cellxgene VIP unleashes full power of interactive visualization, plotting and analysis of scRNA-seq and spatial transcriptomics data in the scale of millions of cells
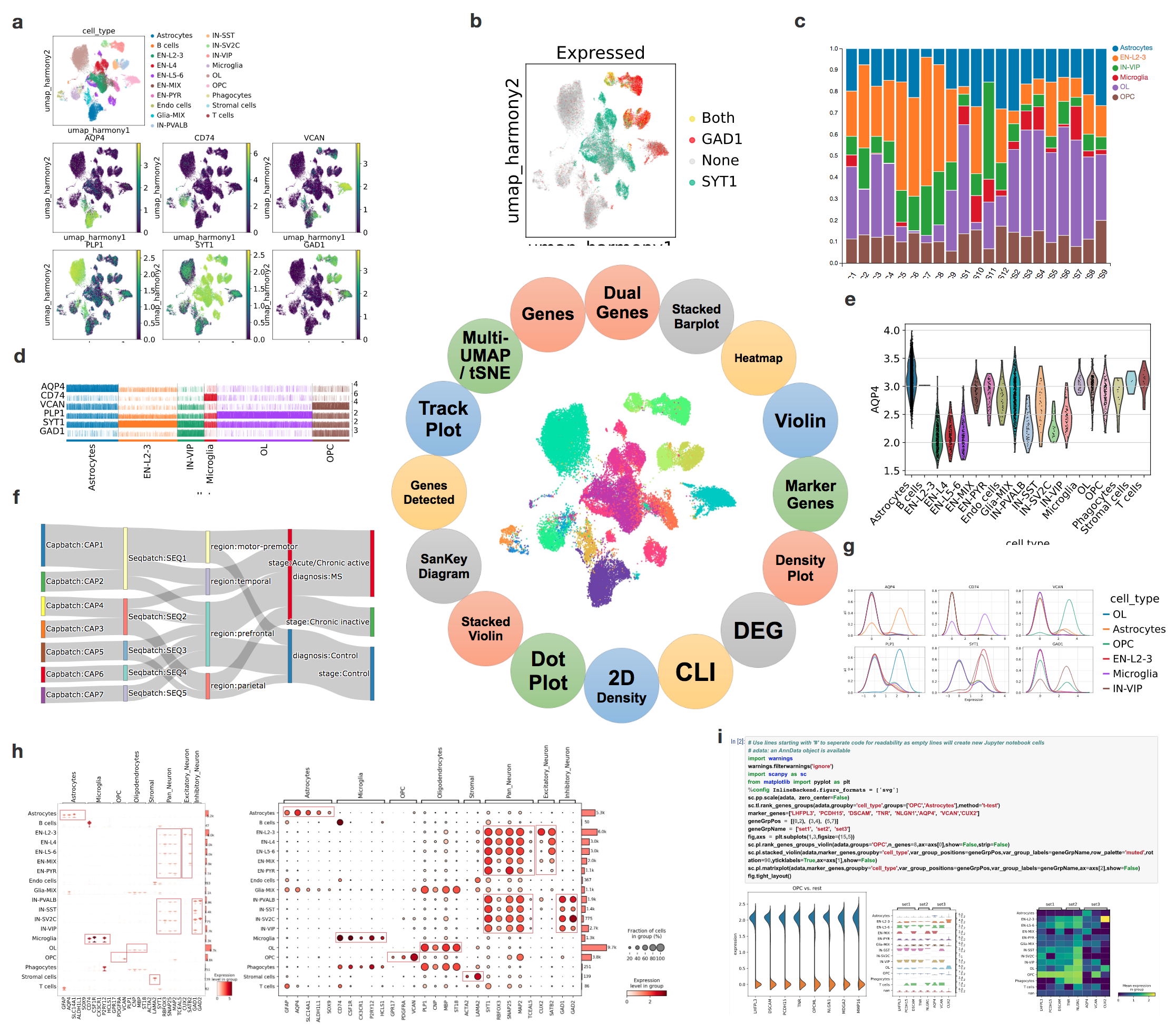
To meet the growing demands from scientists to effectively extract deep insights from single cell and single nuclei RNA-seq datasets, we developed cellxgene VIP, a frontend interactive visualization plugin to cellxgene framework, which greatly expanded capabilities of the base tool in the following aspects. First, it generates a comprehensive set of eighteen commonly used quality control and analytical plots in high resolution with rich customization in real time. Second, it provides more advanced analytical functions, such as differential gene expression analysis, gene set enrichment and marker gene identification. Third, it empowers advanced users to perform analysis in a Jupyter notebook like Command Line Interface (CLI) environment by programming in Python and/or R directly without limitation of available interactive modules. Finally, it pioneers methods to visualize spatial transcriptomics embedding aligned with histological image on one slice or multiple slices in a grid format to fully leverage the aforementioned functionalities. Taken together, the open-source tool makes large scale scRNA-seq data visualization and analysis more accessible in a user-friendly manner and fosters computational reproducibility by simplifying data and code reuse through the CLI. Going forward, it has the potential to become an ecosystem for the scientific community to contribute even more modules to the Swiss Army knife of scRNA-seq data exploration tool.
