pygraph is a library to work with graphs:
-
Searches: DFS, BFS and Dijkstra algorithms
-
Minimum Spanning Tree: Kruskal, Inverse Kruskal and Prism algorithms
-
Render with graphviz
Creation of random graphs with models:
-
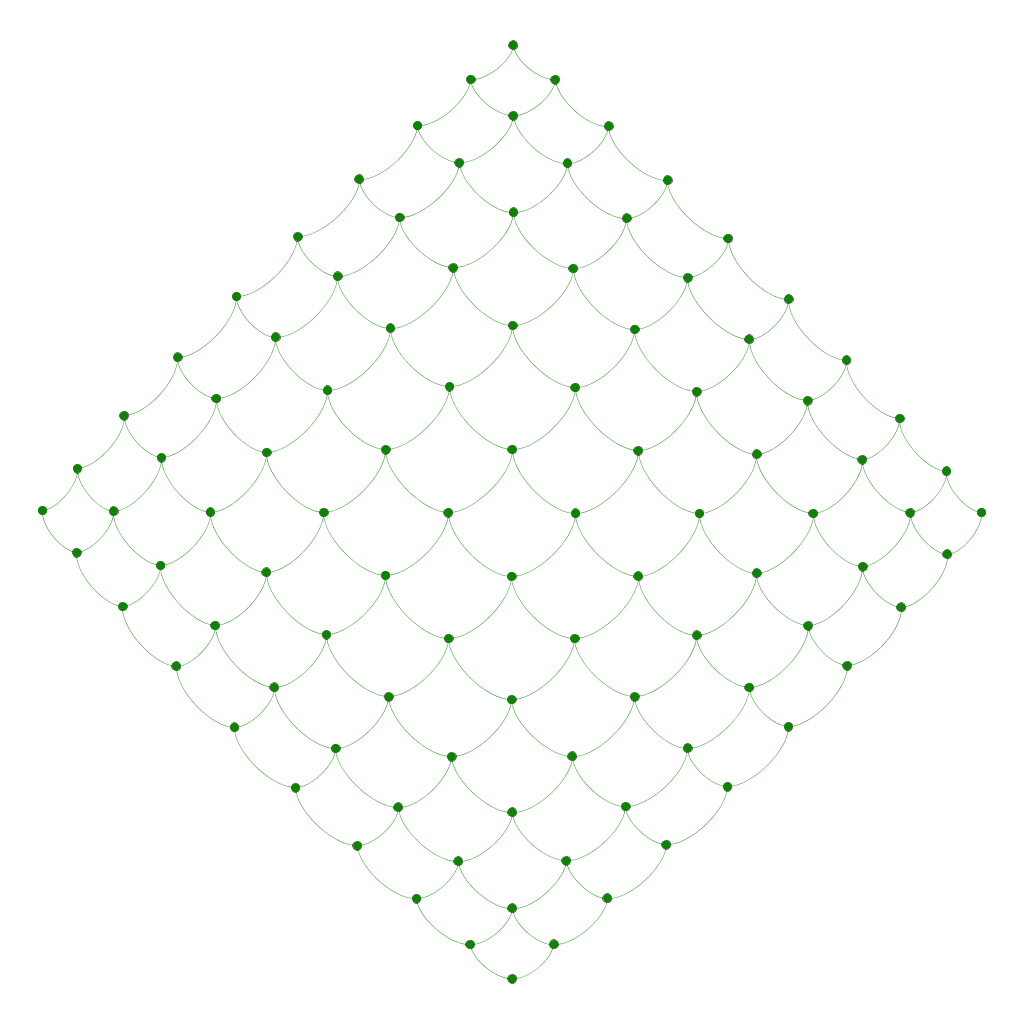
Mesh model
Graph of m*n nodes representing a meshing model.
