
theme_minimal() +
theme(legend.position = "none") +
theme(axis.text = element_blank(), axis.ticks = element_blank(),
axis.title = element_blank(), panel.grid = element_blank())
g_3 <- ggplot(colour, aes(x = c3, color = c3,
fill = c3)) +
geom_bar(position = "stack") +
coord_flip() +
scale_color_manual(values = colour$c3) +
scale_fill_manual(values = colour$c3) +
theme_minimal() +
theme(legend.position = "none") +
theme(axis.text = element_blank(), axis.ticks = element_blank(),
axis.title = element_blank(), panel.grid = element_blank())
g_4 <- ggplot(colour, aes(x = c4, color = c4,
fill = c4)) +
geom_bar(position = "stack") +
coord_flip() +
scale_color_manual(values = colour$c4) +
scale_fill_manual(values = colour$c4) +
theme_minimal() +
theme(legend.position = "none") +
theme(axis.text = element_blank(), axis.ticks = element_blank(),
axis.title = element_blank(), panel.grid = element_blank())
# cowplot 'em all together:
p <- plot_grid(g0, g, g_1, g_2, g_3, g_4, align = "h", ncol = 6,
rel_widths = c(.75, 1.05, 1.05, 1.05, 1.05, 1.05))
title <- ggdraw() + draw_label("Colours range from the bare name to the 4th hue (if it exists)\n i.e. azure azure1 azure2 azure3 azure4 ")
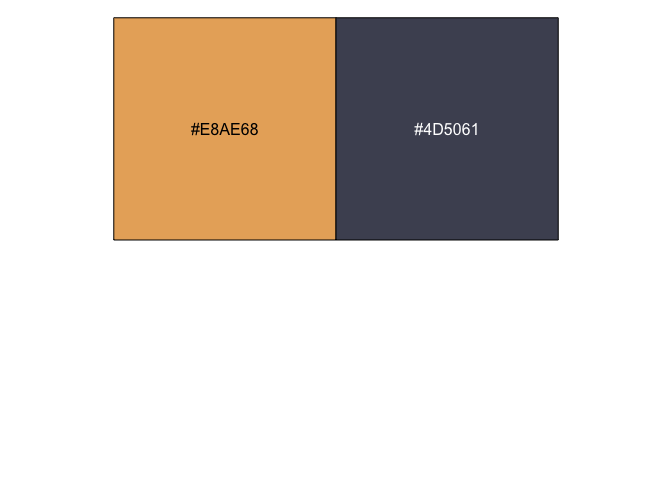
plot_grid(title, p, ncol = 1, rel_heights=c(0.05, 1))I was trying to get a nice colour for a geom_vline() recently, and I
was looking for a way to quickly print a colour in R. The scales
package to the rescue!
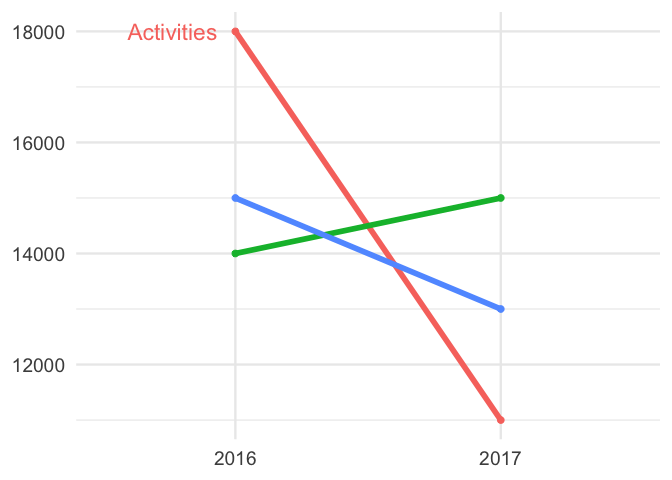
scales::show_col(c("#E8AE68", "#4D5061"))You can use curly braces ({}) to avail of data wrangling in the middle
of your ggplot2 code, as
Alistair once
explained to me. I can’t find the code I was working on at the time, but
the idea was to make some slopegraphs. This is a toy/ugly little
example:
Click to see code
library(dplyr); library(ggplot2)
df <- tibble(
area = rep(c("Health", "Education"), 6),
sub_area = rep(c("Staff", "Projects", "Activities"), 4),
year = c(rep(2016, 6), rep(2017, 6)),
value = c(15000, 12000, 18000, 24000, 14000, 12000, 13000, 16000, 11000,
8000, 15000, 19000)
) %>% arrange(area)
df %>% filter(area == "Health") %>% {
ggplot(.) + # add . to specify to insert results here
