Slitflow is a Python package that aims to construct a fully reproducible and universally accessible workflow for single-molecule analysis—namely, Single-molecule Localization-Integrated Trajectory analysis workFLOW.
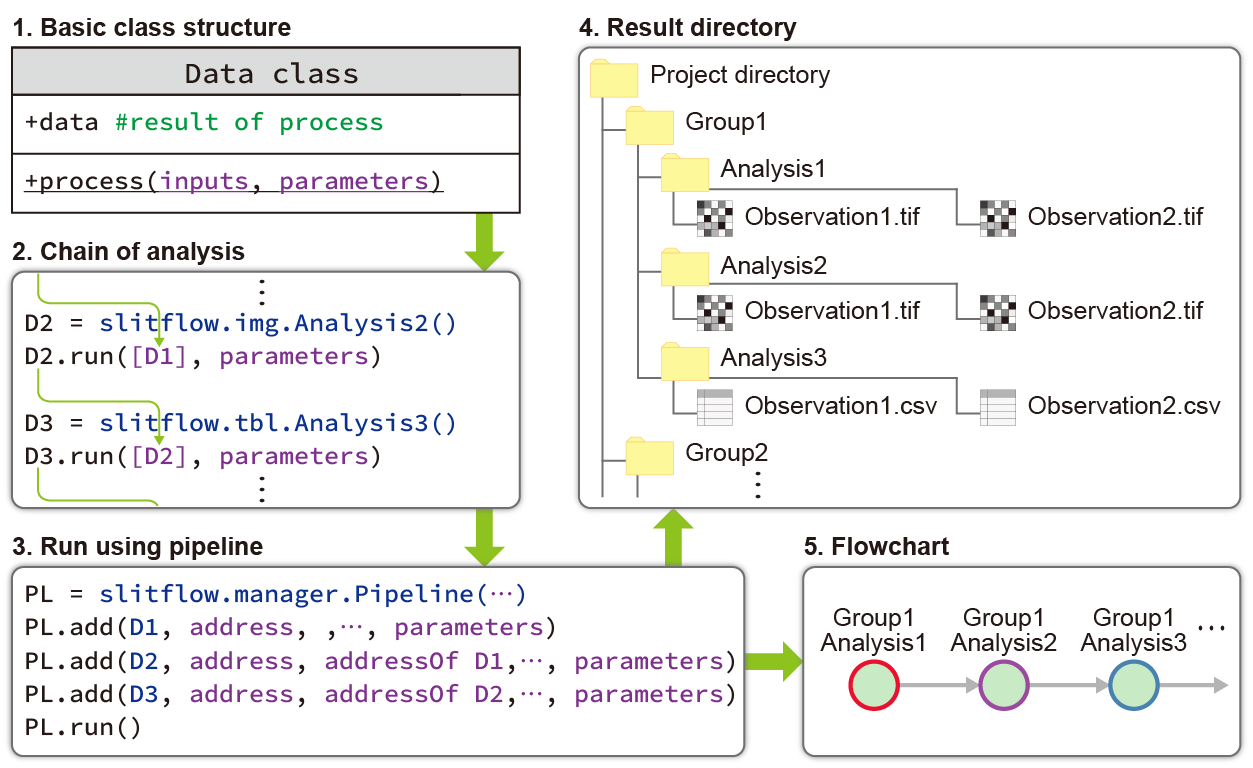
Slitflow comprises a flexible Data class that executes a task and stores the resulting data. A Data object can be input to the next Data object, the network of Data objects forming the entire workflow of complex single-molecule analysis, from image pre-processing to publication-quality figure creation.
Slitflow was designed by considering users who developed analysis tools, validated multiple analysis methods, reproduced workflows without programming skills, and used the results without installing software.
Please see documentation for more information about Slitflow.
Slitflow can be installed from PyPI.
pip install slitflowThe simplest script to make an index table is as follows:
import slitflow as sf
D = sf.tbl.create.Index()
D.run([], {"type": "trajectory", "index_counts": [2, 2], "split_depth": 0})
print(D.data[0])
# img_no trj_no
# 1 1
# 1 2
# 2 1
# 2 2Please see "Getting Started Basic" to overview the functionality by analyzing the trajectories of simulated random walks.
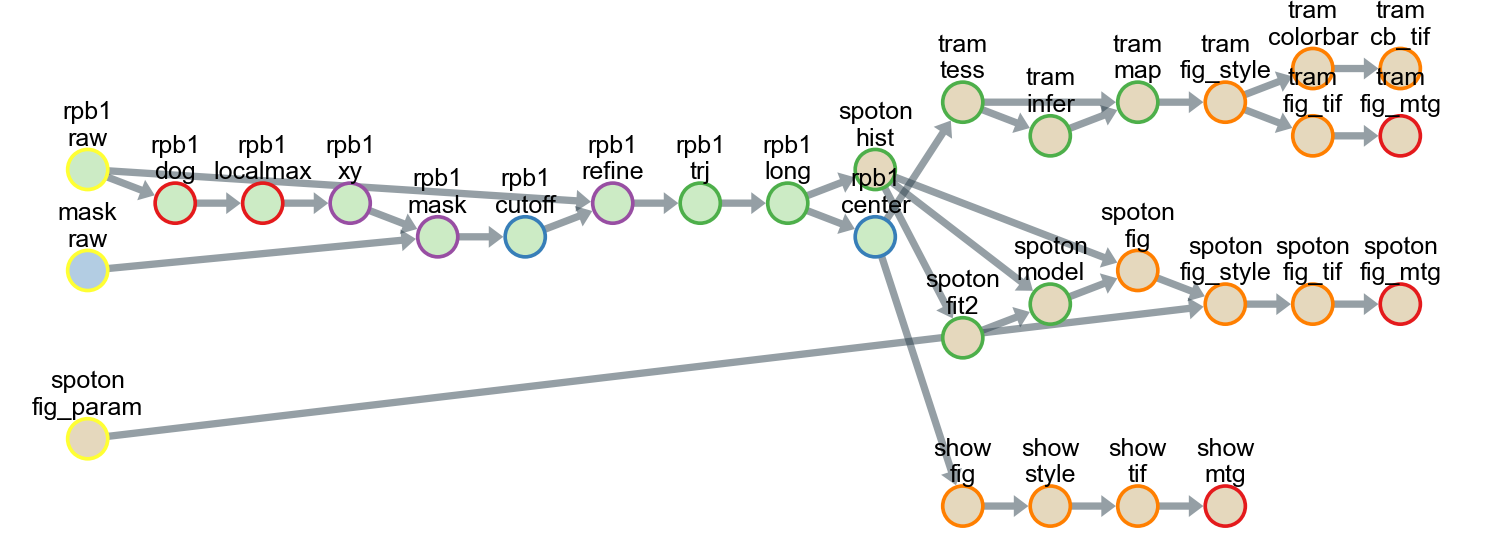
"Getting Started Advanced" demonstrated the integrity and flexibility of the workflow using 1) a cherry-picked tracking method combining popular tools and 2) various state-of-the-art analyses using single-molecule movies.
If Slitflow was useful for your research, please consider citing the following our paper:
- Ito, Y., Hirose, M., and Tokunaga, M. (2023). Slitflow: A Python framework for single-molecule dynamics and localization analysis. SoftwareX 23, 101462. 10.1016/j.softx.2023.101462
Slitflow welcomes any contributions such as bug reports, bug fixes, enhancements, and documentation improvements from interested individuals and groups. Please see documentation. All contributors to this project are expected to abide by our code of conduct.
Slitflow is distributed under the BSD 3-Clause License.