A package that construct an interface for filtering wavelet coefficients (using PyWavelets) with statistical methods
First you can import some modules like:
import pywt
import numpy as np
import statsWaveletFilt.filtration as fil
import statsWaveletFilt.signals as sig
import statsWaveletFilt.miscellaneous as miscAnd follow some steps to make your filtrations:
In many test for filtration and/or estimation of error, showed in articles, are supose an known ideal signal, how show the code above:
import matplotlib.pyplot as plt
dimension = 1024
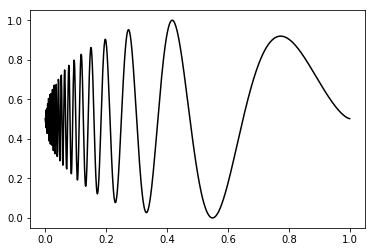
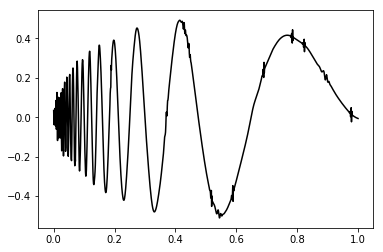
t, idealSignal = sig.dopplerFunction(dimension)
plt.plot(t, idealSignal, 'k-')
plt.show()This package also gives a support for anothers three ideal signal: the heavsine (sig.heavsineFunction()), bump (sig.bumpFunction()) and block (sig.blockFunction()).
And add to this a kind of noise, generally gaussian or, also called, normal.
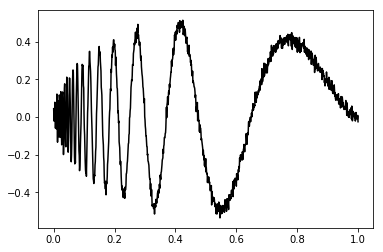
noise = np.random.normal(0, .02, size=dimension)
noisySignal = idealSignal + noise
plt.plot(t, noisySignal, 'k-')
plt.show()Let's do this using the pywt.wavedec function. With this you can 1) make a wavelet transform of your noisy signal, 2) choice wich wavelet function is used ('haar', 'db', 'gaus', 'mexh' among others) and 3) choice any levels you want to evaluate the transform.
noisyCoeff = pywt.wavedec(noisySignal, 'db8', level=5)The first position = noiseCoeff[0] has the scale coefficents of the last nível. The consecutive elements = noiseCoeff[1:] has the wavelet coefficients of the last to the first level of transform.
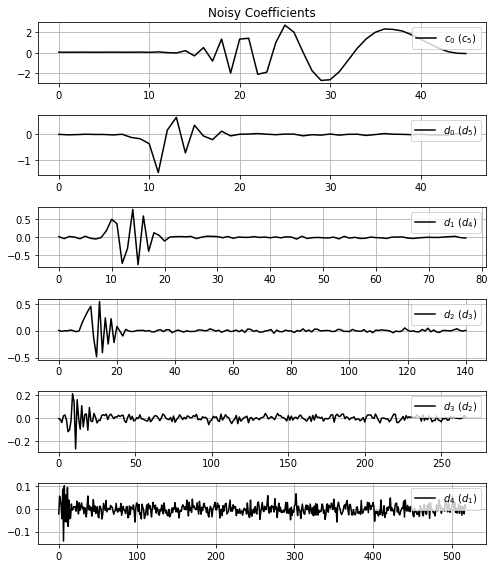
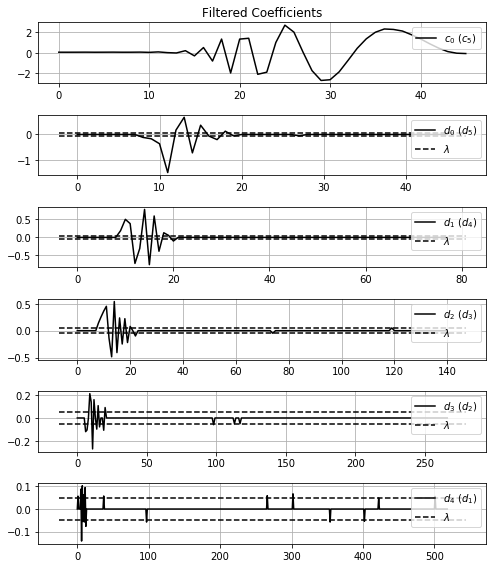
Using misc.showWaveletCoeff function you can show all coefficients returned of the transform.
misc.showWaveletCoeff(noisyCoeff, title='Noisy Coefficients')With the fil.filtration function you can put yours 1) coefficients for filtration, 2) choice the method ('visu', 'sure', 'bayes', 'spc') and, if necessary, 3) set method own parameters (more details in documentation).
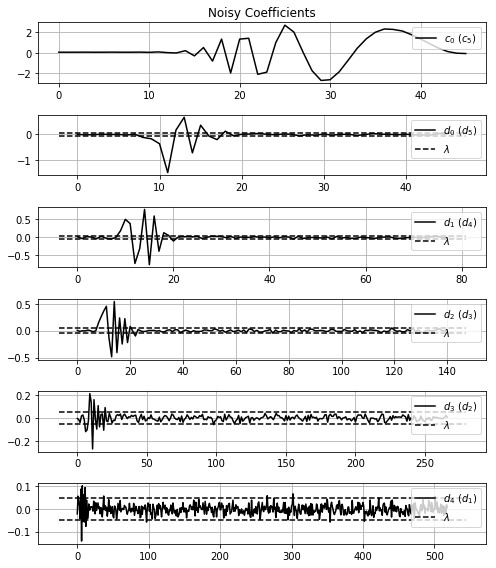
filtrateCoeff, limiars = fil.filtration(noisyCoeff, method='visu')And, again, you can show the wavelet coefficients before and after the filtration, with the limiar used using the misc.showWaveletCoeff function.
misc.showWaveletCoeff(noisyCoeff, threshold_value=limiars, title='Noisy Coefficients')
print('==============================================================')
misc.showWaveletCoeff(filtrateCoeff, threshold_value=limiars, title='Filtered Coefficients')Using the pywt.waverec function we can revovery the signal.
filtratedSignal = pywt.waverec(filtrateCoeff, 'db8')
plt.plot(t, filtratedSignal, 'k-')
plt.show()Using sig.differential_snr_dB we can avaliate the filtration using some figures of merith of different forms of evaluate the snr or cnr (see more in Documentation) of the signal.
sig.differential_snr_dB(noisySignal, filtratedSignal, method='variances', idealSignal=idealSignal)- The
misc.generateDatafunction generate data using the signals showed here with different variances of noise and save in format.npy. - The
misc.normalizeDatafunction normalize a data between a min and max value, this function is used in all signals modeled in sig module.
- DONOHO, JOHNSTONE (1992), Ideal Spatial Adaptation by Wavelet Shrinkage
- DONOHO, JOHNSTONE (1994), Adapting to Unknown Smoothness via Wavelet Shrinkage
- CHANG, YU, VETTERLI (2000),Adaptive Wavelet Thresholding for Image Denoisingand Compression
- BAYER, KOZAKEVICIUS (2010), SPC-Threshold: Uma Proposta de Limiarização para Filtragem Adaptativa de Sinais
- KOZAKEVICIUS, BAYER (2014), Filtragem de sinais via limiarização de coeficientes wavelet