Foldseek enables fast and sensitive comparisons of large structure sets.
- Foldseek
- Webserver
- Installation
- Memory requirments
- Tutorial Video
- Documentation
- Quick Start
- Main Modules
- Examples
Search your protein structures against the AlphaFoldDB and PDB in seconds using our Foldseek webserver: search.foldseek.com 🚀
# Linux AVX2 build (check using: cat /proc/cpuinfo | grep avx2)
wget https://mmseqs.com/foldseek/foldseek-linux-avx2.tar.gz; tar xvzf foldseek-linux-avx2.tar.gz; export PATH=$(pwd)/foldseek/bin/:$PATH
# Linux SSE4.1 build (check using: cat /proc/cpuinfo | grep sse4_1)
wget https://mmseqs.com/foldseek/foldseek-linux-sse41.tar.gz; tar xvzf foldseek-linux-sse41.tar.gz; export PATH=$(pwd)/foldseek/bin/:$PATH
# MacOS
wget https://mmseqs.com/foldseek/foldseek-osx-universal.tar.gz; tar xvzf foldseek-osx-universal.tar.gz; export PATH=$(pwd)/foldseek/bin/:$PATH
# Conda installer (Linux and macOS)
conda install -c conda-forge -c bioconda foldseek
Other precompiled binaries for ARM64 amd SSE2 are available at https://mmseqs.com/foldseek.
For optimal software performance, consider three options based on your RAM and search requirements:
-
With Cα info (default). Use this formula to calculate RAM -
(6 bytes Cα + 1 3Di byte + 1 AA byte) * (database residues). The 54M AFDB50 entries require 151GB. -
Without Cα info. By disabling
--sort-by-structure-bits 0, RAM requirement reduces to 35GB. However, this alters hit rankings and final scores but not E-values. Structure bits are mostly relevant for hit ranking for E-value > 10^-1. -
Single query searches. Use the
--prefilter-mode 1, which isn't memory-limited and computes all ungapped alignments. This option optimally utilizes foldseek's multithreading capabilities for single queries.
We presented a Foldseek tutorial at the SBGrid where we demonstrate the webserver and command line interface of foldseek. Check it out here.
Many of Foldseek's modules (subprograms) rely on MMseqs2. For more information about these modules, refer to the MMseqs2 wiki. For documentation specific to Foldseek, checkout the Foldseek wiki here.
The easy-search module allows to search single or multiple query structures, formatted in PDB/mmCIF format (flat or gzipped), against a target database, folder or single protein structures. In default it outputs the alignment information as a tab-separated file but we support also Superposed Cα PDBs or a HTML output.
foldseek easy-search example/d1asha_ example/ aln tmpFolder
The default fields are containing the following fields: query,target,fident,alnlen,mismatch,gapopen,qstart,qend,tstart,tend,evalue,bits but they can be customized with the --format-output option e.g. --format-output "query,target,qaln,taln" returns the query and target accession and the pairwise alignments in tab separated format. You can choose many different output columns.
| Code | Description |
|---|---|
| query | Query sequence identifier |
| target | Target sequence identifier |
| qca | Calpha corrdinates of the query |
| tca | Calpha corrdinates of the target |
| alntmscore | TM-score of the alignment |
| qtmscore | TM-score normalized by the query length |
| ttmscore | TM-score normalized by the target length |
| u | Rotation matrix (computed to by TM-score) |
| t | Translation vector (computed to by TM-score) |
| lddt | Average LDDT of the alignment |
| lddtfull | LDDT per aligned position |
| prob | Estimated probability for query and target to be homologous (e.g. being within the same SCOPe superfamily) |
Check out the MMseqs2 documentation for more format output codes.
Foldseek's --format-mode 5 generates PDB files with all Cα atoms superimposed based on the aligned coordinates on to the query structure.
For each pairwise alignment it will write a single PDB files, so be carefull when using this options for large searches.
