MedRAG a systematic toolkit for Retrieval-Augmented Generation (RAG) on medical question answering (QA). MedRAG is used to implement various RAG systems for the benchmark study on our MIRAGE (Medical Information Retrieval-Augmented Generation Evaluation).
- (06/19/2024) Add supports for openai>=1.0.0. MedRAG now allows pre-determined snippets/snippet ids as input.
- (05/16/2024) Our paper has been accepted by ACL 2024 Findings!
- (04/26/2024) Add supports for
Google/gemini-1.0-proandmeta-llama/Meta-Llama-3-70B-Instruct. - (02/26/2024) The code has been updated. It supports all corpora and retrievers introduced in our paper now.
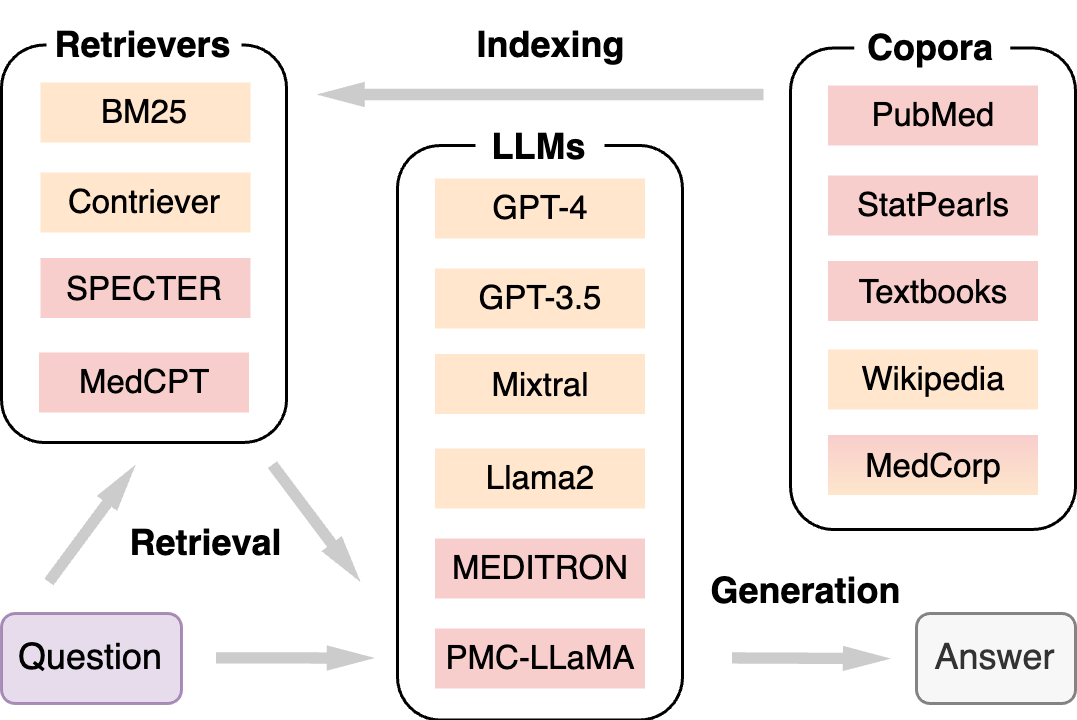
The following figure shows that MedRAG consists of three major components: Corpora, Retrievers, and LLMs.
For corpora used in MedRAG, we collect raw data from four different sources, including the commonly used PubMed for all biomedical abstracts, StatPearls for clinical decision support, medical Textbooks for domain-specific knowledge, and Wikipedia for general knowledge. We also provide a MedCorp corpus by combining all four corpora, facilitating cross-source retrieval. Each corpus is chunked into short snippets.
| Corpus | #Doc. | #Snippets | Avg. L | Domain |
|---|---|---|---|---|
| PubMed | 23.9M | 23.9M | 296 | Biomed. |
| StatPearls | 9.3k | 301.2k | 119 | Clinics |
| Textbooks | 18 | 125.8k | 182 | Medicine |
| Wikipedia | 6.5M | 29.9M | 162 | General |
| MedCorp | 30.4M | 54.2M | 221 | Mixed |
(#Doc.: numbers of raw documents; #Snippets: numbers of snippets (chunks); Avg. L: average length of snippets.)
For the retrieval algorithms, we only select some representative ones in MedRAG, including a lexical retriever (BM25), a general-domain semantic retriever (Contriever), a scientific-domain retriever (SPECTER), and a biomedical-domain retriever (MedCPT).
| Retriever | Type | Size | Metric | Domain |
|---|---|---|---|---|
| BM25 | Lexical | -- | BM25 | General |
| Contriever | Semantic | 110M | IP | General |
| SPECTER | Semantic | 110M | L2 | Scientific |
| MedCPT | Semantic | 109M | IP | Biomed. |
(IP: inner product; L2: L2 norm)
We select several frequently used LLMs in MedRAG, including the commercial GPT-3.5 and GPT-4, the open-source Mixtral and Llama2, and the biomedical domain-specific MEDITRON and PMC-LLaMA. Temperatures are set to 0 for deterministic outputs.
| LLM | Size | Context | Open | Domain |
|---|---|---|---|---|
| GPT-4 | N/A | 32,768 | No | General |
| GPT-3.5 | N/A | 16,384 | No | General |
| Mixtral | 8×7B | 32,768 | Yes | General |
| Llama2 | 70B | 4,096 | Yes | General |
| MEDITRON | 70B | 4,096 | Yes | Biomed. |
| PMC-LLaMA | 13B | 2,048 | Yes | Biomed. |
(Context: context length of the LLM; Open: Open-source.)