-
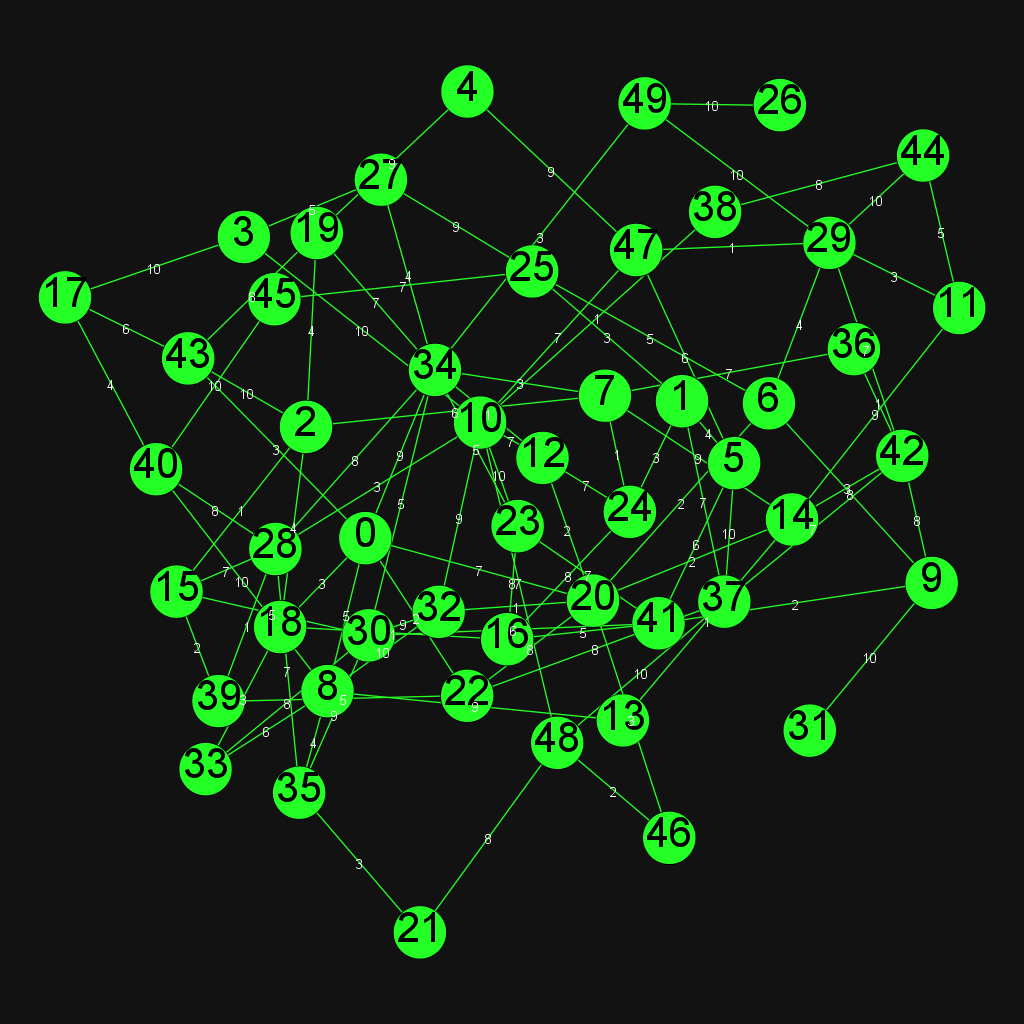
Erdős–Rényi model
In the G(n,p) model, a graph is constructed by connecting labeled nodes randomly. Each edge is included in the graph with probability p, independently from every other edge.
-
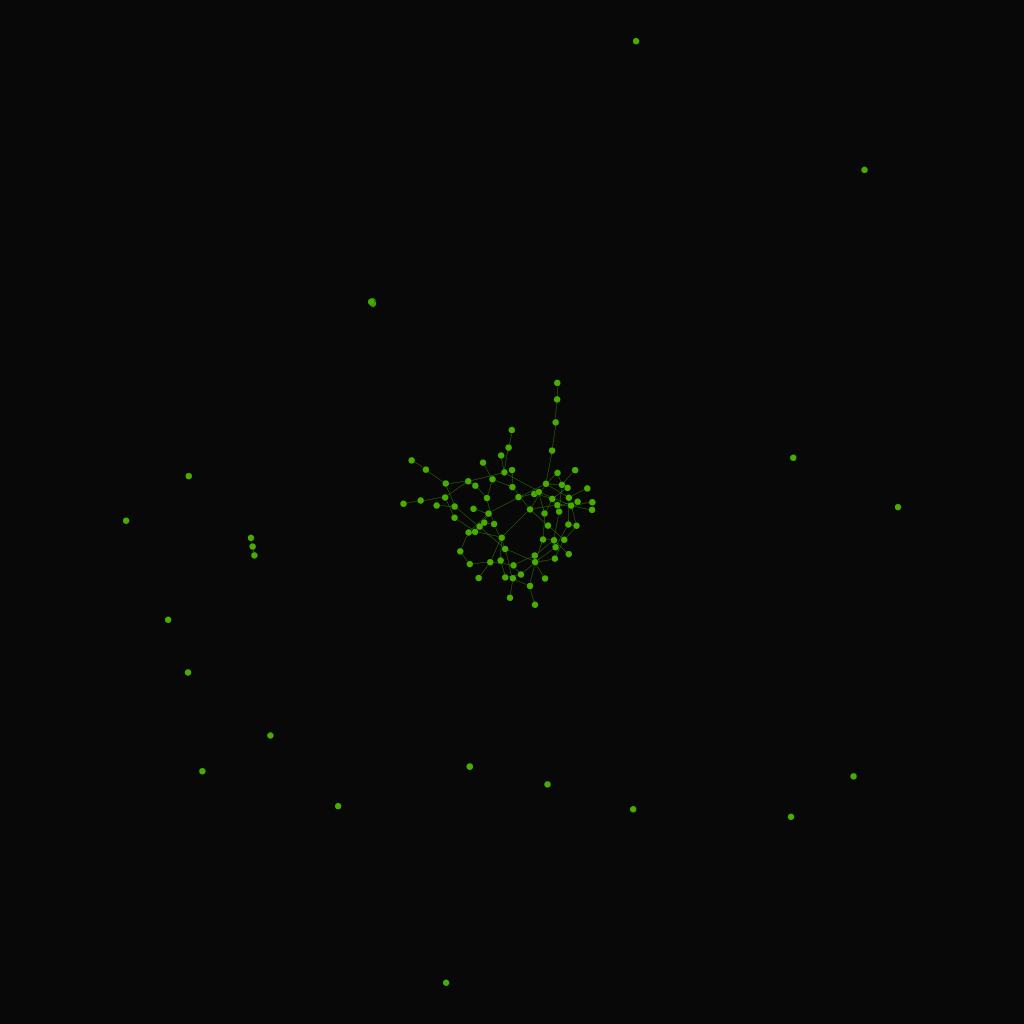
Gilbert model
A random graph obtained by starting with a set of n isolated vertices and adding successive edges between them at random.