geom_line(aes(x = as.factor(year), y = value,
group = sub_area, color = sub_area), size = 2) +
geom_point(aes(x = as.factor(year), y = value,
group = sub_area, color = sub_area), size = 2) +
theme_minimal(base_size = 18) +
geom_text(data = dplyr::filter(.,
year == 2016 & sub_area == "Activities"), # and here
aes(x = as.factor(year), y = value,
color = sub_area, label = sub_area), size = 6,
hjust = 1.2) +
xlab(NULL) + ylab(NULL) + theme(legend.position = "none")
}
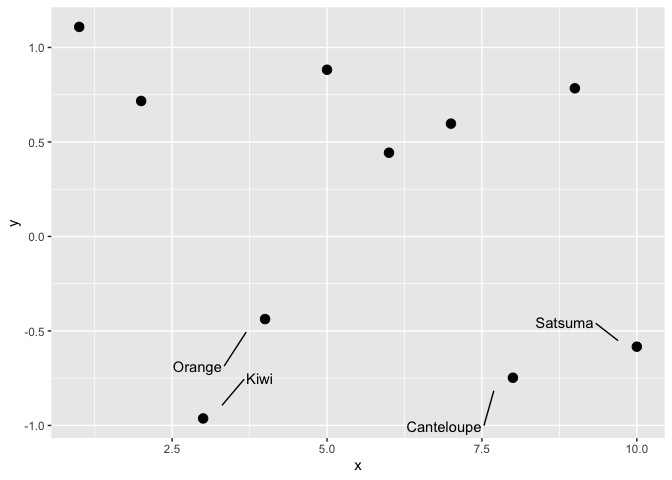
Neat little trick from James Goldie – you can also use
dplyr::case_when() to highlight certain points on a plot:
Click to see code
library(ggplot2); library(ggrepel); library(dplyr)
df <- tibble(
x = 1:10,
y = rnorm(10),
name = c("Apple", "Banana", "Kiwi", "Orange", "Watermelon",
"Grapes", "Pear", "Canteloupe", "Tomato", "Satsuma")) %>%
mutate(name_poor = case_when(
y < 0 ~ name,
TRUE ~ ""))
ggplot(df, aes(x = x, y = y)) +
geom_point(size = 3) +
geom_text_repel(aes(label = name_poor), point.padding = 2)First off, an informative SO
discussion
on the topic. Take-away: use specialized libraries (such as data.table
and Rccp) if you really need speed.
Sometimes for() loops may be necessary in R. They get a bad
reputation, but it’s not totally deserved (and the apply() family of
functions don’t necessarily speed things up, they’re just wrappers for
for() loops). This is old news in R, so let’s dig out an old bookmark
of
mine
on the subject, by Utkarsh Upadhyay. (From 2012!! Wow. Another
interesting
article on the
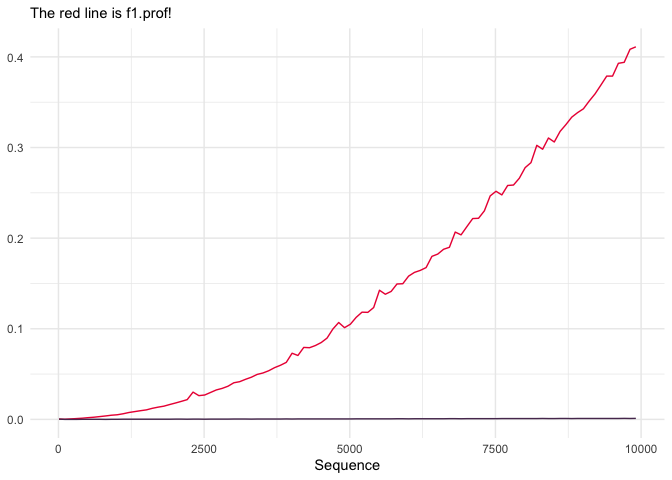
subject, by Noam Ross, led me to it.) What’s happening here? We’re
pre-allocating vectors of the size we need before we run our for
loops. Makes all the difference:
Click to see code
library(dplyr)
f1 <- function (n) {
l <- list()
for(i in 1:n) {
l <- append(l, i)
}
return(l)
}
f2 <- function (n) {
l <- vector("list", n) ## pre-allocate the size
for(i in 1:n) {
l[[i]] <- i
}
return(l)
}
warm.up <- function(f, n, times) {
system.time(sapply(1:times, function (i) f(n)), gcFirst = T)
}
run.all <- function (reps = 10) {
timesSeq <- seq(from = 10, to = 10000, by = 100)
message("Running f1 ...")
