This repo was created to support "Machine Learning & Artificial Intelligence" exam.
In this code i'm using a Brain Tumor dataset for trying some of the classical machine learning alghoritms.
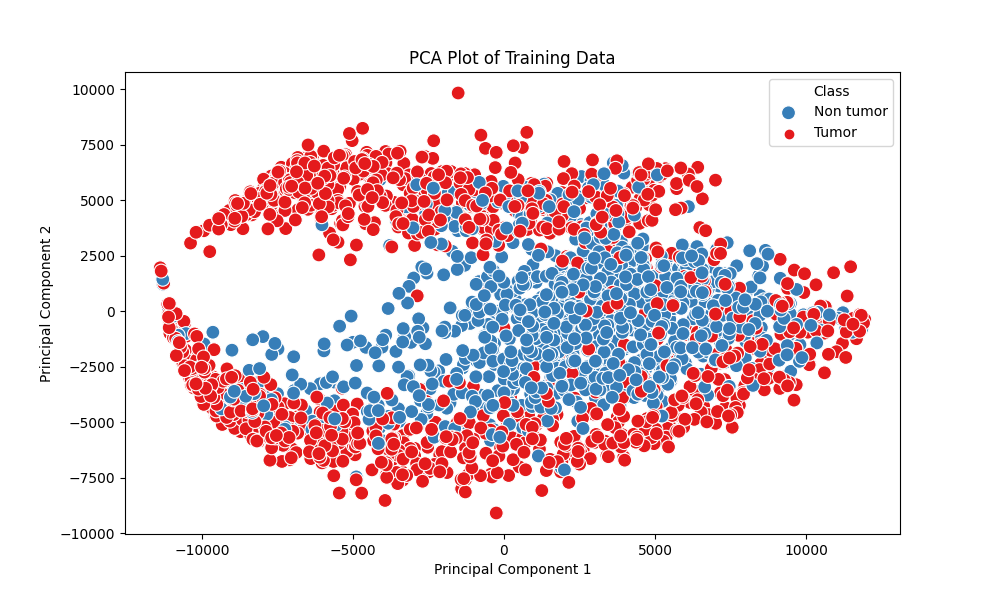
- PCA
- KNN with differents K
- PCA + KNN
- SVM with differents Kernels
- PCA + SVM
- A neural network (CNN)
In order to run the code you need to download the dataset and extract into the code folder. The codespace need to look like that:
UNIVR-MachineLearning-Exam:
|- dataset/
| |- Brain Tumor/
| |- all the images are here
| |- Brain Tumor.csv
| |- bt_dataset_t3.csv
|
|- README.md
|- .gitignore
|- requirements.txt
|- NeuralNetwork.py
|- main.py
number of components = 100
-
k=1, accuracy=95.41%
-
k=3, accuracy=95.76%
-
k=5, accuracy=92.23%
-
k=7, accuracy=91.17%
-
k=9, accuracy=91.52%
-
k=3 achieved highest accuracy of 95.76% on validation data
EVALUATION ON TESTING DATA
precision recall f1-score support
0 0.95 0.94 0.95 526
1 0.93 0.94 0.93 415
accuracy 0.94 941
macro avg 0.94 0.94 0.94 941
weighted avg 0.94 0.94 0.94 941
number of components for PCA = 2
-
k=1, accuracy=77.39%
-
k=3, accuracy=80.92%
-
k=5, accuracy=81.63%
-
k=7, accuracy=81.63%
-
k=9, accuracy=82.33%
-
k=9 achieved highest accuracy of 82.33% on validation data
EVALUATION ON TESTING DATA
precision recall f1-score support
0 0.82 0.83 0.82 526
1 0.78 0.76 0.77 415
accuracy 0.80 941
macro avg 0.80 0.80 0.80 941
weighted avg 0.80 0.80 0.80 941
EVALUATION ON TESTING DATA
-
Linear
-
The model is 75% accurate
-
Precision: 0.693
-
Recall: 0.785
precision recall f1-score support Non tumor 0.81 0.73 0.77 526 Tumor 0.69 0.79 0.74 415 accuracy 0.75 941 macro avg 0.75 0.76 0.75 941 weighted avg 0.76 0.75 0.75 941 -
Elapsed time on linear: 50.239 s
-
-
Polynomial
-
The model is 48.990% accurate
-
Precision: 0.463
-
Recall: 0.997
