This repository attempts to answer the question:
Is Python too slow?
- Short answer: Yes.
- Long answer: Still yes. But you got options!
- Adding numba decorators & compiling your code (LLVM)
- Adding cython type annotations & compiling your code (C)
- Leveraging an external c implementation with ctypes.
Consider a simple function to test if a number is prime:
Given a number
nfind if there's a divisor in the firstn/2numbers.
Of course, we can come with a better prime-testing solution, but we won't. This solution is exactly what we need: inefficient and cpu-intensive.
The goal: Compare the python implementation to a pure C solution.
C implementation:
int isPrime(int num) {
for(int i=2; i<=num/2; i++)
if (!(num%i))
return 0;
return 1;
}This translates to python as:
def is_prime(n: int) -> int:
for i in range(2, (n // 2) + 1):
if not (n % i):
return 0
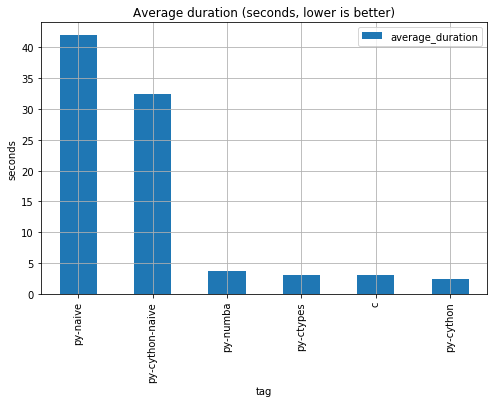
return 1As expected, plain vanilla python is painfully slow. By compiling the plain python code without and modification via cython
we can get a 29% performance increase. If we don't bother adding a simple numba decorator, we can get a 10X performance
increase. These 2 solutions are the easiest to implement because doesn't require changing our is_prime function definitions and requires
minimal boilerplate.
If you want even better performance:
- Use directly a c external function via Python CTypes.
- Add cython type annotations before compiling.
| benchmark_tag | samples | min_duration | average_duration | max_duration |
|---|---|---|---|---|
| py-naive | 10.0 | 41.59 | 41.93 | 42.68 |
| py-cython-naive | 10.0 | 32.15 | 32.40 | 32.94 |
| py-numba | 10.0 | 3.71 | 3.75 | 3.87 |
| py-ctypes | 10.0 | 3.03 | 3.08 | 3.16 |
| c | 10.0 | 3.04 | 3.05 | 3.07 |
| py-cython | 10.0 | 2.39 | 2.42 | 2.44 |
These tests were performed using python 3.7.5. We recommend creating a virtualenv and
installing the dependencies:
$ pip install -r requirements.txt
We need to compile the c, numba & cython implementation. For this, simple run:
$ bash build.sh --c
- The expected output is a
bencharkcfile. - Alternatively:
gcc -o benchmarkc benchmark.c
$ bash build.sh --py-numba
- The expected output is a
numba_impl.*file. - Alternatively:
python numba_source.py
$ bash build.sh --py-cython
- Expected output are the compiled code for
cython_impl.pyandcython_naive_impl.py.
Run benchmarks individually:
- C:
bash run.sh --c - Python Naive:
bash run.sh --py-naive - Python Numba:
bash run.sh --py-numba - Python Cython:
bash run.sh --py-cython - Python CTypes:
bash run.sh --py-ctypes
Or run all at once:
BENCH_PARALLELISM=false python benchmarks.py 1
- The output will be appended to a file:
benchmark-outputs-xxxx.json - Each benchmark is executed on an independent process. If
BENCH_PARALLELISMisfalse, then the processes will be executed sequentially. - Change the
1if you want to run each benchmark more than once (consider that different processes will be used for each).