f1.prof <- sapply(timesSeq, function (arg) warm.up(f1, arg, reps)[1] / reps)
message("Running f2 ...")
f2.prof <- sapply(timesSeq, function (arg) warm.up(f2, arg, reps)[1] / reps)
return(tibble(
timesSeq = timesSeq,
f1.prof = f1.prof,
f2.prof = f2.prof
))
}
x <- run.all()
library(ggplot2)
ggplot(x, aes(x = timesSeq, group = 1)) +
geom_line(aes(y = f1.prof), colour = "#ec0b43") +
geom_line(aes(y = f2.prof), colour = "#58355e") +
ylab(NULL) + theme_minimal() + xlab("Sequence") +
labs(subtitle = "The red line is f1.prof!")You might be surprised to find out that using extra brackets and parentheses can slow your code down. To wit:
library(tictoc)
x <- 1:10
tic()
if(any(x > 5)) print("hey!")
#> [1] "hey!"
toc()
#> 0.001 sec elapsed
tic()
if(any(x > 5)) {
print("hey!")
}
#> [1] "hey!"
toc()
#> 0.001 sec elapsedI’ve seen this in a few places, but the most recent I remember was from Colin Fay on Twitter (Colin’s a fountain of little R tips, particularly for purrr).
You can save a lot of time by setting cache = TRUE in the knitr
options. I usually set warning and message to FALSE too, which
avoids package messages (like those from dplyr, for example) printing in
your RMarkdown document.
knitr::opts_chunk$set(
cache = TRUE,
message = FALSE,
warning = FALSE
)Cmd + option + i gives you an R code chunk. That’s handy.
If you use results = 'asis' in the head of your code chunk,
i.e. ```{r results = 'asis'}, RMarkdown will keep the result of the
chunk ‘alive’, so to speak, in the document, for you to use. A good
example of this is with JavaScript, which you can use to make a nifty
d3 plot. A few bloggers have noted this, although I
think the first was a blogger named
Alice.
The example given by Nick
Strayer
is a good one. Here, I’ll keep things ultra-simple. First we save our
data to the JSON format:
Click to see code
library(dplyr)
library(jsonlite)
send_df_to_js <- function(df){
cat(
paste(
'<script>
var data = ', toJSON(df),';
</script>', sep = "")
)
}
our_data <- data_frame(
country = c("Denmark", "Belgium", "Croatia", "Germany", "Greece",
"Italy", "Ireland", "Spain", "Switzerland", "France"),
gdp = c(53400, 41000, 12100, 42000, 18100, 30500, 62000,
26000, 73000, 36000)
)
send_df_to_js(our_data)So far, so good – the data is in our browser, as you can see from
opening the inspector on this page, going to the console, and typing
data (since it’s a JavaScript variable, defined with var = data), or
just from the mess printed here. You’ll need to paste the following
script into your document before the d3 script too: <script src="//d3js.org/d3.v4.min.js"></script> (here I’m using version 4, if
you use another version, obviously that part changes).