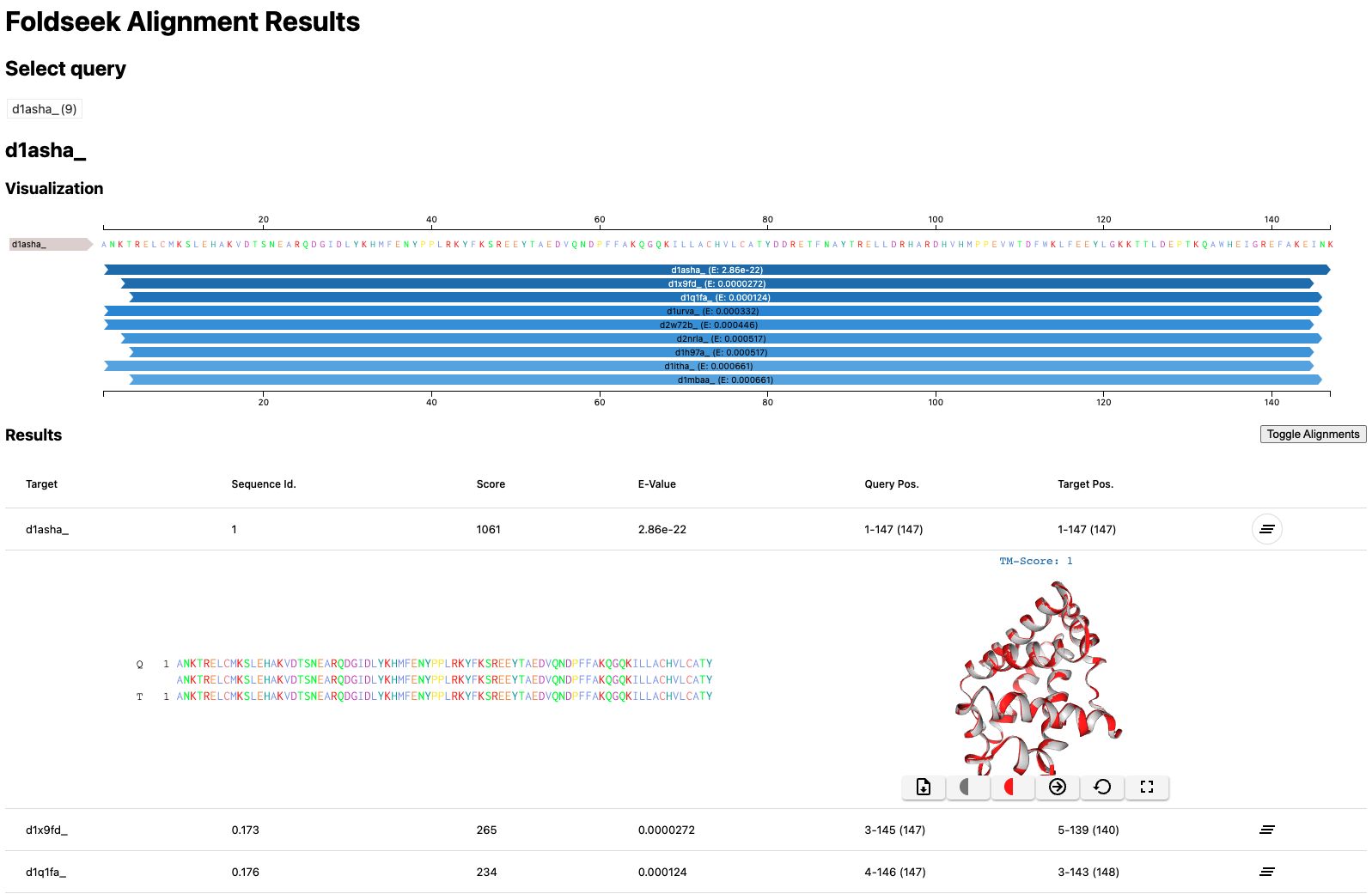
Foldseek can locally generate a search result HTML similiar to the webserver by specifying the format mode --format-mode 3
foldseek easy-search example/d1asha_ example/ result.html tmp --format-mode 3
| Option | Category | Description |
|---|---|---|
| -s | Sensitivity | Adjust sensitivity to speed trade-off; lower is faster, higher more sensitive (fast: 7.5, default: 9.5) |
| --exhaustive-search | Sensitivity | Skips prefilter and performs an all-vs-all alignment (more sensitive but much slower) |
| --max-seqs | Sensitivity | Adjust the amount of prefilter handed to alignment; increasing it can lead to more hits (default: 1000) |
| -e | Sensitivity | List matches below this E-value (range 0.0-inf, default: 0.001); increasing it reports more distant structures |
| --alignment-type | Alignment | 0: 3Di Gotoh-Smith-Waterman (local, not recommended), 1: TMalign (global, slow), 2: 3Di+AA Gotoh-Smith-Waterman (local, default) |
| -c | Alignment | List matches above this fraction of aligned (covered) residues (see --cov-mode) (default: 0.0); higher coverage = more global alignment |
| --cov-mode | Alignment | 0: coverage of query and target, 1: coverage of target, 2: coverage of query |
In default Foldseek uses its local 3Di+AA strutural alignment but it also supports to realign hits using the global TMalign as well as rescoring alignments using TMscore.
foldseek easy-search example/d1asha_ example/ aln tmp --alignment-type 1
In case of the alignment type (--alignment-type 1) tmalign, we sort the results by the TMscore normalized by query length. We write the TMscore into the e-value=(qTMscore+tTMscore)/2 as well as into the score(=qTMscore*100) field. All output fields (like pident, fident, and alnlen) are calculated from the TMalign alignment.
The databases command downloads pre-generated databases like PDB or AlphaFoldDB.