-
Geographic model
Create a random graph with simple method geographic.
-
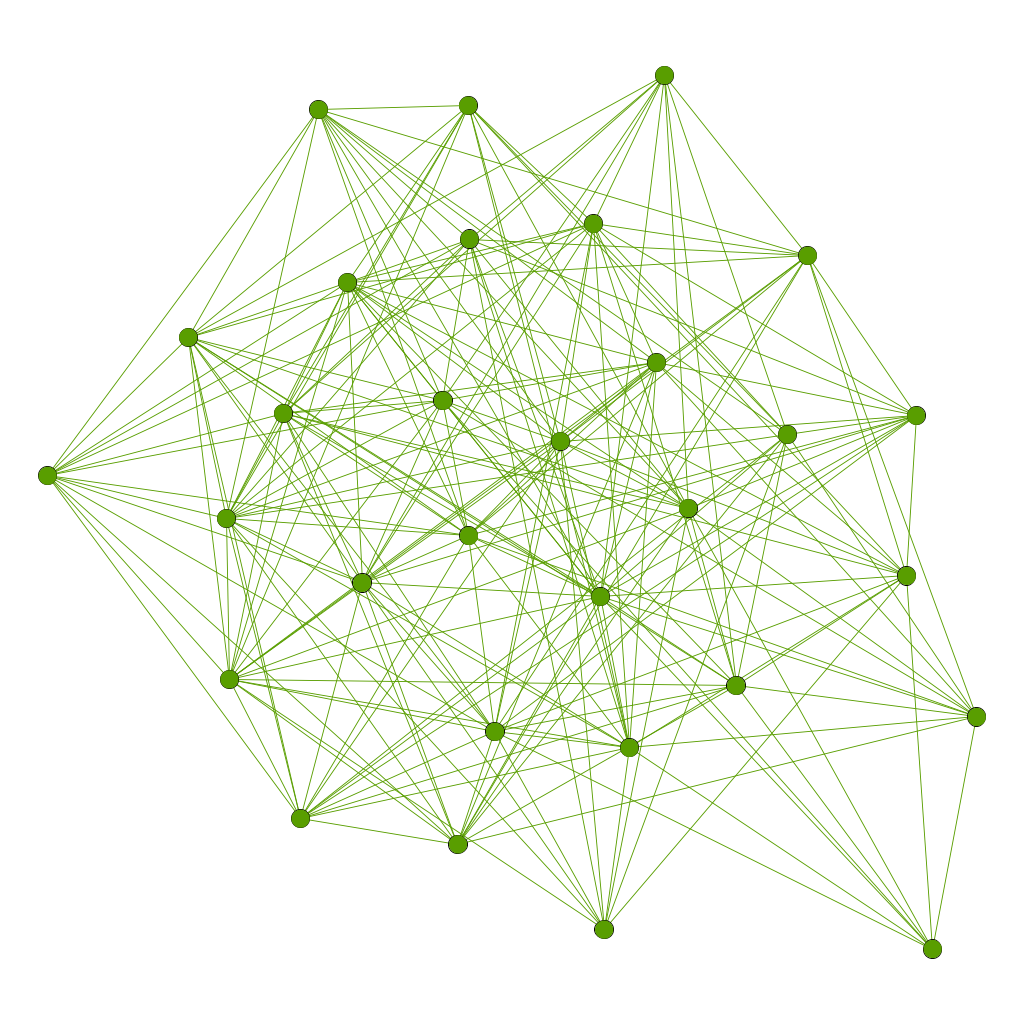
Barabási-Albert
The graph begins with an initial connected network of m0 nodes. New nodes are added to the network one at a time. Each new node is connected to m < m0 existing nodes with a probability that is proportional to the number of links that the existing nodes already have.
-
Dorogovtsev-Mendes
A graph with edge set E, and denote the degree of a vertex v (that is, the number of edges incident to v) by deg(v).
-
Breadth-first search (BFS)
BFS is an algorithm for traversing or searching tree or graph data structures. It starts at the tree root (or some arbitrary node of a graph, sometimes referred to as a 'search key'), and explores all of the neighbor nodes at the present depth prior to moving on to the nodes at the next depth level.