-
First, install PyTorch suitable for your system's CUDA version by following the official instructions (2.1.1+cu121 in our case).
-
Then, install the remaining requirements using:
pip install -r requirements.txt, -
For GPT-3.5/GPT-4, an OpenAI API key is needed. Replace the placeholder with your key in
src/config.py. -
Git-lfsis required to download and load corpora for the first time. -
Javais requried for using BM25.
from src.medrag import MedRAG
question = "A lesion causing compression of the facial nerve at the stylomastoid foramen will cause ipsilateral"
options = {
"A": "paralysis of the facial muscles.",
"B": "paralysis of the facial muscles and loss of taste.",
"C": "paralysis of the facial muscles, loss of taste and lacrimation.",
"D": "paralysis of the facial muscles, loss of taste, lacrimation and decreased salivation."
}
## CoT Prompting
cot = MedRAG(llm_name="OpenAI/gpt-3.5-turbo-16k", rag=False)
answer, _, _ = cot.answer(question=question, options=options)
## MedRAG
medrag = MedRAG(llm_name="OpenAI/gpt-3.5-turbo-16k", rag=True, retriever_name="MedCPT", corpus_name="Textbooks")
### MedRAG without pre-determined snippets
answer, snippets, scores = medrag.answer(question=question, options=options, k=32) # scores are given by the retrieval system
### MedRAG with pre-determined snippets
snippets = [{'id': 'InternalMed_Harrison_30037', 'title': 'InternalMed_Harrison', 'content': 'On side of lesion Horizontal and vertical nystagmus, vertigo, nausea, vomiting, oscillopsia: Vestibular nerve or nucleus Facial paralysis: Seventh nerve Paralysis of conjugate gaze to side of lesion: Center for conjugate lateral gaze Deafness, tinnitus: Auditory nerve or cochlear nucleus Ataxia: Middle cerebellar peduncle and cerebellar hemisphere Impaired sensation over face: Descending tract and nucleus fifth nerve On side opposite lesion Impaired pain and thermal sense over one-half the body (may include face): Spinothalamic tract Although atheromatous disease rarely narrows the second and third segments of the vertebral artery, this region is subject to dissection, fibromuscular dysplasia, and, rarely, encroachment by osteophytic spurs within the vertebral foramina.', 'contents': 'InternalMed_Harrison. On side of lesion Horizontal and vertical nystagmus, vertigo, nausea, vomiting, oscillopsia: Vestibular nerve or nucleus Facial paralysis: Seventh nerve Paralysis of conjugate gaze to side of lesion: Center for conjugate lateral gaze Deafness, tinnitus: Auditory nerve or cochlear nucleus Ataxia: Middle cerebellar peduncle and cerebellar hemisphere Impaired sensation over face: Descending tract and nucleus fifth nerve On side opposite lesion Impaired pain and thermal sense over one-half the body (may include face): Spinothalamic tract Although atheromatous disease rarely narrows the second and third segments of the vertebral artery, this region is subject to dissection, fibromuscular dysplasia, and, rarely, encroachment by osteophytic spurs within the vertebral foramina.'}]
answer, _, _ = medrag.answer(question=question, options=options, snippets=snippets)
### MedRAG with pre-determined snippet ids
snippets_ids = [{"id": s["id"]} for s in snippets]
answer, snippets, _ = medrag.answer(question=question, options=options, snippets_ids=snippets_ids)We've tested the following LLMs on our MedRAG toolkit:
- OpenAI/gpt-4
- OpenAI/gpt-3.5-turbo
- Google/gemini-1.0-pro
- meta-llama/Meta-Llama-3-70B-Instruct
- meta-llama/Llama-2-70b-chat-hf
- mistralai/Mixtral-8x7B-Instruct-v0.1
- epfl-llm/meditron-70b
- axiong/PMC_LLaMA_13B
@inproceedings{xiong-etal-2024-benchmarking,
title = "Benchmarking Retrieval-Augmented Generation for Medicine",
author = "Xiong, Guangzhi and
Jin, Qiao and
Lu, Zhiyong and
Zhang, Aidong",
editor = "Ku, Lun-Wei and
Martins, Andre and
Srikumar, Vivek",
booktitle = "Findings of the Association for Computational Linguistics ACL 2024",
month = aug,
year = "2024",
address = "Bangkok, Thailand and virtual meeting",
publisher = "Association for Computational Linguistics",
url = "https://aclanthology.org/2024.findings-acl.372",
pages = "6233--6251",
abstract = "While large language models (LLMs) have achieved state-of-the-art performance on a wide range of medical question answering (QA) tasks, they still face challenges with hallucinations and outdated knowledge. Retrieval-augmented generation (RAG) is a promising solution and has been widely adopted. However, a RAG system can involve multiple flexible components, and there is a lack of best practices regarding the optimal RAG setting for various medical purposes. To systematically evaluate such systems, we propose the Medical Information Retrieval-Augmented Generation Evaluation (MIRAGE), a first-of-its-kind benchmark including 7,663 questions from five medical QA datasets. Using MIRAGE, we conducted large-scale experiments with over 1.8 trillion prompt tokens on 41 combinations of different corpora, retrievers, and backbone LLMs through the MedRAG toolkit introduced in this work. Overall, MedRAG improves the accuracy of six different LLMs by up to 18{\%} over chain-of-thought prompting, elevating the performance of GPT-3.5 and Mixtral to GPT-4-level. Our results show that the combination of various medical corpora and retrievers achieves the best performance. In addition, we discovered a log-linear scaling property and the {``}lost-in-the-middle{''} effects in medical RAG. We believe our comprehensive evaluations can serve as practical guidelines for implementing RAG systems for medicine.",
}