library(devtools)
install_github('ievaKer/aPEAR')library(devtools)
install_github('ievaKer/aPEAR@development')library(aPEAR)
library(clusterProfiler)
library(org.Hs.eg.db)
library(DOSE)
data(geneList)
enrich <- gseGO(geneList, OrgDb = org.Hs.eg.db, ont = 'CC')
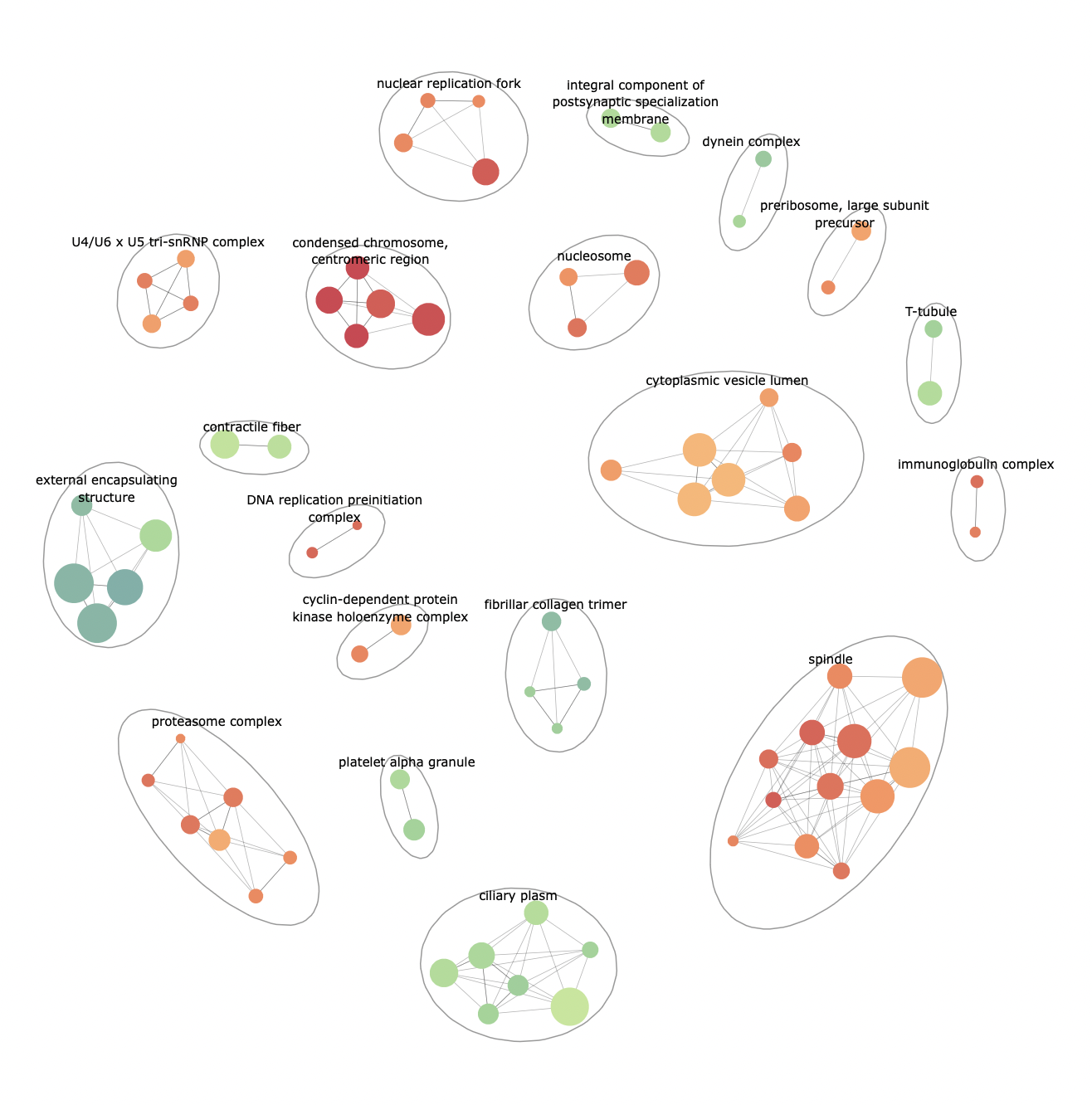
p <- enrichmentNetwork(enrich@result, drawEllipses = TRUE, fontSize = 2.5)
pTo create interactive plots, use plotly:
library(plotly)
ggplotly(p, tooltip=c('ID', 'Cluster', 'Cluster size'))You may find example plots in results directory of this repository.