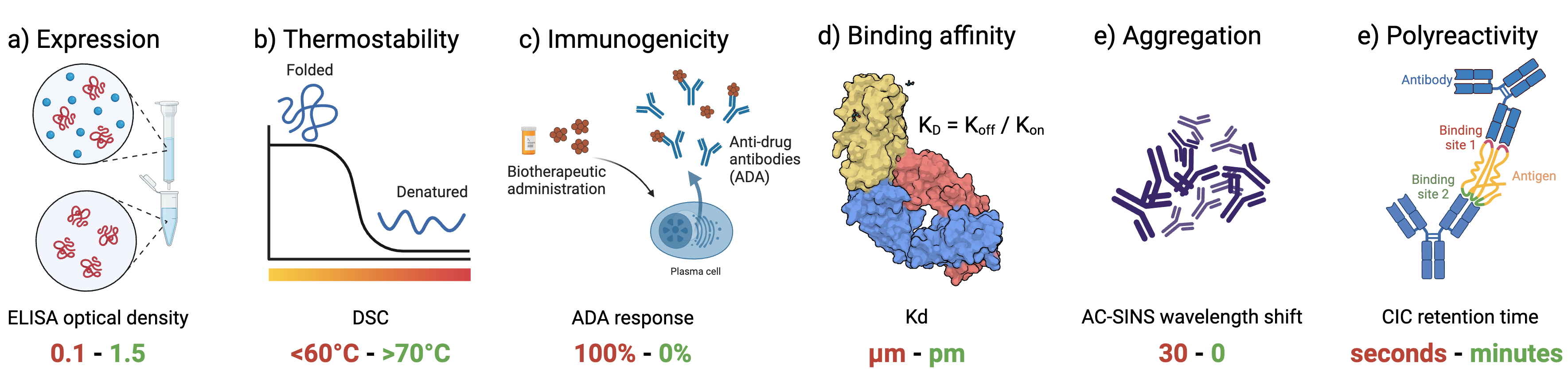
Official repository for FLAb: Benchmarking deep learning methods for antibody fitness prediction. FLAb provides experimental data for six properties of therapeutic antibodies: Expression, themrostability, immunogenicity, aggregation, polyreactivity, and binding affinity. We use FLAb to assess the performance of several widely used deep learning models (AntiBERTy, IgLM, ProtGPT2, ProGen2, ProteinMPNN, ESM-IF) and compare them to physics-based Rosetta.
For easiest use, create a conda environment for each scoring and structure prediction method:
$ conda env create --name ENV_NAME --file envs/[ENV] Where [ENV] ∈ antiberty.yml, esmif.yml, iglm.yml, mpnn.yml, progen.yml, pyrosetta.yml
FLAb supports structure prediction with IgFold and perplexity scoring with AntiBERTy, ProGen2, IgLM, ESM-2, ESM-IF, proteinMPNN, and Rosetta energy.
Antibody sequences must be provided as a csv of sequences, where the columns are chains heavy and light and column values are the sequences. This step is necessary to complete first before scoring with structure-based methods (i.e. ESM-IF, proteinMPNN, Rosetta energy).
$ sbatch sbatch/structure.sh data/tm/Hie2022_C143_Tm.csv After the script completes, antibody structures will be saved in a new directory path structures/tm/Hie2022_C143_Tm/
Calculate perplexity for a csv of sequences with the columns heavy for heavy chain sequences, light for light chain sequences, and fitness for some experimental antibody fitness metric.
$ sbatch sbatch/score_seq.sh data/tm/Hie2022_C143_Tm.csv [MODEL] [SIZE] Where [MODEL] ∈ antiberty, esmif, iglm, mpnn, progen, pyrosetta
If using progen: [SIZE] ∈ small, medium, base, oas, large, BFD90, xlarge. Otherwise leave [SIZE] blank.
For structure-based scoring methods, structures must first be predicted.
$ sbatch sbatch/score_struc.sh data/tm/Hie2022_C143_Tm.csv esmifAfter the script completes, the CSV with heavy and light sequences will be updated with a new column for uncertainty. The CSV will be saved in a new directory path within scores/tm/Hie2022_C143_Tm/
FLAb is a living benchmark: We are motivated to continually expand the antibody fitness data utilized and methods evaluated. We invite contributions and encourage contributors to add data or test new models (e.g. ESM-2, CDConv, ProNet, MaSIF, MIF, CARP, ProtBERT, UniRep, ProteinBERT).
If you run into any problems while using FLAb, please create a Github issue with a description of the problem and the steps to reproduce it.
@article{chungyoun2023flab,
title = {FLAb: Benchmarking tasks in fitness landscape inference for antibodies},
author = {Chungyoun, Michael and Ruffolo, Jeff and Gray, Jeffrey J},
journal = {submitted to NeurIPS datasets and benchmarking track},
year = {2023}
}