-
DFS Depth-first search (DFS)
DFS is an algorithm for traversing or searching tree or graph data structures. The algorithm starts at the root node (selecting some arbitrary node as the root node in the case of a graph) and explores as far as possible along each branch before backtracking.
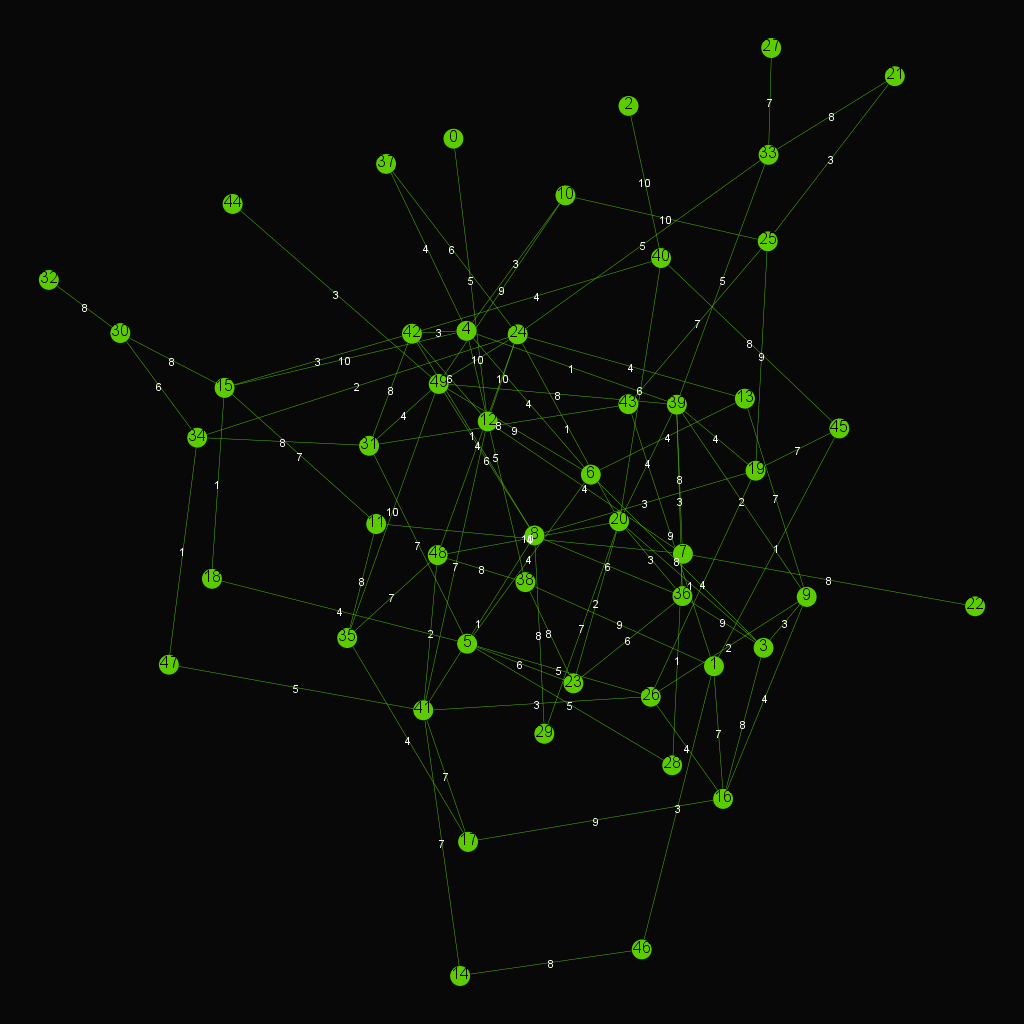
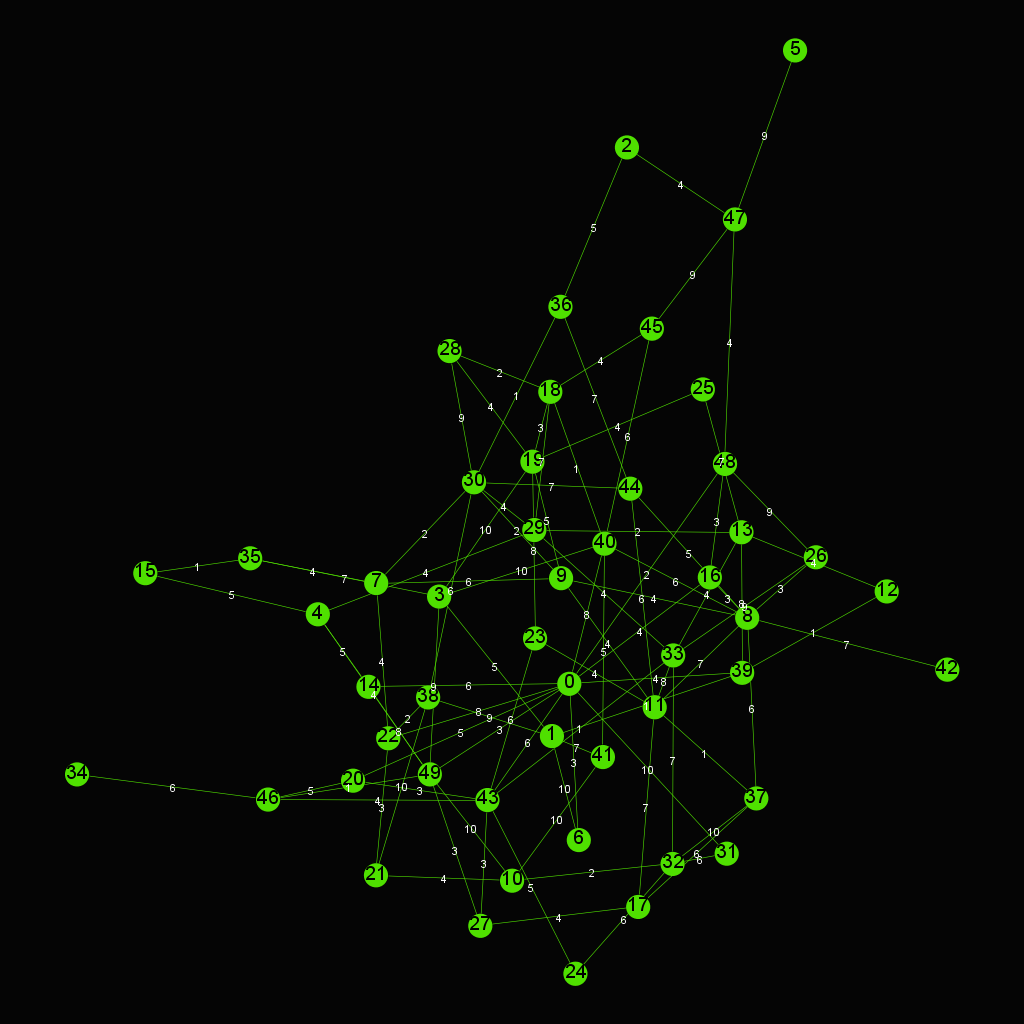
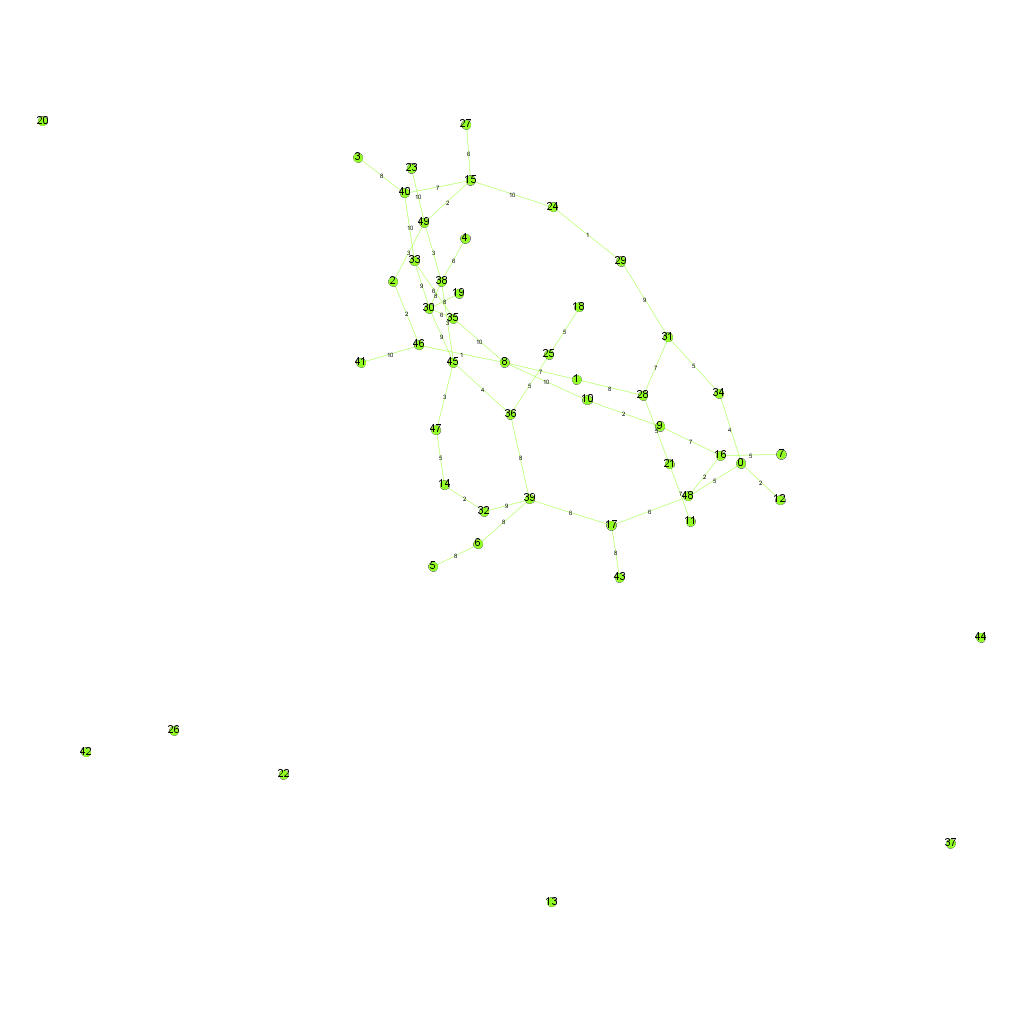
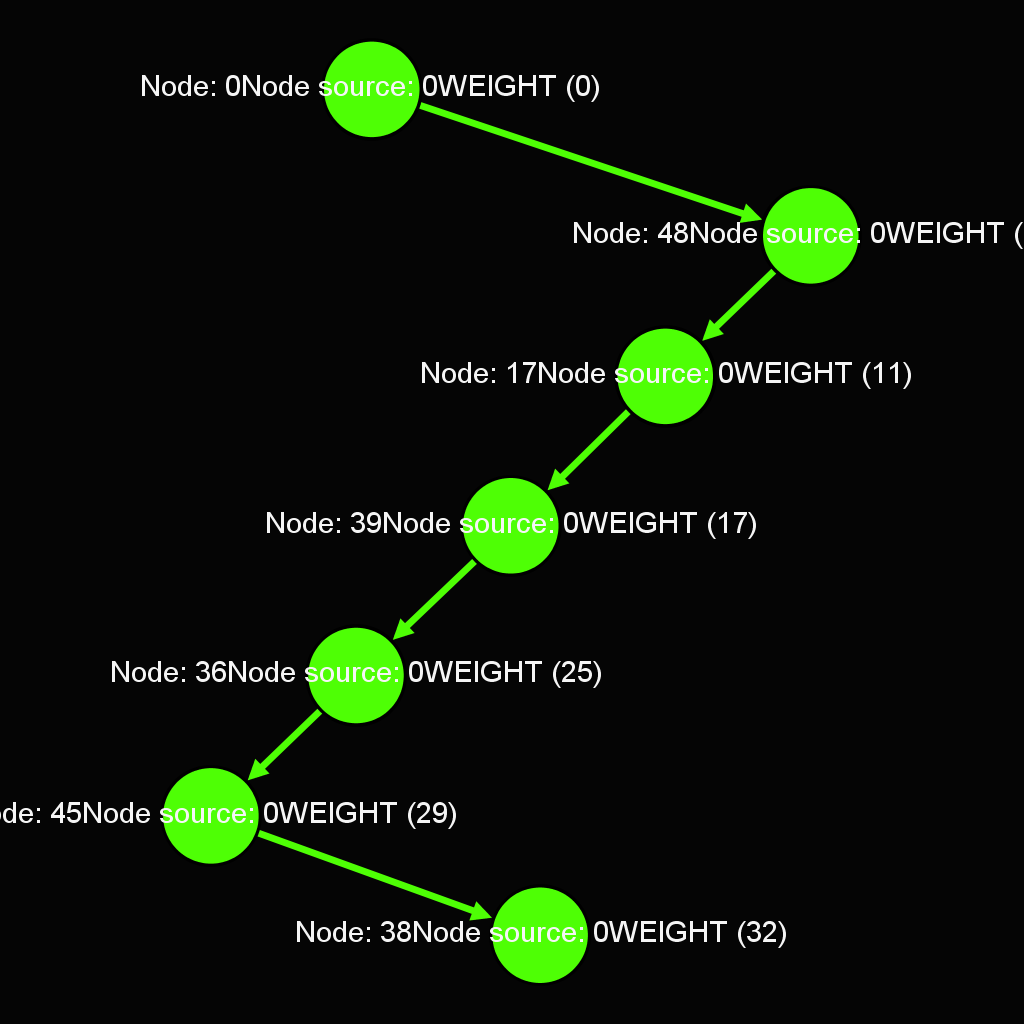
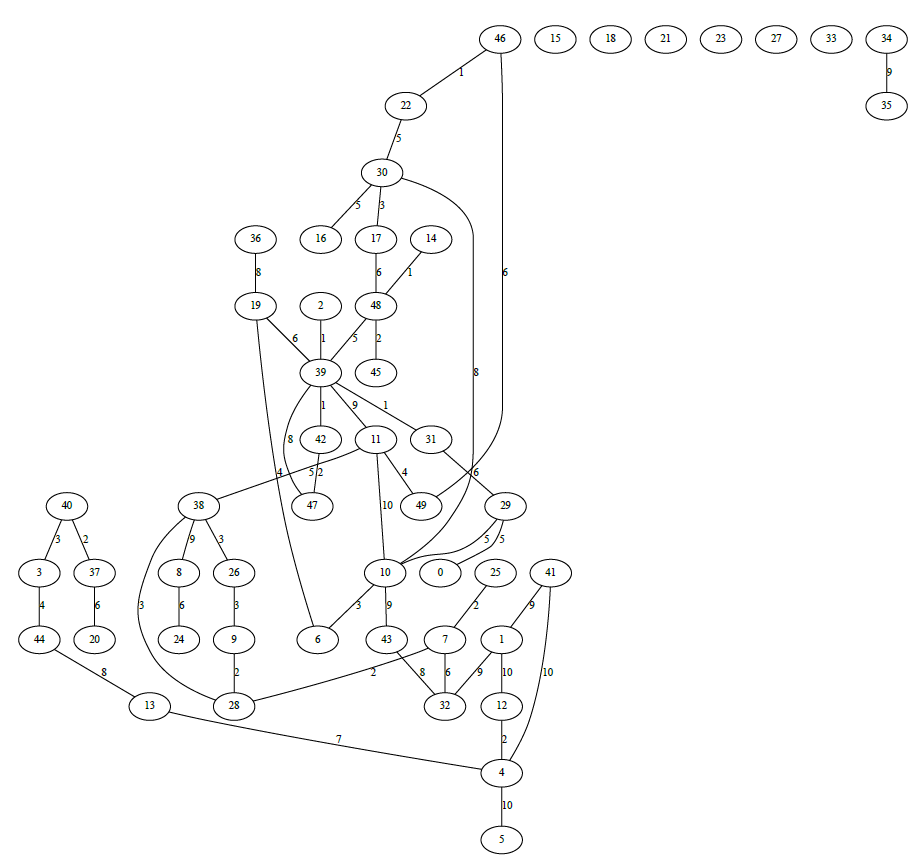
- Dijkstra's algorithm
Dijkstra's algorithm is an algorithm for finding the shortest paths between nodes in a graph
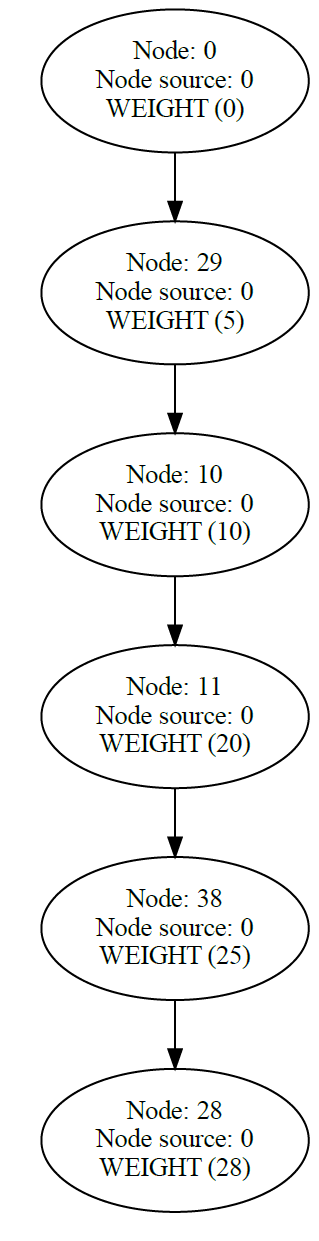
Another example of Dijkstra's algorithm to find the shortest path from Node 0 to Node 28