Then, following Nick, I’ve created a div element to put the
visualization in (it doesn’t show on GitHub, but make a simple test .Rmd
and you’ll see it works), it’s div id="viz"></div>. Then the
JavaScript code to make a simple bar graph (from
here):
// set the dimensions and margins of the graph
var margin = {top: 20, right: 20, bottom: 30, left: 40},
width = 960 - margin.left - margin.right,
height = 500 - margin.top - margin.bottom;
// set the ranges
var x = d3.scaleBand()
.range([0, width])
.padding(0.1);
var y = d3.scaleLinear()
.range([height, 0]);
var svg = d3.select("#viz").append("svg")
.attr("width", width + margin.left + margin.right)
.attr("height", height + margin.top + margin.bottom)
.append("g")
.attr("transform",
"translate(" + margin.left + "," + margin.top + ")")
x.domain(data.map(function(d) { return d.country; }));
y.domain([0, d3.max(data, function(d) { return d.gdp; })]);
svg.selectAll(".bar")
.data(data)
.enter().append("rect")
.attr("class", "bar")
.attr("x", function(d) { return x(d.country); })
.attr("width", x.bandwidth())
.attr("y", function(d) { return y(d.gdp); })
.attr("height", function(d) { return height - y(d.gdp); });
svg.append("g")
.attr("transform", "translate(0," + height + ")")
.call(d3.axisBottom(x));
// add the y Axis
svg.append("g")
.call(d3.axisLeft(y));And there you go, you’ll have a nice d3 bar graph with data you made/tidied with R.
I often find the syntax for negating %in% to be weird. For example:
Click to see code
library(dplyr)
library(tibble)
mtcars %>%
rownames_to_column(var = "cars") %>%
filter(!cars %in% c("Toyota Corolla", "Honda Civic", "Datsun 710"))
#> cars mpg cyl disp hp drat wt qsec vs am gear carb
#> 1 Mazda RX4 21.0 6 160.0 110 3.90 2.620 16.46 0 1 4 4
#> 2 Mazda RX4 Wag 21.0 6 160.0 110 3.90 2.875 17.02 0 1 4 4
#> 3 Hornet 4 Drive 21.4 6 258.0 110 3.08 3.215 19.44 1 0 3 1
#> 4 Hornet Sportabout 18.7 8 360.0 175 3.15 3.440 17.02 0 0 3 2
#> 5 Valiant 18.1 6 225.0 105 2.76 3.460 20.22 1 0 3 1
#> 6 Duster 360 14.3 8 360.0 245 3.21 3.570 15.84 0 0 3 4
#> 7 Merc 240D 24.4 4 146.7 62 3.69 3.190 20.00 1 0 4 2
#> 8 Merc 230 22.8 4 140.8 95 3.92 3.150 22.90 1 0 4 2
#> 9 Merc 280 19.2 6 167.6 123 3.92 3.440 18.30 1 0 4 4
#> 10 Merc 280C 17.8 6 167.6 123 3.92 3.440 18.90 1 0 4 4
#> 11 Merc 450SE 16.4 8 275.8 180 3.07 4.070 17.40 0 0 3 3
#> 12 Merc 450SL 17.3 8 275.8 180 3.07 3.730 17.60 0 0 3 3
#> 13 Merc 450SLC 15.2 8 275.8 180 3.07 3.780 18.00 0 0 3 3
#> 14 Cadillac Fleetwood 10.4 8 472.0 205 2.93 5.250 17.98 0 0 3 4
#> 15 Lincoln Continental 10.4 8 460.0 215 3.00 5.424 17.82 0 0 3 4
#> 16 Chrysler Imperial 14.7 8 440.0 230 3.23 5.345 17.42 0 0 3 4
#> 17 Fiat 128 32.4 4 78.7 66 4.08 2.200 19.47 1 1 4 1
#> 18 Toyota Corona 21.5 4 120.1 97 3.70 2.465 20.01 1 0 3 1
#> 19 Dodge Challenger 15.5 8 318.0 150 2.76 3.520 16.87 0 0 3 2
#> 20 AMC Javelin 15.2 8 304.0 150 3.15 3.435 17.30 0 0 3 2
#> 21 Camaro Z28 13.3 8 350.0 245 3.73 3.840 15.41 0 0 3 4
