This repo contains pre-trained Meta-matching models. If you want to train your own meta-matching model from scratch, please visit our CBIG repo.
- He, T., An, L., Chen, P., Chen, J., Feng, J., Bzdok, D., Holmes, A.J., Eickhoff, S.B. and Yeo, B.T., 2022. Meta-matching as a simple framework to translate phenotypic predictive models from big to small data, Nature Neuroscience 25, 795-804.
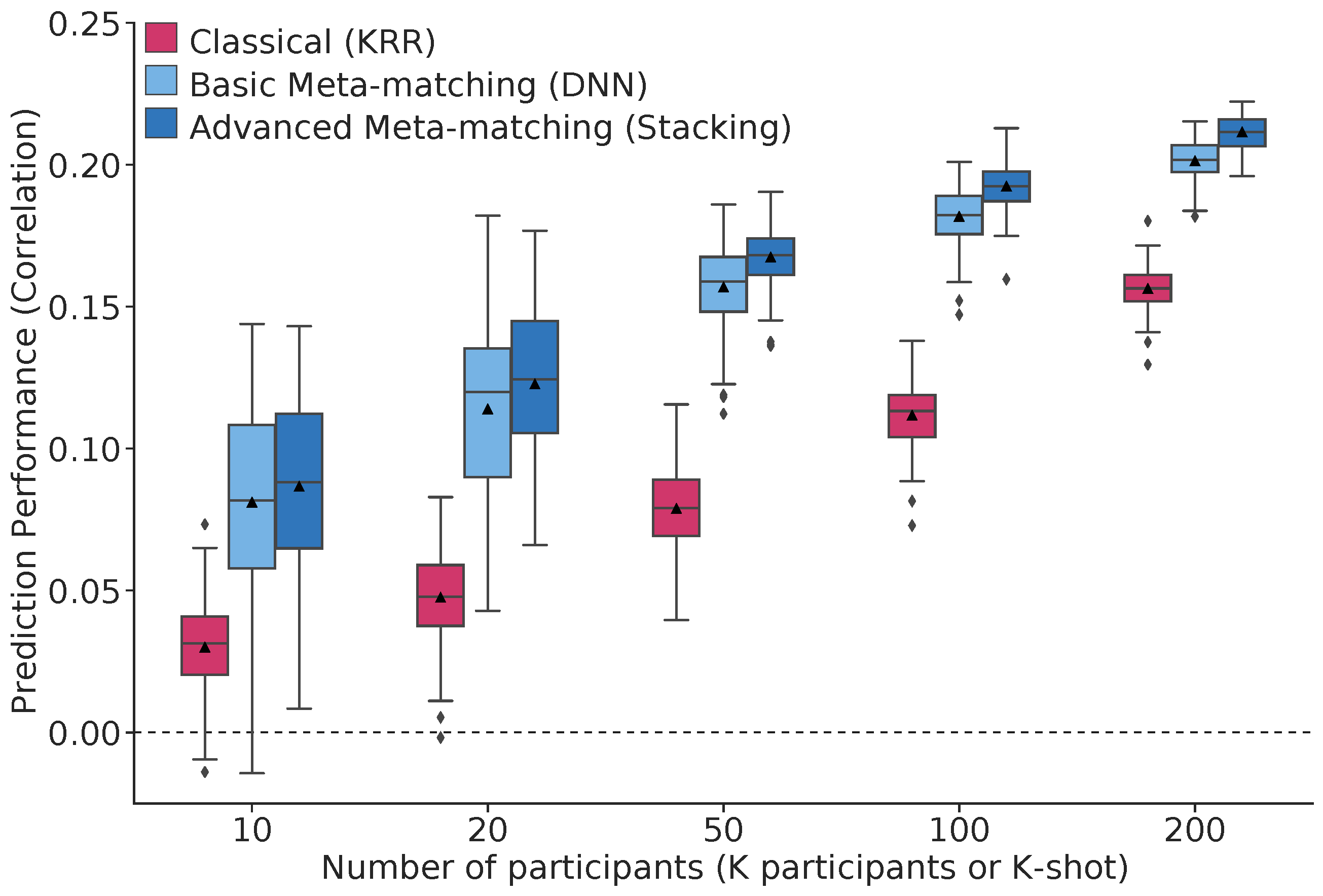
There is significant interest in using brain imaging to predict phenotypes, such as cognitive performance or clinical outcomes. However, most prediction studies are underpowered. We propose a simple framework – meta-matching – to translate predictive models from large-scale datasets to new unseen non-brain-imaging phenotypes in small-scale studies. The key consideration is that a unique phenotype from a boutique study likely correlates with (but is not the same as) related phenotypes in some large-scale dataset. Meta-matching exploits these correlations to boost prediction in the boutique study. We apply meta-matching to predict non-brain-imaging phenotypes from resting-state functional connectivity. Using the UK Biobank (N=36,848) and HCP (N=1,019) datasets, we demonstrate that meta-matching can greatly boost the prediction of new phenotypes in small independent datasets in many scenarios. For example, translating a UK Biobank model to 100 HCP participants yields an 8-fold improvement in variance explained with an average absolute gain of 4.0% (min=-0.2%, max=16.0%) across 35 phenotypes.
Please check the detailed readme under each folder.
- It contains first release of the meta-matching model.
See our LICENSE file for license rights and limitations (MIT).
Please contact He Tong at hetong1115@gmail.com, Lijun An at anlijun.cn@gmail.com, Pansheng Chen at chenpansheng@gmail.com and Thomas Yeo at yeoyeo02@gmail.com.
Happy researching!