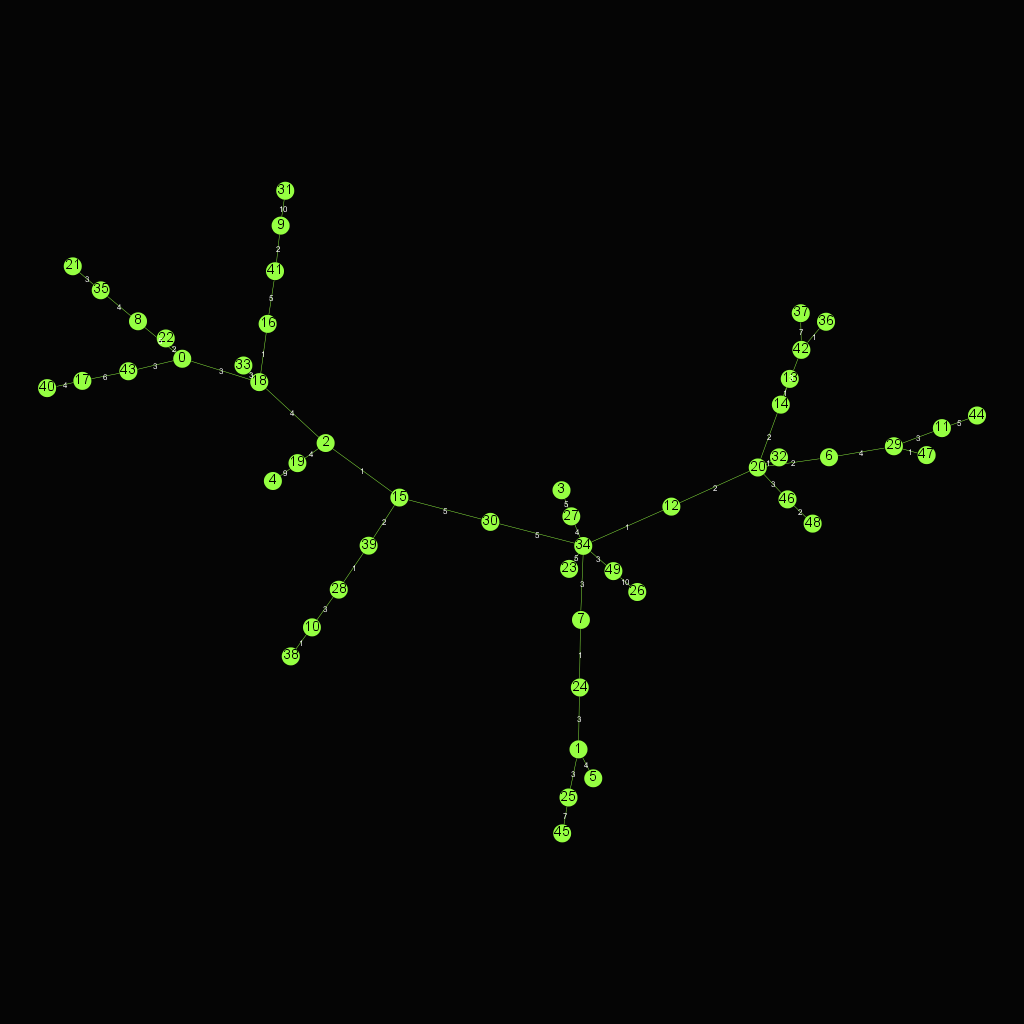
- Kruskal's algorithm
Kruskal's algorithm finds a minimum spanning forest of an undirected edge-weighted graph
- Inverse Kruskal's algorithm
Inverse Kruskal's algorithm (also known as reverse-delete algorithm) is an algorithm in graph theory used to obtain a minimum spanning tree from a given connected, edge-weighted graph. If the graph is disconnected, this algorithm will find a minimum spanning tree for each disconnected part of the graph. The set of these minimum spanning trees is called a minimum spanning forest, which contains every vertex in the graph.
- Prim's algorithm
Prim's (also known as Jarník's) algorithm is a greedy algorithm that finds a minimum spanning tree for a weighted undirected graph. This means it