#> 22 Pontiac Firebird 19.2 8 400.0 175 3.08 3.845 17.05 0 0 3 2
#> 23 Fiat X1-9 27.3 4 79.0 66 4.08 1.935 18.90 1 1 4 1
#> 24 Porsche 914-2 26.0 4 120.3 91 4.43 2.140 16.70 0 1 5 2
#> 25 Lotus Europa 30.4 4 95.1 113 3.77 1.513 16.90 1 1 5 2
#> 26 Ford Pantera L 15.8 8 351.0 264 4.22 3.170 14.50 0 1 5 4
#> 27 Ferrari Dino 19.7 6 145.0 175 3.62 2.770 15.50 0 1 5 6
#> 28 Maserati Bora 15.0 8 301.0 335 3.54 3.570 14.60 0 1 5 8
#> 29 Volvo 142E 21.4 4 121.0 109 4.11 2.780 18.60 1 1 4 2‘Not-cars-in-list’ is defo weird for me. So I usually use Negate():
library(dplyr)
library(tibble)
'%ni%' <- Negate('%in%')
mtcars %>%
rownames_to_column(var = "cars") %>%
filter(cars %ni% c("Toyota Corolla", "Honda Civic", "Datsun 710"))
#> cars mpg cyl disp hp drat wt qsec vs am gear carb
#> 1 Mazda RX4 21.0 6 160.0 110 3.90 2.620 16.46 0 1 4 4
#> 2 Mazda RX4 Wag 21.0 6 160.0 110 3.90 2.875 17.02 0 1 4 4
#> 3 Hornet 4 Drive 21.4 6 258.0 110 3.08 3.215 19.44 1 0 3 1
#> 4 Hornet Sportabout 18.7 8 360.0 175 3.15 3.440 17.02 0 0 3 2
#> 5 Valiant 18.1 6 225.0 105 2.76 3.460 20.22 1 0 3 1
#> 6 Duster 360 14.3 8 360.0 245 3.21 3.570 15.84 0 0 3 4
#> 7 Merc 240D 24.4 4 146.7 62 3.69 3.190 20.00 1 0 4 2
#> 8 Merc 230 22.8 4 140.8 95 3.92 3.150 22.90 1 0 4 2
#> 9 Merc 280 19.2 6 167.6 123 3.92 3.440 18.30 1 0 4 4
#> 10 Merc 280C 17.8 6 167.6 123 3.92 3.440 18.90 1 0 4 4
#> 11 Merc 450SE 16.4 8 275.8 180 3.07 4.070 17.40 0 0 3 3
#> 12 Merc 450SL 17.3 8 275.8 180 3.07 3.730 17.60 0 0 3 3
#> 13 Merc 450SLC 15.2 8 275.8 180 3.07 3.780 18.00 0 0 3 3
#> 14 Cadillac Fleetwood 10.4 8 472.0 205 2.93 5.250 17.98 0 0 3 4
#> 15 Lincoln Continental 10.4 8 460.0 215 3.00 5.424 17.82 0 0 3 4
#> 16 Chrysler Imperial 14.7 8 440.0 230 3.23 5.345 17.42 0 0 3 4
#> 17 Fiat 128 32.4 4 78.7 66 4.08 2.200 19.47 1 1 4 1
#> 18 Toyota Corona 21.5 4 120.1 97 3.70 2.465 20.01 1 0 3 1
#> 19 Dodge Challenger 15.5 8 318.0 150 2.76 3.520 16.87 0 0 3 2
#> 20 AMC Javelin 15.2 8 304.0 150 3.15 3.435 17.30 0 0 3 2
#> 21 Camaro Z28 13.3 8 350.0 245 3.73 3.840 15.41 0 0 3 4
#> 22 Pontiac Firebird 19.2 8 400.0 175 3.08 3.845 17.05 0 0 3 2
#> 23 Fiat X1-9 27.3 4 79.0 66 4.08 1.935 18.90 1 1 4 1
#> 24 Porsche 914-2 26.0 4 120.3 91 4.43 2.140 16.70 0 1 5 2
#> 25 Lotus Europa 30.4 4 95.1 113 3.77 1.513 16.90 1 1 5 2
#> 26 Ford Pantera L 15.8 8 351.0 264 4.22 3.170 14.50 0 1 5 4
#> 27 Ferrari Dino 19.7 6 145.0 175 3.62 2.770 15.50 0 1 5 6
#> 28 Maserati Bora 15.0 8 301.0 335 3.54 3.570 14.60 0 1 5 8
#> 29 Volvo 142E 21.4 4 121.0 109 4.11 2.780 18.60 1 1 4 2Often, the daily builds of RStudio have some interesting things happening (although, of course, they might be a bit buggy). Using a combination of scripts from Bob Rudis and Aron Atkins, with a few adjustments, we can download and install the latest build at a certain time each day (I’ve stuck with Bob’s choice of 7:30 am). This is for Mac OS only (Aron’s original script also allows for Ubuntu).
