Citation | Installation | Example | Usage | Command line | API doc | Interface classes | Website | Acknowledgment
An IPython/Jupyter widget to interactively view molecular structures and trajectories. Utilizes the embeddable NGL Viewer for rendering. Support for showing data from the file-system, RCSB PDB, simpletraj and from objects of analysis libraries mdtraj, pytraj, mdanalysis, ParmEd, rdkit, ase, HTMD, biopython, cctbx, pyrosetta, schrodinger's Structure
Should work with Python 3. If you experience problems, please file an issue.
Ask question about usage? Please post here
- Installation
- Example
- Showcase from users
- Usage
- Contributing
- Command line
- API doc
- Interface classes
- Changelog
- FAQ
- Website
- Acknowledgment
- Cite
- License
-
Available on
conda-forgechannelconda install nglview -c conda-forge # might need: jupyter-nbextension enable nglview --py --sys-prefix # if you already installed nglview, you can `upgrade` conda upgrade nglview --force # might need: jupyter-nbextension enable nglview --py --sys-prefix
-
Available on PyPI
pip install nglview
# might need: jupyter-nbextension enable nglview --py --sys-prefixJupyterlab: nglview works best with jupyterlab >= 3.0 and no further steps needed.
If you are using notebook v5.0, you need to increase the iopub_data_rate_limit
to visualize big structure (e.g: solvated system)
jupyter notebook --NotebookApp.iopub_data_rate_limit=10000000
Requirement: ipywidgets >= 7.0, notebook >= 4.2
The development version can be installed directly from github:
git clone https://github.com/arose/nglview
cd nglview
python setup.py install
# if you edit files in ./js folder, make sure to rebuild the code
cd js
npm install