finds a subset of the edges that forms a tree that includes every vertex, where the total weight of all the edges in the tree is minimized.
Available commands:
make install Install dependencies.
make tests Run tests
make linter Execute linter
- Barabási-Albert model
g = models.barabasi(30, 30, True)
g.create_graphviz('Barabasi_directed_30')- Gilbert model
g = models.gilbert(30, 0.3)
g.create_graphviz('Gilbert_30')There are more examples in test_graph.py
- Breadth-first search (BFS)
g = models.erdos_rengy(100, 150)
bfs = g.bfs(0)
bfs.create_graphviz('BFS_100')There are more examples in test_bfs.py
- Depth-first search (DFS)
g = models.erdos_rengy(100, 150)
dfs = g.dfs(0)
dfs.create_graphviz('DFS_100')- Depth-first search (DFS) recursive
g = models.erdos_rengy(100, 150)
dfs = g.dfs_r(0)
dfs.create_graphviz('DFS_R_100')There are more examples in test_dfs.py
- Dijkstra
# Find shortest path from node 0 to node 38
g = models.erdos_rengy(100, 150)
# Assign random weight to each edge
for e in g.edges.values():
e.attr["WEIGHT"] = randint(1,10)
graph_dijkstra = g.dijkstra(0, 38)
graph_dijkstra.create_graphviz('dijkstra_calculado',"WEIGHT",0)There are more examples in test_dijkstra.py
- Kruskal
# Find minimum spanning forest
g = models.erdos_rengy(50, 100)
# Assign random weight to each edge
for e in g.edges.values():
e.attr["WEIGHT"] = randint(1,10)
kruskal_graph = g.KruskalD()
kruskal_graph.create_graphviz('KruskalD_50_calculado', attr_label_edge="WEIGHT", source=0)There are more examples in test_kruskal.py
- Inverse Kruskal
# Find minimum spanning forest
g = models.erdos_rengy(50, 100)
# Assign random weight to each edge
for e in g.edges.values():
e.attr["WEIGHT"] = randint(1,10)
kruskal_graph = g.Kruskal()
kruskal_graph.create_graphviz('inverse_Kruskal_50_calculado', attr_label_edge="WEIGHT", source=0)There are more examples in test_kruskal.py
- Prim
# Find minimum spanning forest
g = models.erdos_rengy(50, 100)
# Assign random weight to each edge
for e in g.edges.values():
e.attr["WEIGHT"] = randint(1,10)
prim_graph = g.Prim()
prim_graph.create_graphviz('prim_50_calculado', attr_label_edge="WEIGHT", source=0)There are more examples in test:_kruskal.py
Run test
make tests
To execute specific test
python -m unittest test.test_graph.TestGraph -v