What’s Behind the Mask: Understanding Masked Graph Modeling for Graph Autoencoders (KDD 2023)
MaskGAE: Masked Graph Modeling Meets Graph Autoencoders (arXiv 2022)Jintang Li, Ruofan Wu, Wangbin Sun, Liang Chen, Sheng Tian, Liang Zhu, Changhua Meng, Zibin Zheng, Weiqiang Wang
This repository is an official PyTorch implementation of MaskGAE.
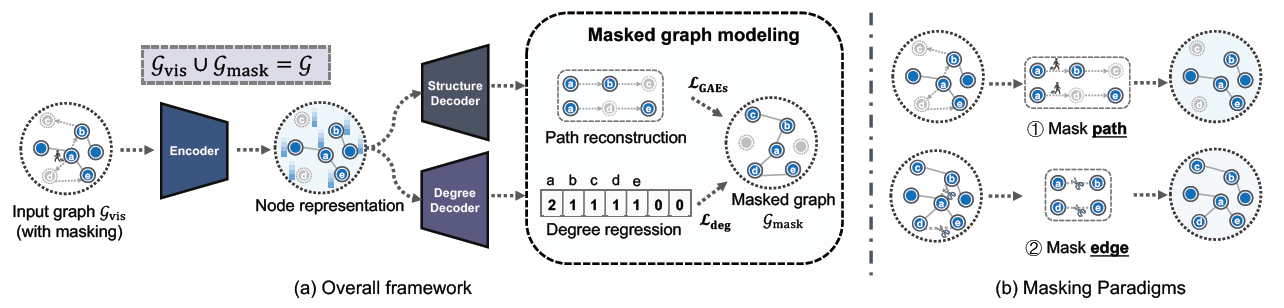
Fig. 1. MaskGAE framework and masking strategies.
The last years have witnessed the emergence of a promising self-supervised learning strategy, referred to as masked autoencoding. However, there is a lack of theoretical understanding of how masking matters on graph autoencoders (GAEs). In this work, we present masked graph autoencoder (MaskGAE), a self-supervised learning framework for graph-structured data. Different from standard GAEs, MaskGAE adopts masked graph modeling (MGM) as a principled pretext task - masking a portion of edges and attempting to reconstruct the missing part with partially visible, unmasked graph structure. To understand whether MGM can help GAEs learn better representations, we provide both theoretical and empirical evidence to comprehensively justify the benefits of this pretext task. Theoretically, we establish close connections between GAEs and contrastive learning, showing that MGM significantly improves the self-supervised learning scheme of GAEs. Empirically, we conduct extensive experiments on a variety of graph benchmarks, demonstrating the superiority of MaskGAE over several state-of-the-arts on both link prediction and node classification tasks.
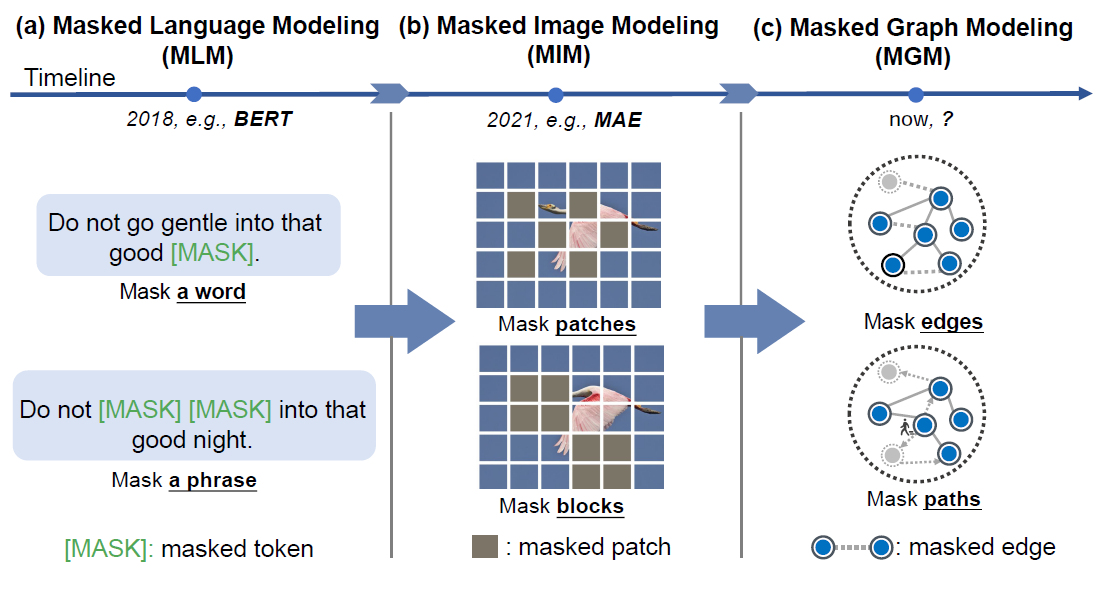
Fig. 2. Comparison of masked language modeling (MLM), masked image modeling (MIM) and masked graph modeling (MGM).
Higher versions should be also available.