# probably need to activate widgetsnbextension
# python -m ipykernel install --sys-prefix
# jupyter nbextension enable --py --sys-prefix widgetsnbextension
# jupyter nbextension enable --py --sys-prefix nglview
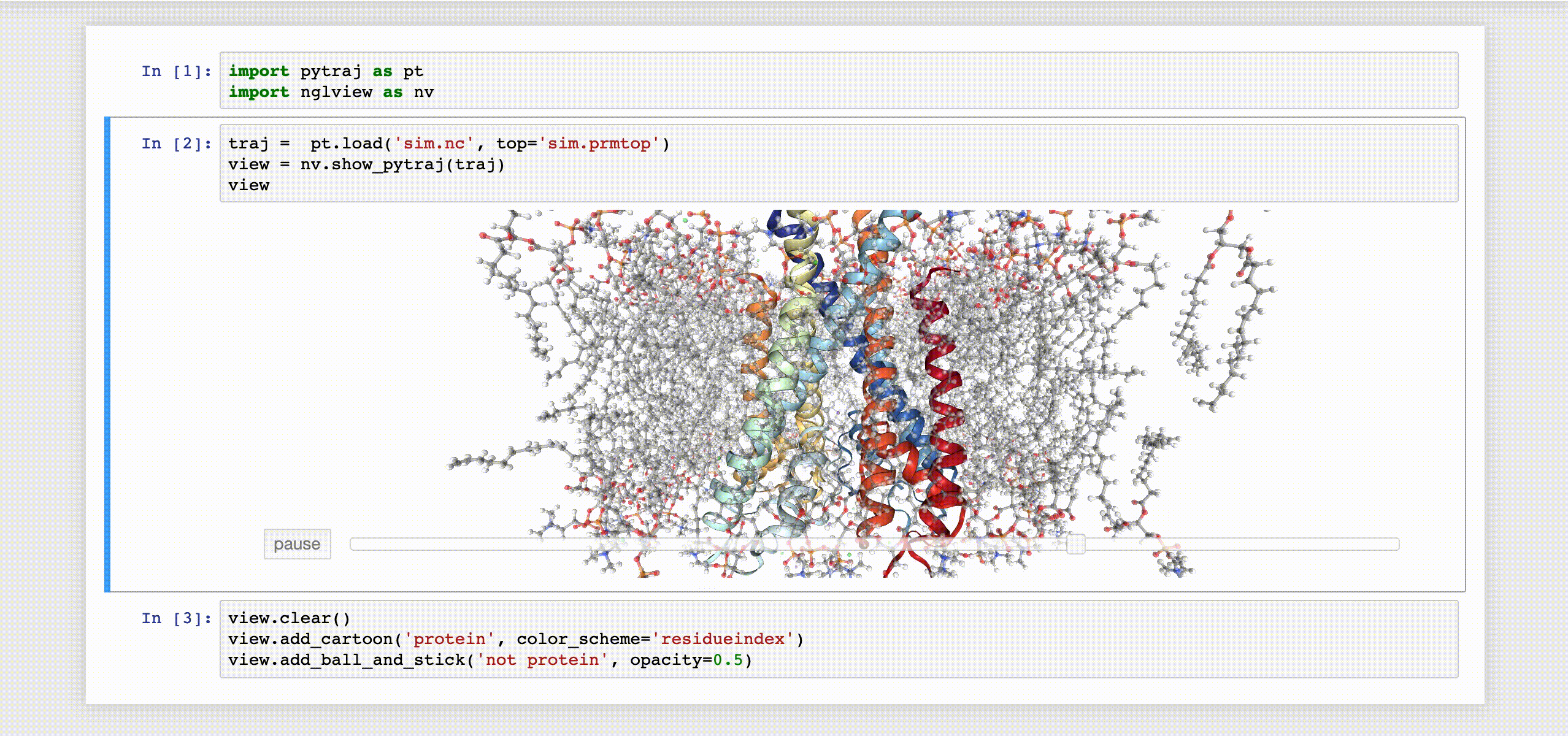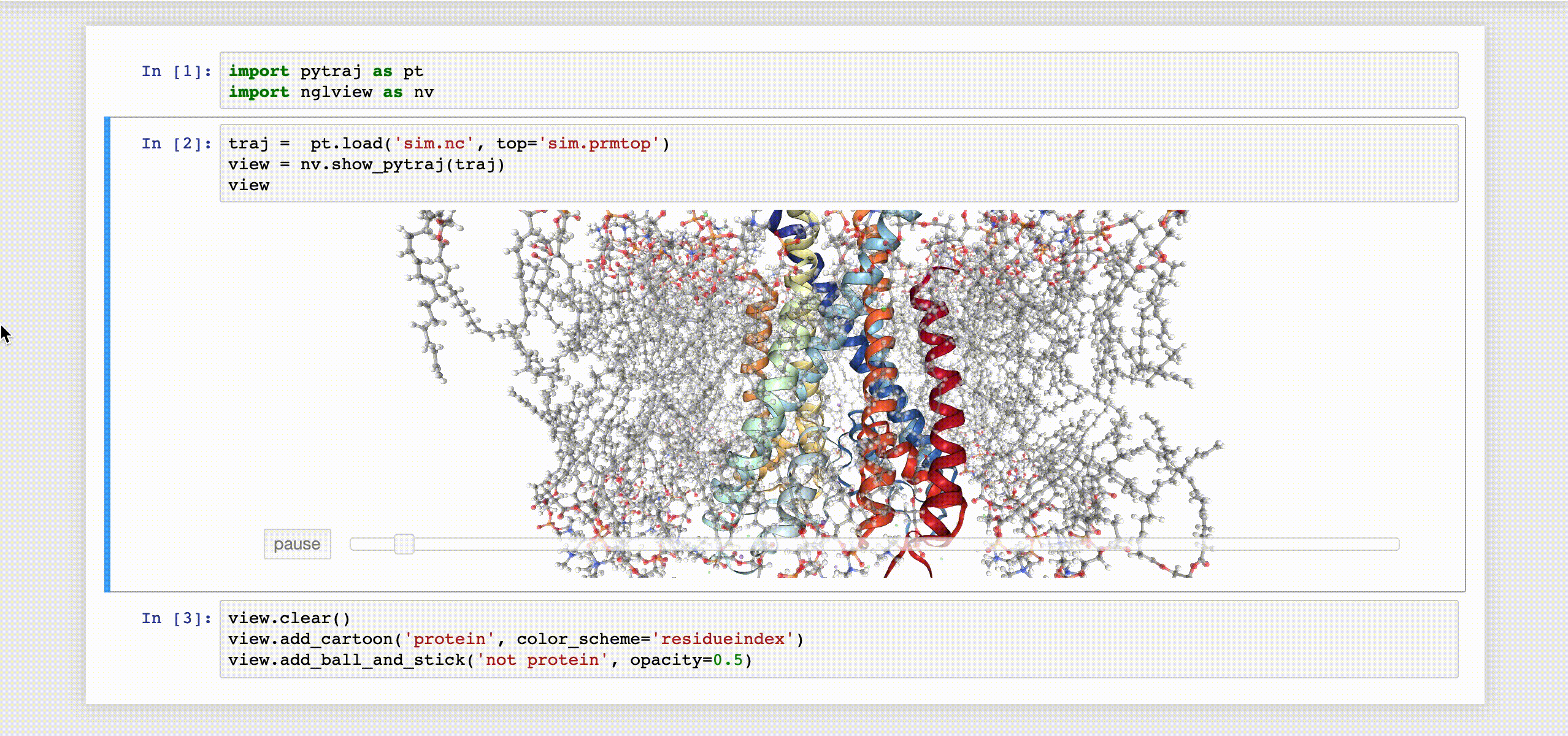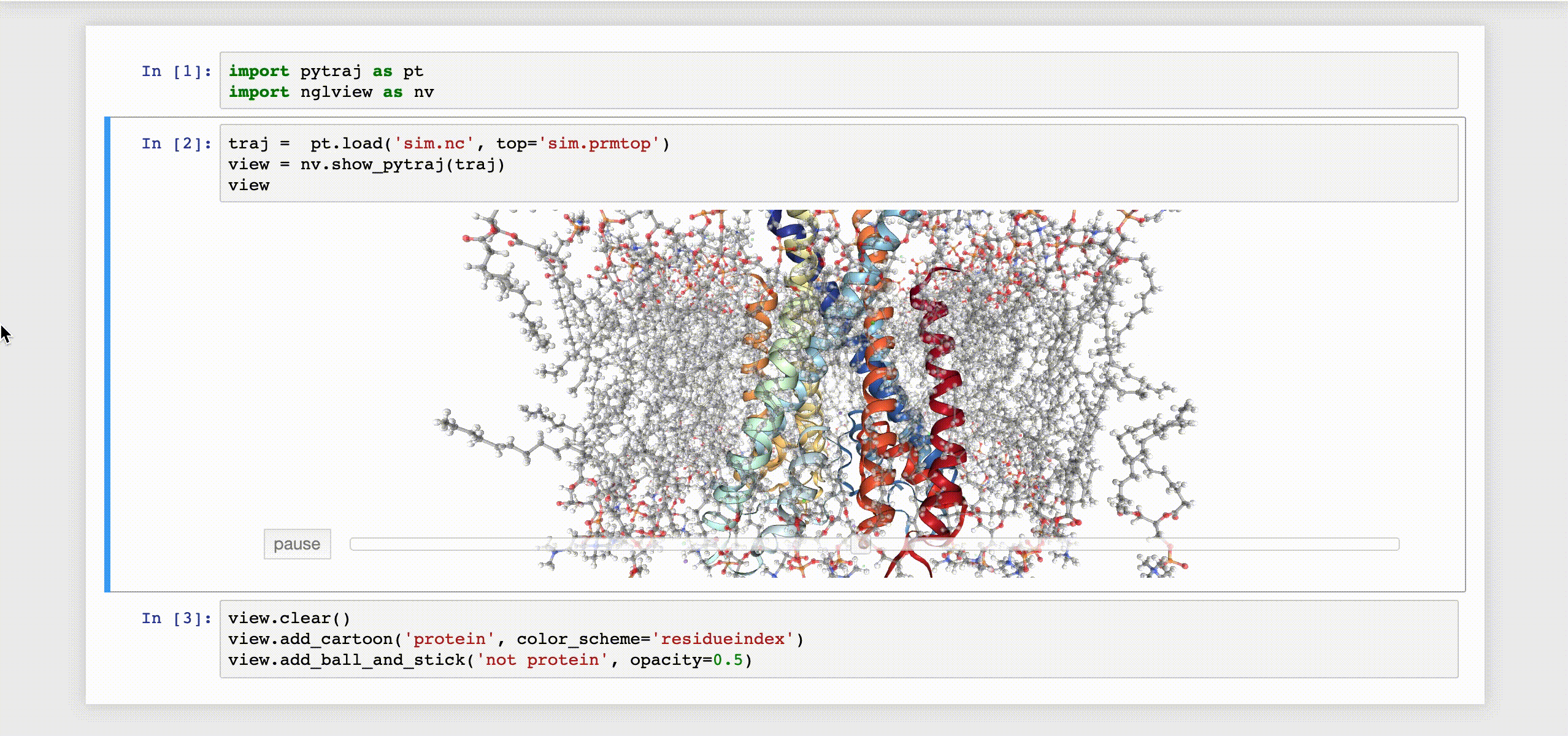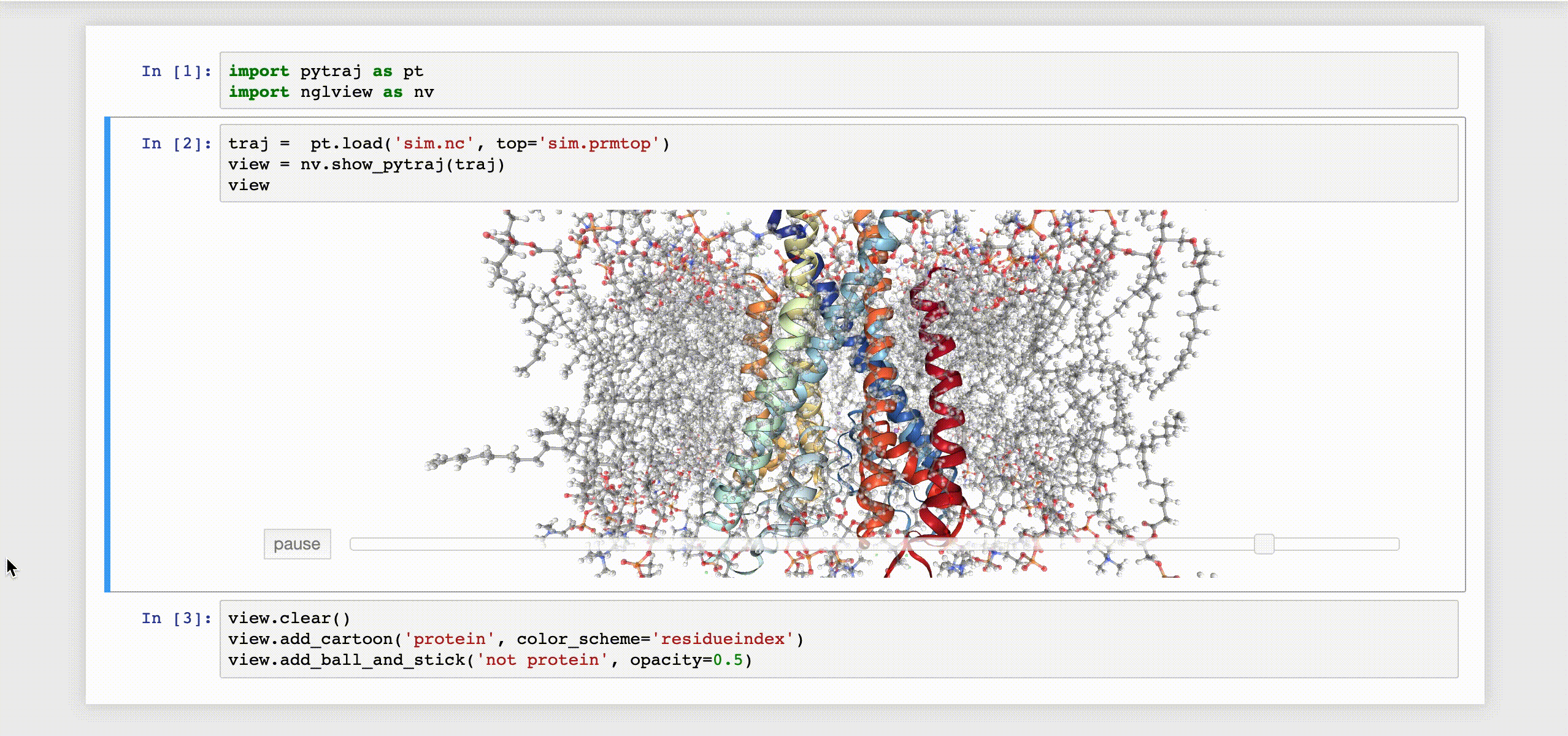
# tested with ipywidgets 5.2.2, notebook 4.2.1- Notebooks: please see our Jupyter notebook examples
- Simple demo for trajectory (take time to load): biomembrane
- Please check user examples. Feel free to contribute.
