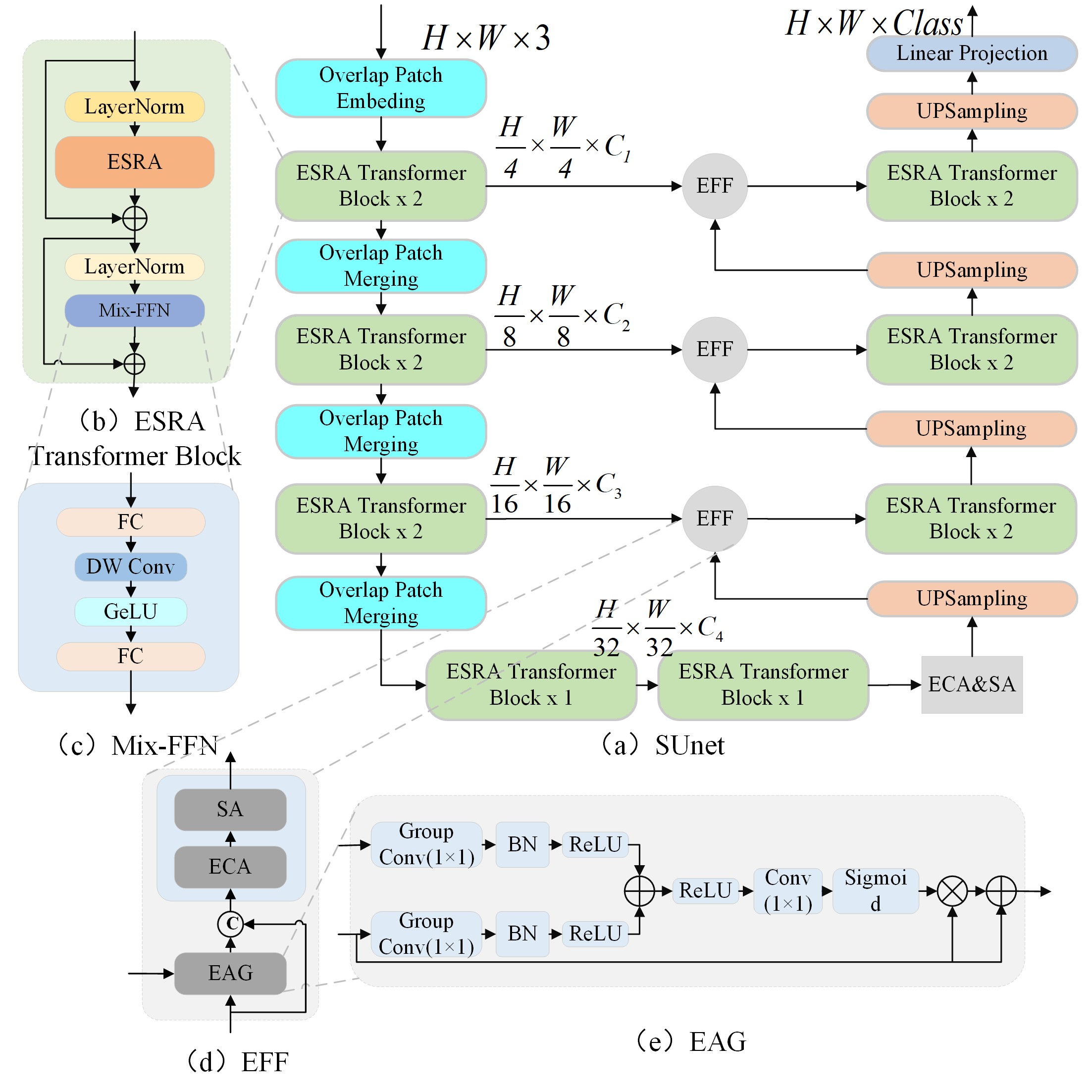
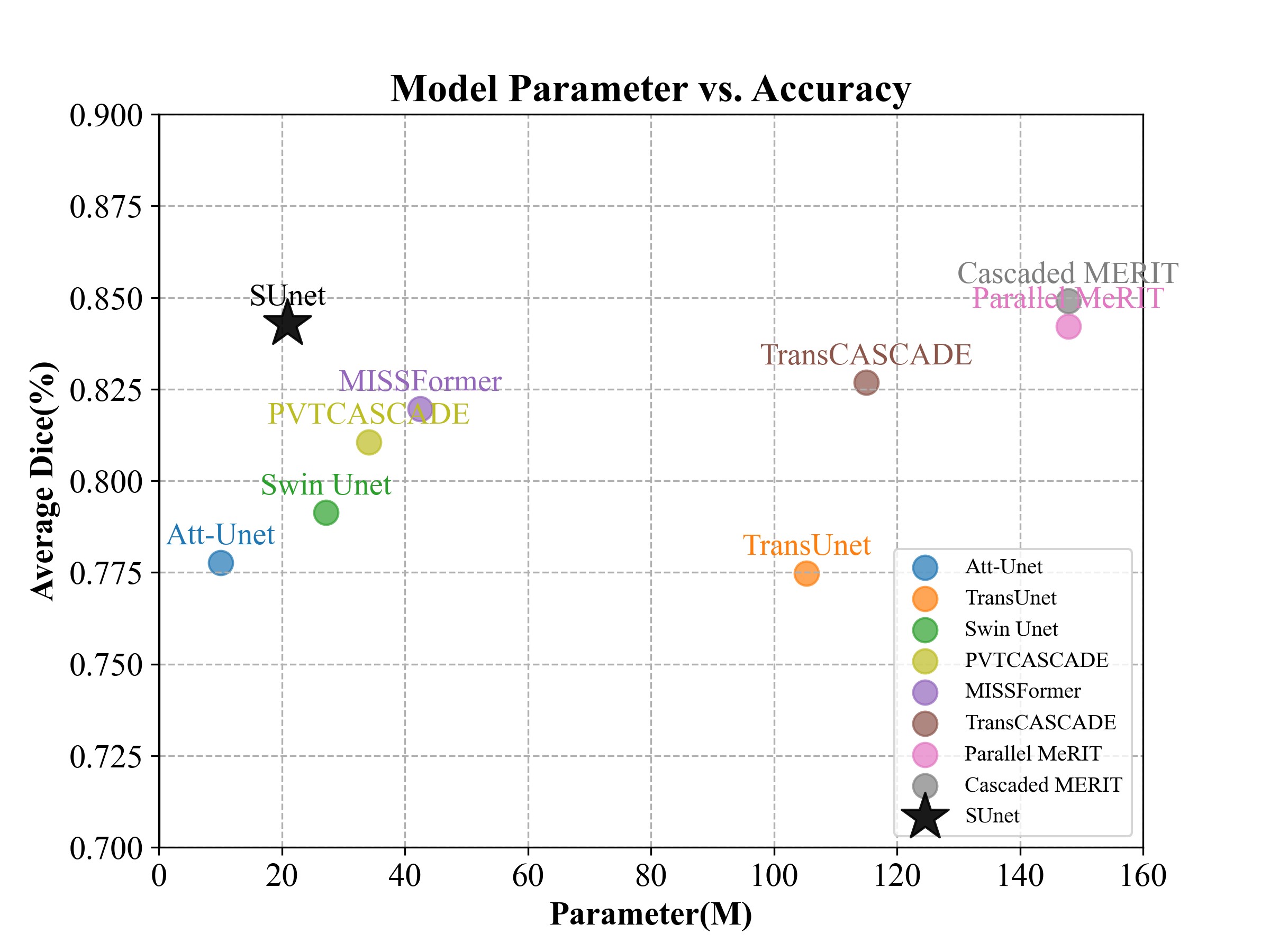
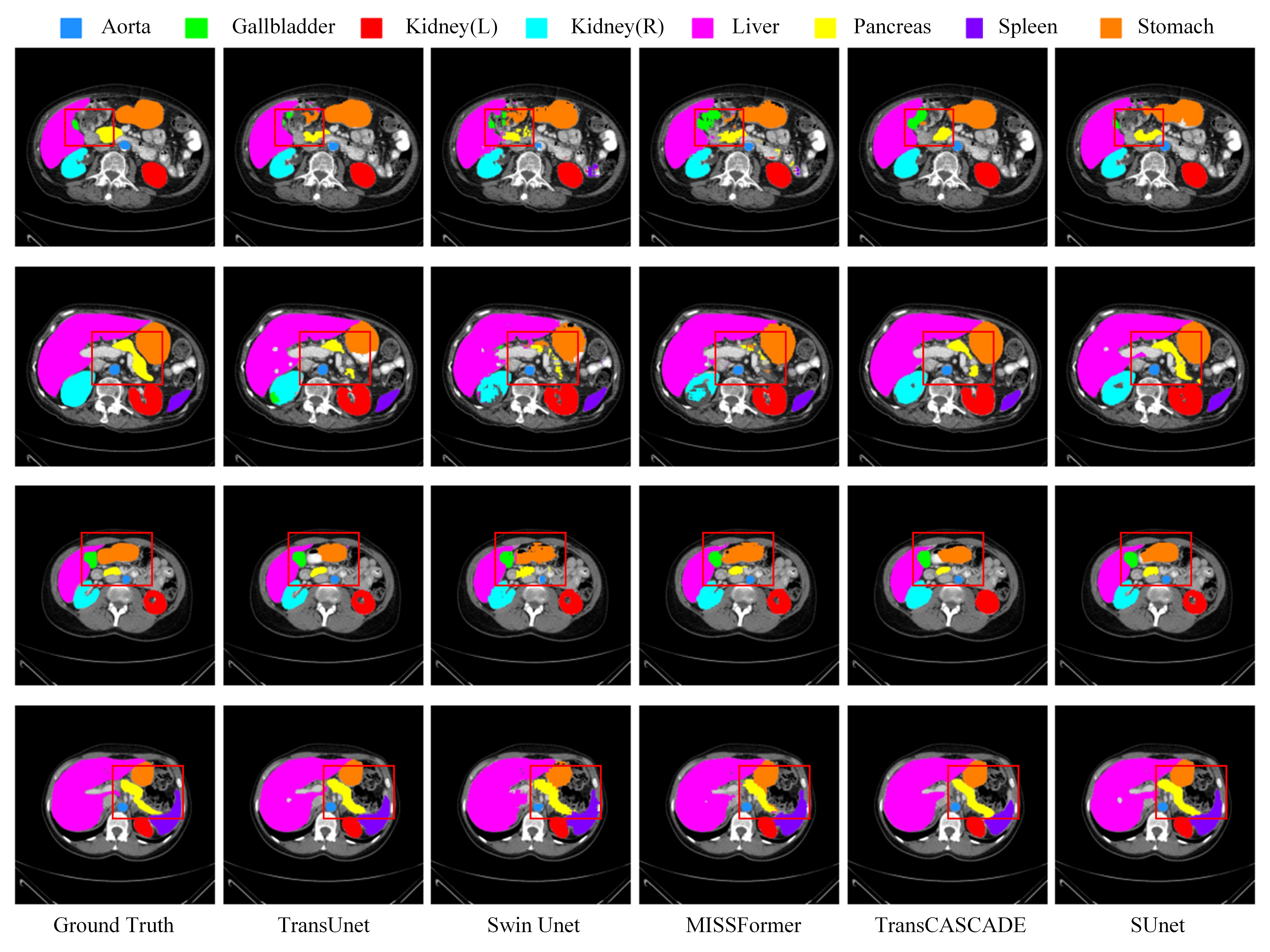
The official implementation of "SUnet: A multi-organ segmentation network based on multiple attention"
Python 3.8 + Pytorch 1.11.0 + torchvision 0.12.0 conda create -n 'your environment name' python==3.8
conda activite 'your environment name'
conda install pytorch==1.11.0 torchvision==0.12.0 torchaudio==0.11.0 cudatoolkit=11.3 -c pytorchAnd please use pip install -r requirements.txt to install the dependencies.
Sign up in the official Synapse website and download the dataset. Then split the 'RawData' folder into 'TrainSet' (18 scans) and 'TestSet' (12 scans) following