In this exercise we will build a simple model that can predict the carbon dioxide uptake in MOFs.
You can find more background information in the accompanying notes.
If you find some errors, typos or issues feel free to open an issue or directly make a pull request.
Some parts of this exercise will need more computational resources than most of the other MolSim exercises. If you have a modern laptop, we recommend you run them on the laptop. If you do not want to use your machine or the cluster, you can also run the exercises on Google Colab or Binder.
First, you need to create a conda environment using the environment.yml
file we provide in this repository.
If you do not already have anaconda installed, head over to https://docs.conda.io/en/latest/miniconda.html and install a Python 3.7 version for your operating system.
Then, create a new folder and clone this repository
git clone --depth 1 https://github.com/kjappelbaum/ml_molsim2020.git
cd ml_molsim2020
Now you can create a new conda environment.
conda env create --file environment.yml
After this completed, active the environment and open the jupyter notebook
conda activate molsim_ml
jupyter notebook molsim_ml.ipynb
Here, you can use relatively powerful computing resources from Google for free (like GPUs and TPUs). Click the "Open in Colab" button on the top and run the first three cells to install the dependencies. Then you should be able to use the notebook in Colab.
Make sure to make a copy into your Google Drive and work on this copy. And not on the shared notebook!
Note: If you have a Google Account from your organization, e.g. university, you might need to log out and use your personal account as many organizations block third-party applications.
MyBinder is a free service to run jupyter notebooks from git repositories. Compared to the Colab, computing resources are limited, and it can take some time to start your instance. Just click on the Binder button and wait for your instance to start.
Note: If you encounter a 404 error (as shown below), click on the jupyter symbol and you will be redirected
to a file browser where you can select molsim_ml.ipynb.
We also host a Kaggle competition for this exercise.
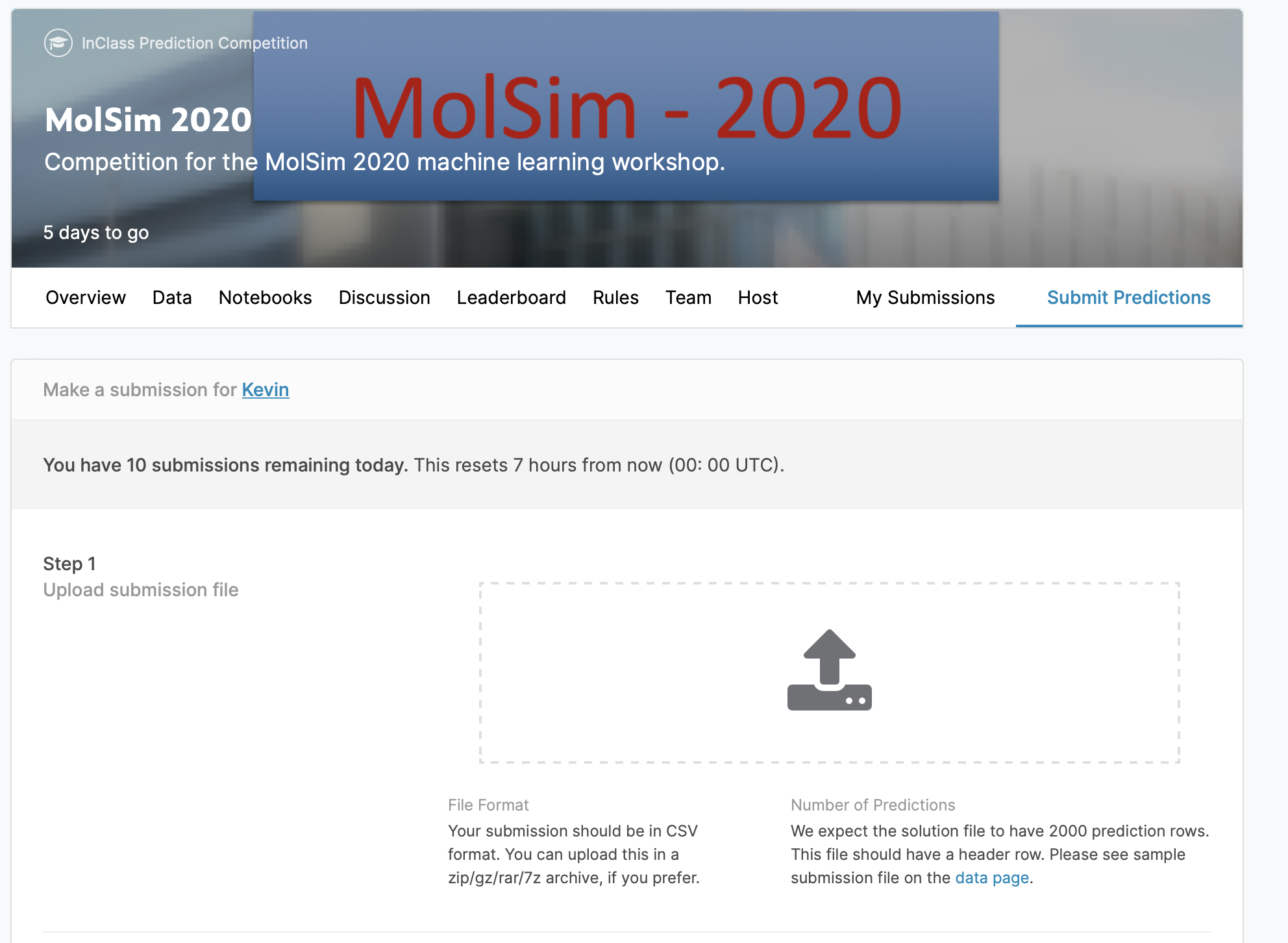
To join this competition, you need to create an account on www.kaggle.com and use the features (features.csv) you can find on the competition site or also in the data directory to predict the high-pressure uptake of carbon dioxide.
You can either drag and drop your solution .csv on the submission
page.
An example submission file can be found in data/submission.csv.
Please share your code with a Kaggle notebook!
We want to thank Leopold Talirz for incredibly valuable feedback and input, Peter Alexander Knudsen for spotting typos, and Berend Smit for sponsoring the main prize for the Kaggle competition.