- numpy==1.21.6
- torch==1.12.1+cu102
- torch-cluster==1.6.0
- torch_geometric>=2.4.0
- torch-scatter==2.0.9
- torch-sparse==0.6.14
- scipy==1.7.3
- texttable==1.6.2
- CUDA 10.2
- CUDNN 7.6.0
pip install -r requirements.txt| Dataset | #Nodes | #Edges | #Features | #Classes | Density |
|---|---|---|---|---|---|
| Cora | 2,708 | 10,556 | 1,433 | 7 | 0.144% |
| CiteSeer | 3,327 | 9,104 | 3,703 | 6 | 0.082% |
| Pubmed | 19,717 | 88,648 | 500 | 3 | 0.023% |
| Photo | 7,650 | 238,162 | 745 | 8 | 0.407% |
| Computer | 13,752 | 491,722 | 767 | 10 | 0.260% |
| arXiv | 16,9343 | 2,315,598 | 128 | 40 | 0.008% |
| MAG | 736,389 | 10,792,672 | 128 | 349 | 0.002% |
| Collab | 235,868 | 1,285,465 | 128 | - | 0.002% |
All datasets used throughout experiments are publicly available in PyTorch Geometric library.
- Cora
python train_linkpred.py --dataset Cora --bn
python train_linkpred.py --dataset Cora --bn --mask Edge- Citeseer
python train_linkpred.py --dataset Citeseer --bn
python train_linkpred.py --dataset Citeseer --bn --mask Edge- Pubmed
python train_linkpred.py --dataset Pubmed --bn --encoder_dropout 0.2
python train_linkpred.py --dataset Pubmed --bn --encoder_dropout 0.2 --mask Edge- Collab
python train_linkpred_ogb.py
python train_linkpred_ogb.py --mask Edge- Cora
python train_nodeclas.py --dataset Cora --bn --l2_normalize --alpha 0.004
python train_nodeclas.py --dataset Cora --bn --l2_normalize --alpha 0.003 --mask Edge --eval_period 10- Citeseer
python train_nodeclas.py --dataset Citeseer --bn --l2_normalize --nodeclas_weight_decay 0.1 --alpha 0.001 --lr 0.02
python train_nodeclas.py --dataset Citeseer --bn --l2_normalize --nodeclas_weight_decay 0.1 --alpha 0.001 --lr 0.02 --mask Edge --eval_period 20- Pubmed
python train_nodeclas.py --dataset Pubmed --bn --l2_normalize --alpha 0.001 --encoder_dropout 0.5 --decoder_dropout 0.5
python train_nodeclas.py --dataset Pubmed --bn --l2_normalize --alpha 0.001 --encoder_dropout 0.5 --mask Edge- Photo
python train_nodeclas.py --dataset Photo --bn --nodeclas_weight_decay 5e-3 --decoder_channels 128 --lr 0.005
python train_nodeclas.py --dataset Photo --bn --nodeclas_weight_decay 5e-3 --decoder_channels 64 --mask Edge- Computers
python train_nodeclas.py --dataset Computers --bn --encoder_dropout 0.5 --alpha 0.002 --encoder_channels 128 --hidden_channels 256 --eval_period 20
python train_nodeclas.py --dataset Computers --bn --encoder_dropout 0.5 --alpha 0.003 --encoder_channels 128 --hidden_channels 256 --eval_period 10 --mask Edge- arxiv
python train_nodeclas.py --dataset arxiv --bn --decoder_channels 128 --decoder_dropout 0. --decoder_layers 4 \
--encoder_channels 256 --encoder_dropout 0.2 --encoder_layers 4 \
--hidden_channels 512 --lr 0.0005 --nodeclas_weight_decay 0 --weight_decay 0.0001 --epochs 100 \
--eval_period 10
python train_nodeclas.py --dataset arxiv --bn --decoder_channels 128 --decoder_dropout 0. --decoder_layers 4 \
--encoder_channels 256 --encoder_dropout 0.2 --encoder_layers 4 \
--hidden_channels 512 --lr 0.0005 --nodeclas_weight_decay 0 --weight_decay 0.0001 --epochs 100 \
--eval_period 10 --mask Edge- MAG
python train_nodeclas.py --dataset mag --alpha 0.003 --bn --decoder_channels 128\
--encoder_channels 256 --encoder_dropout 0.7 --epochs 100 \
--hidden_channels 128 --nodeclas_weight_decay 1e-5 --weight_decay 5e-5 --eval_period 10
python train_nodeclas.py --dataset mag --alpha 0.003 --bn --decoder_channels 128
--encoder_channels 256 --encoder_dropout 0.7 --epochs 100 \
--hidden_channels 128 --nodeclas_weight_decay 1e-5 --weight_decay 5e-5 --eval_period 10 --mask Edge You can also simply run node_classification.ipynb to reproduce the results.
@inproceedings{maskgae,
author = {Jintang Li and
Ruofan Wu and
Wangbin Sun and
Liang Chen and
Sheng Tian and
Liang Zhu and
Changhua Meng and
Zibin Zheng and
Weiqiang Wang},
title = {What's Behind the Mask: Understanding Masked Graph Modeling for Graph Autoencoders},
booktitle = {KDD},
publisher = {{ACM}},
year = {2023}
}