- Also check a series of excelent tutorials about/using nglview from volkamerlab
Open a notebook
jupyter notebook
and issue
import nglview
view = nglview.show_pdbid("3pqr") # load "3pqr" from RCSB PDB and display viewer widget
viewA number of convenience functions are available to quickly display data from the file-system, RCSB PDB, simpletraj and from objects of analysis libraries mdtraj, pytraj, mdanalysis, ParmEd, rdkit, HTMD, biopython.
| Function | Description |
|---|---|
show_file(path) |
Shows any NGL supported file formats (pdb, gro, mol2, sdf, dx, ..) in path |
show_pdbid(pdbid) |
Shows pdbid fetched from RCSB PDB |
show_simpletraj(struc_path, traj_path) |
Shows structure & trajectory loaded with simpletraj |
show_mdtraj(traj) |
Shows MDTraj trajectory traj |
show_pytraj(traj) |
Shows PyTraj trajectory traj |
show_parmed(structure) |
Shows ParmEd structure |
show_mdanalysis(univ) |
Shows MDAnalysis Universe or AtomGroup univ |
show_rdkit(mol) |
Shows rdkit rdkit.Chem.rdchem.Mol |
show_ase(atoms) |
Shows ase Atoms |
show_asetraj(traj) |
Shows ase trajectory traj |
show_pymatgen(struct) |
Shows pymatgen Structure |
show_htmd(mol) |
Shows HTMD Molecules |
show_biopython(mol) |
Shows Biopython structural entities |
show_iotbx(mol) |
Shows cctbx's iotbx structure |
show_rosetta(pose) |
Shows pyrosetta's Pose |
show_iodata(obj) |
Shows iodata's IOData |
show_psi4(obj) |
Shows psi4's Molecule |
show_qcelemental |
Shows QCelementary's Molecule |
show_openbabel |
Shows openbabel's OMol |
show_prody |
Shows prody's Ensemble or AtomGroup |
view.add_representation('cartoon', selection='protein')
# or shorter
view.add_cartoon(selection="protein")
view.add_surface(selection="protein", opacity=0.3)
# specify color
view.add_cartoon(selection="protein", color='blue')
# specify residue
view.add_licorice('ALA, GLU')
# clear representations
view.clear_representations()
# update parameters for ALL cartoons of component 0 (default)
view.update_cartoon(opacity=0.4, component=0)
# remove ALL cartoons of component 0 (default)
view.remove_cartoon(opacity=0.4, component=0)
# Not using default representation
view = nv.show_file('your.pdb', default=False)
view.center()
view.add_rope()Representations can also be changed by overwriting the representations property
of the widget instance view. The available type and params are described
in the NGL Viewer documentation.
view.representations = [
{"type": "cartoon", "params": {
"sele": "protein", "color": "residueindex"
}},
{"type": "ball+stick", "params": {
"sele": "hetero"
}}
]The widget constructor also accepts a representation argument:
initial_repr = [
{"type": "cartoon", "params": {
"sele": "protein", "color": "sstruc"
}}
]
view = nglview.NGLWidget(struc, representation=initial_repr)
view# set the frame number
view.frame = 100# parameters for the NGL stage object
view.stage.set_parameters(**{
# "percentages, "dist" is distance too camera in Angstrom
"clipNear": 0, "clipFar": 100, "clipDist": 10,
# percentages, start of fog and where on full effect
"fogNear": 0, "fogFar": 100,
# background color
"backgroundColor": "black",
})
# note: NGLView accepts both origin camel NGL keywords (e.g. "clipNear")
# and snake keywords (e.g "clip_near")# parameters to control the `delay` between snapshots
# change `step` to play forward (positive value) or backward (negative value)
# note: experimental code
view.player.parameters = dict(delay=0.04, step=-1)# update camera type
view.camera = 'orthographic'# change background color
view.background = 'black'# adding new trajectory
view.add_trajectory(traj)
# traj could be a `pytraj.Trajectory`, `mdtraj.Trajectory`, `MDAnalysis.Universe`,
# `parmed.Structure`, `htmd.Molecule` or derived class of `nglview.Trajectory`
# change representation
view[0].add_cartoon(...) # equal to view.add_cartoon(component=0)
view[1].add_licorice(...) # equal to view.add_licorice(component=1)# Density volumes (MRC/MAP/CCP4, DX/DXBIN, CUBE)
# Or adding derived class of `nglview.Structure`
view.add_component('my.ccp4')