Demo site: https://cellxgenevip-ms.bxgenomics.com
Online tutorial: https://interactivereport.github.io/cellxgene_VIP/tutorial/docs
Pre-print: https://www.biorxiv.org/content/10.1101/2020.08.28.270652v1
Installation instruction
1. Install anaconda if not available on server (https://docs.anaconda.com/anaconda/install/linux/)
bash ~/Downloads/Anaconda3-2020.02-Linux-x86_64.sh2. Create and enable conda environment
git clone https://github.com/interactivereport/cellxgene_VIP.git
cd cellxgene_VIP
source <path to Anaconda3>/etc/profile.d/conda.sh (Default: /opt/anaconda3/etc/profile.d/conda.sh)
conda config --set channel_priority flexible
conda env create -n <env name, such as: VIP> -f VIP.yml (system-wide R) or VIP_conda_R.yml (local R under conda, no root privilege needed)
For Mac User, conda env create -n <env name, such as: VIP> -f VIP.macOS.yml
conda activate <env name, such as: VIP>
or
source activate <env name>3. Install cellxgene by running config.sh in "cellxgene_VIP" directory
./config.sh
For Mac User, ./config.macOS.sh4. Install R packages
export LIBARROW_MINIMAL=false
# ensure that the right instance of R is used. e.g. system-wide: /bin/R or /usr/bin/R ; local R under conda: ~/.conda/envs/VIP_conda_R/bin/R
which R
R -q -e 'if(!require(devtools)) install.packages("devtools",repos = "http://cran.us.r-project.org")'
R -q -e 'if(!require(Cairo)) devtools::install_version("Cairo",version="1.5-12",repos = "http://cran.us.r-project.org")'
R -q -e 'if(!require(foreign)) devtools::install_version("foreign",version="0.8-76",repos = "http://cran.us.r-project.org")'
R -q -e 'if(!require(ggpubr)) devtools::install_version("ggpubr",version="0.3.0",repos = "http://cran.us.r-project.org")'
R -q -e 'if(!require(ggrastr)) devtools::install_version("ggrastr",version="0.1.9",repos = "http://cran.us.r-project.org")'
R -q -e 'if(!require(arrow)) devtools::install_version("arrow",version="2.0.0",repos = "http://cran.us.r-project.org")'
R -q -e 'if(!require(Seurat)) devtools::install_version("Seurat",version="3.2.3",repos = "http://cran.us.r-project.org")'
R -q -e 'if(!require(rmarkdown)) devtools::install_version("rmarkdown",version="2.5",repos = "http://cran.us.r-project.org")'
R -q -e 'if(!require(tidyverse)) devtools::install_version("tidyverse",version="1.3.0",repos = "http://cran.us.r-project.org")'
R -q -e 'if(!require(viridis)) devtools::install_version("viridis",version="0.5.1",repos = "http://cran.us.r-project.org")'
R -q -e 'if(!require(hexbin)) devtools::install_version("hexbin",version="1.28.2",repos = "http://cran.us.r-project.org")'
R -q -e 'if(!require(ggforce)) devtools::install_version("ggforce",version="0.3.3",repos = "http://cran.us.r-project.org")'
R -q -e 'if(!require(RcppRoll)) devtools::install_version("RcppRoll",version="0.3.0",repos = "http://cran.r-project.org")'
R -q -e 'if(!require(fastmatch)) devtools::install_version("fastmatch",version="1.1-3",repos = "http://cran.r-project.org")'
R -q -e 'if(!require(BiocManager)) devtools::install_version("BiocManagerre",version="1.30.10",repos = "http://cran.us.r-project.org")'
R -q -e 'if(!require(fgsea)) BiocManager::install("fgsea")'
R -q -e 'if(!require(rtracklayer)) BiocManager::install("rtracklayer")'
# These should be already installed as dependencies of above packages
R -q -e 'if(!require(dbplyr)) devtools::install_version("dbplyr",version="1.0.2",repos = "http://cran.us.r-project.org")'
R -q -e 'if(!require(RColorBrewer)) devtools::install_version("RColorBrewer",version="1.1-2",repos = "http://cran.us.r-project.org")'
R -q -e 'if(!require(glue)) devtools::install_version("glue",version="1.4.2",repos = "http://cran.us.r-project.org")'
R -q -e 'if(!require(gridExtra)) devtools::install_version("gridExtra",version="2.3",repos = "http://cran.us.r-project.org")'
R -q -e 'if(!require(ggrepel)) devtools::install_version("ggrepel",version="0.8.2",repos = "http://cran.us.r-project.org")'
R -q -e 'if(!require(MASS)) devtools::install_version("MASS",version="7.3-51.6",repos = "http://cran.us.r-project.org")'
R -q -e 'if(!require(data.table)) devtools::install_version("data.table",version="1.13.0",repos = "http://cran.us.r-project.org")'5. Run cellxgene by specifiying a h5ad file storing scRNA-seq data along with a host and a port, use "ps" to find used ports to spare, see https://chanzuckerberg.github.io/cellxgene/posts/launch for details.
ps -ef | grep cellxgene
Rscript -e 'reticulate::py_config()'
# Run the following command if the output of the above command doesn't point to the Python in your env.
export RETICULATE_PYTHON=`which python`
cellxgene launch --host <xxx> --port <xxx> --disable-annotations --verbose <h5ad file>6. From web browser (Chrome is preferred, Version 87.0.4280.88 or 87.0.4280.141 is used), access http(s)://host:port
You should be able to see this in Console of Chrome Developer Tools if everything is right.

note: while spinning up the cellxgene from HPC, do NOT use qlogin. ssh directly to the server.
Updating
./update.VIPInterface.sh all # if "interface.html" or "VIPInterface.py" is modified, often.
./update.index_template.sh # if jsPanel is modified, very rare.