My attempt to reproduce graph classification results from recent papers [1, 2] using Graph U-Net. So far, my results using Graph U-Net are worse than the baseline (GCN). I also compare to a recent work on Multigraph GCN (MGCN) [4].
This repository contains all necessary data for the PROTEINS dataset. It can be found here along with similar datasets.
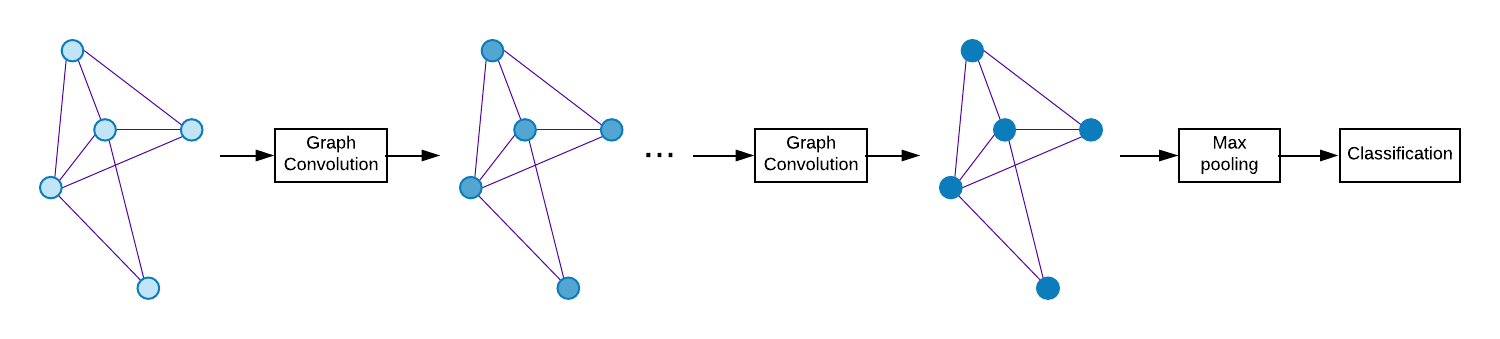
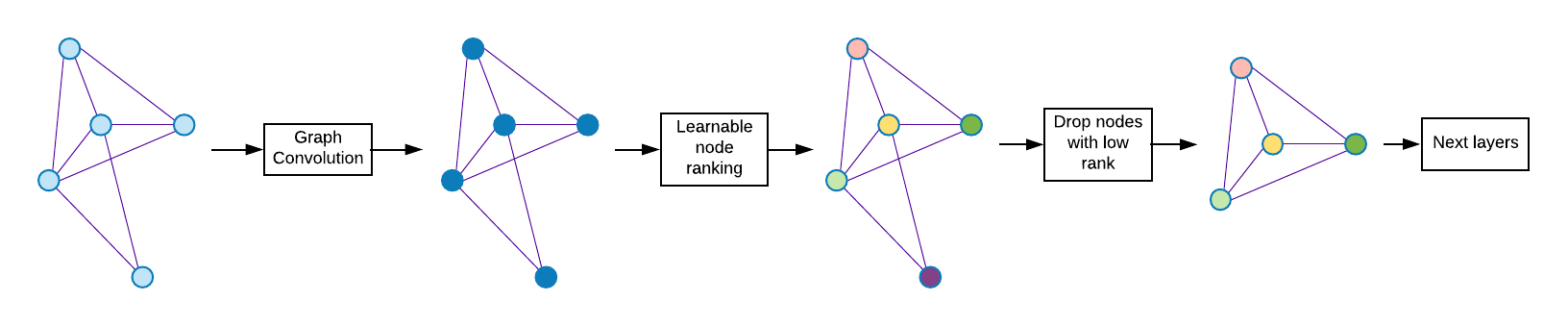
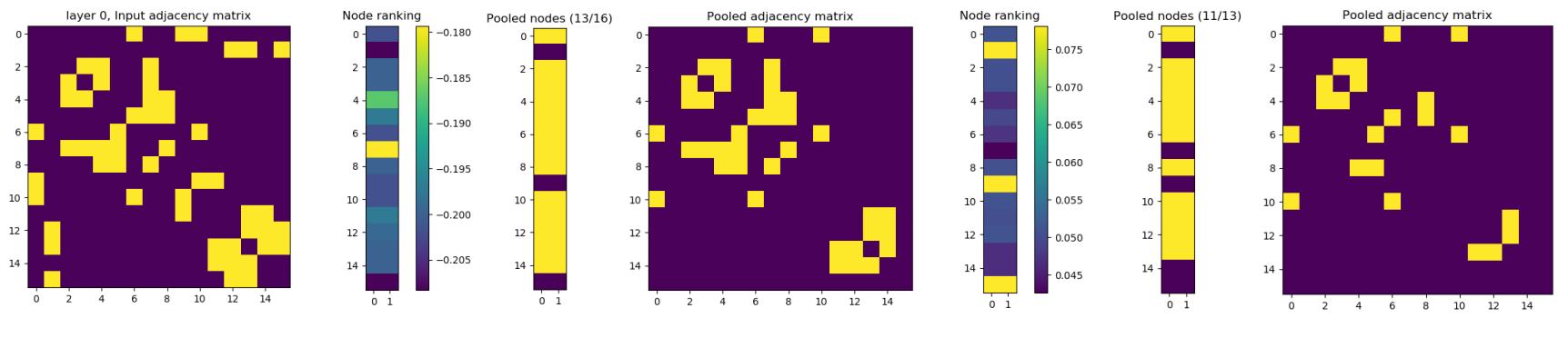
The baseline model is Graph Convolutional Network (GCN) [3]. The decoder part of Graph U-Net is not implemented yet in our code, i.e. the only difference with the baseline is using pooling based on dropping nodes between graph convolution layers.