# add component from url
view.add_component('rcsb://1tsu.pdb')
# NOTE: Trajectory is a special case of component.# coot mouse style (https://en.wikipedia.org/wiki/Coot_(software))
view.stage.set_parameters(mouse_preset='coot')Require: moviepy (pip install moviepy)
from nglview.contrib.movie import MovieMaker
movie = MovieMaker(view, output='my.gif', in_memory=True)
movie.make()# open a notebook and import nglview
nglview
# Require installing pytraj (PR for other backends is welcome)
# open notebook, load `my.pdb` to pytraj's trajectory then display `view`
nglview my.pdb
# load density data
nglview my.ccp4
# open notebook, create trajectory with given topology `my.parm7` and trajecotry file `traj.nc`,
# then display `view`
nglview my.parm7 -c traj.nc
# load all trajectories with filename ending with 'nc'
# make sure to use quote " "
nglview my.parm7 -c "*.nc"
# open notebook, copy content from `myscript.py`
nglview myscript.py
# create a remote notebook
# just follow its instruction
nglview my.pdb --remote
nglview my.parm7 -c traj.nc --remote
nglview mynotebook.ipynb --remote
# demo (don't need pytraj)
nglview demo
# specify web browser
nglview my.pdb --browser=google-chrome(Feel free to make a PR to add/remove your project here. Thanks.)
- AMBER - A package of programs for molecular dynamics simulations of proteins and nucleic acids
- mbuild - A hierarchical, component based molecule builder
- deepchem - Deep-learning models for Drug Discovery and Quantum Chemistry
- htmd - High throughput molecular dynamics simulations
- Moleidoscope - Molecular kaleidoscope
- ssbio - Tools for enabling structural systems biology
- hublib - hublib is a Python library for the HUBzero science gateway platform.
- molPX: ipython API to visualize MD-trajectories along projected trajectories
- nanoribbon
- ase: Atomic Simulation Environment
- pida: Software for analyzing multiple protein-protein interaction docking solutions,
- pytim
- MobleyLab/drug-computing Educational materials for, and related to, UC Irvine's Drug Discovery Computing Techniques course.
- pyiron: an integrated development environment for implementing, testing, and running simulations in computational materials science.
- BioSimSpace: An interoperable framework for biomolecular simulation
- pyrod: PyRod - Tracing water molecules in molecular dynamics simulations
- kugupu: kugupu - a molecular network generator to study charge transport pathways in amorphous materials
- pnab: proto-Nucleic Acid Builder
- opencadd: A Python library for structural cheminformatics
- teachopencadd: TeachOpenCADD: a teaching platform for computer-aided drug design (CADD) using open source packages and data
- query.libretexts.org: query.libretexts.org
- datamol: A python library to work with molecules.
- dynophores: Dynamic pharmacophore modeling of molecular interactions
- pychemcurv: Discrete and local curvature applied to chemistry and chemical reactivity
- AutoSolvate: Automated workflow for generating quantum chemistry calculation of explicitly solvated molecules
- plipify: PLIPify: Protein-Ligand Interaction Frequencies across Multiple Structures
- Melodia: Differential Geometry of Proteins Backbones
- pyrosetta_viewer3d: Display PackedPose objects, Pose objects, or PDB files within a Jupyter notebook and Google Colab
- py4vasp: Python interface for VASP
- eminus: A plane wave density functional theory code.
- MolSysMT: Molecular Systems Multi-Tool
- Funding: Hai Nguyen is supported by NIH Grant GM103297, "The Center for HIV RNA Studies" (2015 to 02-2017).
- Many thanks to
nglviewcontributors - dunovank/jupyter-themes: for
oceans16theme - base64-arraybuffer
- ipywidgets
If you would like to acknowledge our work, feel free to cite:
Hai Nguyen, David A Case, Alexander S Rose; NGLview - Interactive molecular graphics for Jupyter notebooks, Bioinformatics, , btx789, https://doi.org/10.1093/bioinformatics/btx789
Generally MIT, see the LICENSE file for details.