precision recall f1-score support Non tumor 0.98 0.09 0.16 526 Tumor 0.46 1.00 0.63 415 accuracy 0.49 941 macro avg 0.72 0.54 0.40 941 weighted avg 0.75 0.49 0.37 941 -
Elapsed time on polynomial: 56.108 s
-
-
RBF
-
The model is 70.031 accurate
-
Precision: 0.620
-
Recall: 0.824
precision recall f1-score support Non tumor 0.81 0.60 0.69 526 Tumor 0.62 0.82 0.71 415 accuracy 0.70 941 macro avg 0.72 0.71 0.70 941 weighted avg 0.73 0.70 0.70 941 -
Elapsed time on RBF: 60.251 s
-
number of components for PCA = 3
EVALUATION ON TESTING DATA
-
Linear
-
The model is 50.903% accurate
-
Precision: 0.445
-
Recall: 0.465
precision recall f1-score support Non tumor 0.56 0.54 0.55 526 Tumor 0.45 0.47 0.46 415 accuracy 0.51 941 macro avg 0.50 0.50 0.50 941 weighted avg 0.51 0.51 0.51 941 -
Elapsed time on linear: 0.07 s
-
-
Poly
-
The model is 80.871% accurate
-
Precision: 0.723
-
Recall: 0.915
precision recall f1-score support Non tumor 0.92 0.72 0.81 526 Tumor 0.72 0.92 0.81 415 accuracy 0.81 941 macro avg 0.82 0.82 0.81 941 weighted avg 0.83 0.81 0.81 941 -
Elapsed time on poly: 0.073 s
-
-
RBF
-
The model is 87.353% accurate
-
Precision: 0.854
-
Recall: 0.860
precision recall f1-score support Non tumor 0.89 0.88 0.89 526 Tumor 0.85 0.86 0.86 415 accuracy 0.87 941 macro avg 0.87 0.87 0.87 941 weighted avg 0.87 0.87 0.87 941 -
Elapsed time on rbf: 0.076 s
-
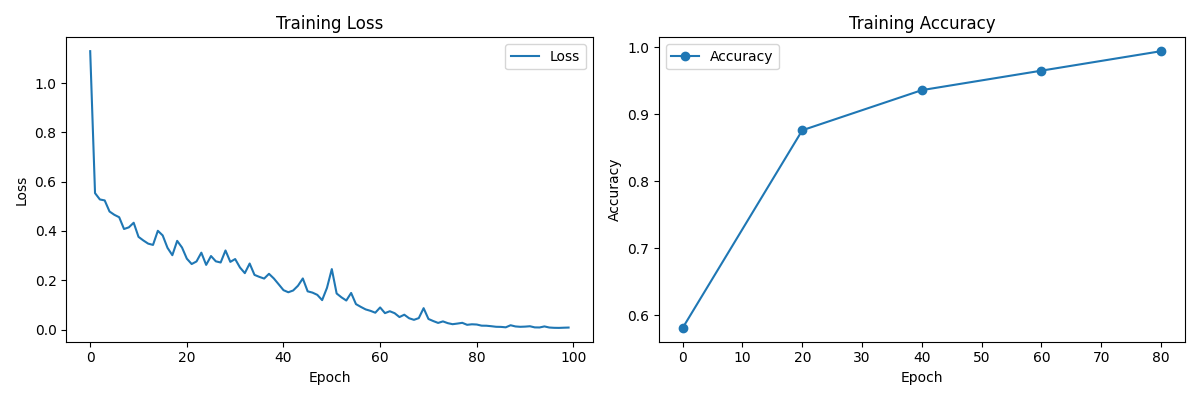
Neural Network CNN
cuda:0 - RTX 3070 ti
- Train Shape : (2821, 240, 240)
- Test Shape(941, 240, 240)
Epoch: 0. Loss: 0.18476. | Accuracy (on trainset/self): 0.58099
Epoch: 20. Loss: 0.14416. | Accuracy (on trainset/self): 0.87593
Epoch: 40. Loss: 0.01653. | Accuracy (on trainset/self): 0.93583
Epoch: 60. Loss: 0.08495. | Accuracy (on trainset/self): 0.96490
Epoch: 80. Loss: 0.02391. | Accuracy (on trainset/self): 0.99397
Accuracy on test set: 0.95324
Negative Positive
Negative 508 (TN) 18 (FP)
Positive 26 (FN) 389 (TP)
Precision : 0.955
Recall : 0.937