Click to see code
First, create a preference list file (.plist) – I’ve called it
‘UpdateRstudio.plist’ (You can do this on Terminal in a mac with
touch UpdateRstudio.plist). The content of this is as follows:
<?xml version="1.0" encoding="UTF-8"?>
<!DOCTYPE plist PUBLIC "-//Apple//DTD PLIST 1.0//EN" "http://www.apple.com/DTDs/PropertyList-1.0.dtd">
<plist version="1.0">
<dict>
<key>Label</key>
<string>UpdateRStudio</string>
<key>Program</key>
<string>/usr/local/bin/rsupd</string>
<key>StartCalendarInterval</key>
<dict>
<key>Hour</key>
<integer>7</integer>
<key>Minute</key>
<integer>30</integer>
</dict>
</dict>
</plist>
If you’d like to update at a different time, you can change the
<key>Hour</key><integer>7</integer><key>Minute</key><integer>30</integer>
part. This file goes in /Library/LaunchAgents/.
Also, notice that the .plist is directed towards
/usr/local/bin/rsupd – that is the location of the next file we’re
going to make. It is:
#!/bin/bash
set -e
install_macos_daily() {
REDIRECT_URL="https://www.rstudio.org/download/latest/daily/desktop/mac/RStudio-latest.dmg"
echo "Discovering daily build from: ${REDIRECT_URL}"
# Perform a HEAD request to find the redirect target. We use the name of the
# file to derive the mounted volume name.
RELEASE_URL=$(curl -s -L -I -o /dev/null -w '%{url_effective}' "${REDIRECT_URL}")
if [ "${RELEASE_URL}" == "" ]; then
echo "Could not extract daily build URL from listing; maybe rstudio.org is having problems?"
echo "Check: ${DAILY_LIST_URL}"
exit 1
fi
echo "Downloading daily build from: ${RELEASE_URL}"
cd /tmp
TARGET=$(basename "${RELEASE_URL}")
# Volume name mirrors the DMG filename without extension.
# Simpler than parsing hdiutil output.
VOLUME_NAME=$(basename "${TARGET}" .dmg)
VOLUME_MOUNT="/Volumes/${VOLUME_NAME}"
curl -L -o "${TARGET}" "${RELEASE_URL}"
hdiutil attach -quiet "${TARGET}"
# Remove any prior installation.
rm -rf /Applications/RStudio.app
cp -R "${VOLUME_MOUNT}/RStudio.app" /Applications
hdiutil detach -quiet "${VOLUME_MOUNT}"
rm "${TARGET}"
echo "Installed ${VOLUME_NAME} to /Applications"
}
if [[ `uname -s` = "Darwin" ]]; then
install_macos_daily
else
echo "This script only works on OSX/macOS."
exit 1
fiAs I said, this is saved as rsupd, into /usr/local/bin. We then make
it an executable with chmod 755 /usr/local/bin/rsupd. Then you load
the preference list with launchctl load -w /Library/LaunchAgents/UpdateRStudio.plist (use unload here when you
want to stop it). Voilà, fresh RStudio for thee everyday.
ts plus Shift and Tab gives you a nice data-stamped code section:
# Sun Mar 25 14:36:54 2018 ------------------------------Looking for something in code or a file in a project? Try Cmd + Shift + F.
Nice.