the TransUNet's lists and put in the './data/synapse/Abdomen/RawData/' folder. Finally, preprocess using python ./utils/preprocess_synapse_data.py or download the preprocessed data and save in the './data/synapse/' folder. Note: If you use the preprocessed data from TransUNet, please make necessary changes (i.e., remove the code segment (line# 88-94) to convert groundtruth labels from 14 to 9 classes) in the utils/dataset_synapse.py.
Download the preprocessed ACDC dataset from Google Drive of MT-UNet and move into './data/ACDC/' folder.
You should download the pretrained pvt_v2_b1 models from Google Drive, and then put it in the './pretrained_pth/pvt/' folder for initialization.
git clone https://github.com/XSforAI/SUnet.git
cd SUnet python train.py python train_ACDC.pyWarnning: Change the parameters when you run the code. For example, root_path
We are very grateful for these excellent works timm, MERIT, CASCADE, Polyp-PVT and TransUNet, which have provided the basis for our framework.
@article{li2023sunet,
title={SUnet: A multi-organ segmentation network based on multiple attention},
author={Li, Xiaosen and Qin, Xiao and Huang, Chengliang and Lu, Yuer and Cheng, Jinyan and Wang, Liansheng and Liu, Ou and Shuai, Jianwei and Yuan, Chang-an},
journal={Computers in Biology and Medicine},
volume={167},
pages={107596},
year={2023},
publisher={Elsevier}
}