# pdb
foldseek databases PDB100 pdb tmp
# alphafold db
foldseek databases Alphafold/Proteome afdb tmp
We currently support the following databases:
Name Type Taxonomy Url
- Alphafold/UniProt Aminoacid yes https://alphafold.ebi.ac.uk/
- Alphafold/UniProt50 Aminoacid yes https://alphafold.ebi.ac.uk/
- Alphafold/Proteome Aminoacid yes https://alphafold.ebi.ac.uk/
- Alphafold/Swiss-Prot Aminoacid yes https://alphafold.ebi.ac.uk/
- ESMAtlas30 Aminoacid - https://esmatlas.com
- PDB Aminoacid yes https://www.rcsb.org
The target database can be pre-processed by createdb. This make sense if searched multiple times.
foldseek createdb example/ targetDB
foldseek createindex targetDB tmp #OPTIONAL generates and stores the index on disk
foldseek easy-search example/d1asha_ targetDB aln.m8 tmpFolder
The easy-cluster algorithm is designed for structural clustering by assigning structures to a representative protein using structural alignment. It accepts input in either PDB or mmCIF format, with support for both flat and gzipped files. By default, easy-cluster generates three output files with the following prefixes: (1) _clu.tsv, (2) _repseq.fasta, and (3) _allseq.fasta. The first file (1) is a tab-separated file describing the mapping from representative to member, while the second file (2) contains only representative sequences, and the third file (3) includes all cluster member sequences.
foldseek easy-cluster example/ res tmp -c 0.9
The provided format represents protein structure clustering in a tab-separated, two-column layout (representative and member). Each line denotes a cluster-representative and cluster-member relationship, signifying that the member shares significant structural similarity with the representative, and thus belongs to the same cluster.
Q0KJ32 Q0KJ32
Q0KJ32 C0W539
Q0KJ32 D6KVP9
E3HQM9 E3HQM9
E3HQM9 F0YHT8
The _repseq.fasta contains all representative protein sequences of the clustering.
>Q0KJ32
MAGA....R
>E3HQM9
MCAT...Q
In _allseq.fasta file all sequences of the cluster are present. A new cluster is marked by two identical name lines of the representative sequence, where the first line stands for the cluster and the second is the name line of the first cluster sequence. It is followed by the fasta formatted sequences of all its members.
>Q0KJ32
>Q0KJ32
MAGA....R
>C0W539
MVGA....R
>D6KVP9
MVGA....R
>D1Y890
MVGV....R
>E3HQM9
>E3HQM9
MCAT...Q
>Q223C0
MCAR...Q
| Option | Category | Description |
|---|---|---|
| -e | Sensitivity | List matches below this E-value (range 0.0-inf, default: 0.001); increasing it reports more distant structures |
| --alignment-type | Alignment | 0: 3Di Gotoh-Smith-Waterman (local, not recommended), 1: TMalign (global, slow), 2: 3Di+AA Gotoh-Smith-Waterman (local, default) |
| -c | Alignment | List matches above this fraction of aligned (covered) residues (see --cov-mode) (default: 0.0); higher coverage = more global alignment |
| --cov-mode | Alignment | 0: coverage of query and target, 1: coverage of target, 2: coverage of query |
| --min-seq-id | Alignment | the minimum sequence identity to be clustered |
easy-searchfast protein structure searcheasy-clusterfast protein structure clusteringcreatedbcreate a database from protein structures (PDB,mmCIF, mmJSON)databasesdownload pre-assembled databases
Easiest way to get the alignment TMscore normalized by min(alnLen,qLen,targetLen) as well as a rotation matrix is through the following command:
foldseek easy-search example/ example/ aln tmp --format-output query,target,alntmscore,u,t
Alternative, it is possible to compute TMscores for the kind of alignment output (e.g. 3Di/AA) using the following commands:
foldseek createdb example/ targetDB
foldseek createdb example/ queryDB
foldseek search queryDB targetDB aln tmpFolder -a
foldseek aln2tmscore queryDB targetDB aln aln_tmscore
foldseek createtsv queryDB targetDB aln_tmscore aln_tmscore.tsv
Output format aln_tmscore.tsv: query and target identifier, TMscore, translation(3) and rotation vector=(3x3)
The following command aligns the input structures all-against-all and keeps only alignments with 80% of the sequence covered by the alignment (-c 0.8) (read more about alignment coverage here). It then clusters the results using greedy set cover algorithm. The clustering mode can be adjusted using --cluster-mode, read more here. The clustering output format is described here.
foldseek createdb example/ db
foldseek search db db aln tmpFolder -c 0.8
foldseek clust db aln clu
foldseek createtsv db db clu clu.tsv
Foldseek can generate a3m based multiple sequence alignments using the following commands.
a3m can be converted to fasta format using reformat.pl (reformat.pl in.a3m out.fas).
foldseek createdb example/ targetDB
foldseek createdb example/ queryDB
foldseek search queryDB targetDB aln tmpFolder -a
foldseek result2msa queryDB targetDB aln msa --msa-format-mode 6
foldseek unpackdb msa msa_output --unpack-suffix a3m --unpack-name-mode 0