Hyperparameters are taken from [2], but learning rate decay and dropout is also applied. The readout layer (last pooling layer over nodes) is also simplified to just max pooling over nodes.
All hyperparameters are the same for the baseline, Graph U-Net and Multigraph GCN (MGCN).
Implementation is very basic without much optimization, so that it is easier to debug and play around with the code.
python graph_unet.py --model gcn # to run baseline GCN
python graph_unet.py --model unet # to run Graph U-Net
python graph_unet.py --model mgcn # to run Multigraph GCNTo use the PyTorch Geometric data loader, add flag --torch-geom.
Repeating 10 times for different seeds:
for i in $(seq 1 10); do seed=$(( ( RANDOM % 10000 ) + 1 )); python graph_unet.py --model gcn --seed $seed | tee logs/gcn_proteins_"$i".log; doneThen reading log files can be done as following:
results_dir = './logs'
acc = []
for f in os.listdir(results_dir):
with open(pjoin(results_dir, f), 'r') as fp:
s = fp.readlines()[-1]
pos1 = s.find(':')
acc.append(float(s[pos1+1:s[pos1:].find('(') + pos1]))
print(len(acc), np.mean(acc), np.std(acc))Average and std of accuracy for 10-fold cross-validation. We also repeat experiments 10 times (as shown above) for different random seeds and report average and std over those 10 times.
| Model | PROTEINS | PROTEINS (10 times) |
|---|---|---|
| GCN [3] | 74.71 ± 3.44* | 74.37 ± 0.31 |
| GCN [3] + A2 | 74.36 ± 4.57 | 74.56 ± 0.26 |
| GCN [3] + A2 + 2I | 74.45 ± 4.91 | 74.23 ± 0.37 |
| Graph U-Net [1, 2] | 72.39 ± 3.34 | 72.45 ± 0.88 |
| Graph U-Net [1, 2] + A2 | 72.90 ± 4.08 | 72.87 ± 0.52 |
| Graph U-Net [1, 2] + A2 + 2I | 73.63 ± 4.67 | 73.18 ± 0.50 |
| Multigraph GCN (MGCN) [4] | 74.62 ± 2.56 | 75.56 ± 0.27 |
*74.72 ± 2.90 with PyTorch 1.0.0.
Some datasets contain additional float-valued node attributes, which can improve graph classification a lot. Note that some algorithms, including Weisfeiler-Lehman (WL) Graph Kernels, are not able to make use of these additional attributes, so algorithms should be compared fairly.
| Model | ENZYMES | ENZYMES + continuous node attributes |
|---|---|---|
| GCN [3] | 32.33 ± 5.071 | 51.17 ± 5.632 |
| Graph U-Net [1, 2] | 33.00 ± 4.88 | 48.33 ± 6.32 |
| Multigraph GCN (MGCN) [4] | 40.50 ± 5.58 | 59.83 ± 6.56 |
These results were obtained by running (similarly for Graph U-Net):
1
python graph_unet.py -D ENZYMES -f 128,128,128 --n_hidden 256 -g --lr 0.0005 --epochs 100 --lr_decay_step 150
2
python graph_unet.py -D ENZYMES -f 128,128,128 --n_hidden 256 -gc --lr 0.0005 --epochs 100 --lr_decay_step 150
Here, lr_decay_step can be any number larger than the number of epochs to avoid learning rate decay.
The code is tested on Ubuntu 16.04 with PyTorch 0.4.1/1.0.0 and Python 3.6.
The jupyter notebook file is kept for debugging purposes.
Optionally: