logKDE: log-transformed kernel density estimation
The goal of logKDE is to provide a set of functions for kernel density
estimation on the positive domain, using log-kernel density functions,
for the R programming environment. The main functions of the package
are the logdensity and logdensity_fft functions. The choice of
functional syntax was made to resemble those of the density function,
for conducting kernel density estimation on the real domain. The
logdensity function conducts density estimation, via first principle
computations, whereas logdensity_fft utilizes fast-Fourier
transformation in order to speed up computation. The use of Rcpp
guarantees that both methods are sufficiently fast for large data
scenarios.
Currently, a variety of kernel functions and plugin bandwidth methods
are available. By default both logdensity and logdensity_fft are set
to use log-normal kernel functions (kernel = 'gaussian') and
Silverman’s rule-of-thumb bandwidth, applied to log-transformed data
(bw = 'nrd0'). However, the following kernels are also available:
- log-Epanechnikov (
kernel = 'epanechnikov'), - log-Laplace (
kernel = 'laplace'), - log-logistic (
kernel = 'logistic'), - log-triangular (
kernel = 'triangular'), - log-uniform (
kernel = 'uniform').
The following plugin bandwidth methods are also available:
- all of the methods that available for density, applied to
log-transformed data (see
?bw.nrdregarding the options), - unbiased cross-validated bandwidths in the positive domain (
bw = 'logcv'), - a Silverman-type rule-of-thumb that optimizes the kernel density
estimator fit, compared to a log-normal density function (
bw = 'logg').
The logdensity and logdensity_fft functions also behave in the same
way as density, when called within the plot function. The usual
assortment of commands that apply to plot output objects can also be
called.
For a comprehensive review of the literature on positive-domain kernel
density estimation, thorough descriptions of the mathematics relating to
the methods that have been described, simulation results, and example
applications of the logKDE package, please consult the package
vignette. The vignette is available via the command
vignette('logKDE'), once the package is installed.
Installation
If devtools has already been installed, then the most current build of
logKDE can be obtained via the command:
devtools::install_github('andrewthomasjones/logKDE',build_vignettes = T)The latest stable build of logKDE can be obtain from CRAN via the
command:
install.packages("logKDE", repos='http://cran.us.r-project.org')An archival build of logKDE is available at
https://zenodo.org/record/1317784. Manual installation instructions
can be found within the R installation and administration manual
https://cran.r-project.org/doc/manuals/r-release/R-admin.html.
Examples
Example 1
In this example, we demonstrate that logdensity has nearly identical
syntax to density. We also show that the format of the outputs are
also nearly identical.
## Load 'logKDE' library.
library(logKDE)
## Set a random seed.
set.seed(1)
## Generate strictly positive data.
## Data are generated from a chi-squared distribution with 12 degrees of freedom.
x <- rchisq(100,6)
## Construct and print the output of the function 'density'.
density(x)
#>
#> Call:
#> density.default(x = x)
#>
#> Data: x (100 obs.); Bandwidth 'bw' = 1.018
#>
#> x y
#> Min. :-2.366 Min. :0.0000475
#> 1st Qu.: 2.547 1st Qu.:0.0072263
#> Median : 7.459 Median :0.0331904
#> Mean : 7.459 Mean :0.0508396
#> 3rd Qu.:12.372 3rd Qu.:0.1013289
#> Max. :17.284 Max. :0.1312107
## Construct and print the output of the function 'logdensity'.
logdensity(x)
#>
#> Call:
#> logdensity(x = x)
#>
#> Data: x (100 obs.); Bandwidth 'bw' = 0.1923
#>
#> x y
#> Min. : 0.1111 Min. :0.00000
#> 1st Qu.: 3.7851 1st Qu.:0.02313
#> Median : 7.4592 Median :0.06527
#> Mean : 7.4592 Mean :0.06707
#> 3rd Qu.:11.1333 3rd Qu.:0.11219
#> Max. :14.8073 Max. :0.13698
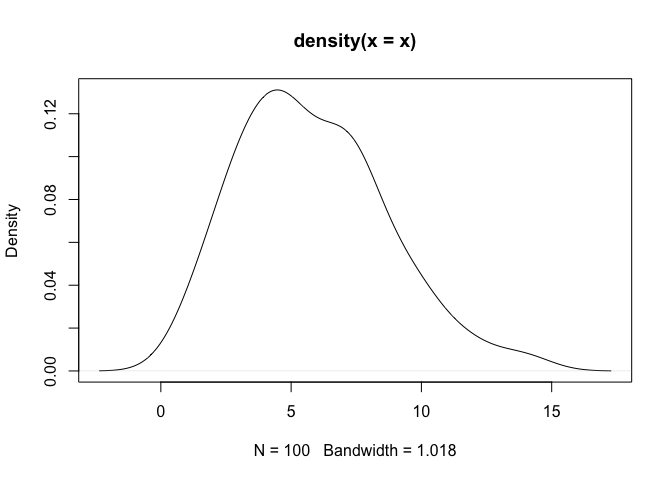
## Plot the 'density' output object.
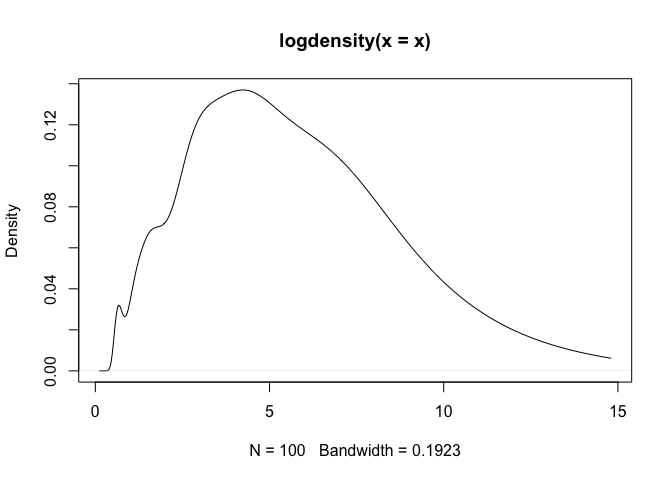
plot(density(x))## Plot the 'logdensity' output object.
plot(logdensity(x))As a note, one can observe that density assigns positive probability
to negative values. Since we know that the chi-squared generative model
generates only positive values, this is an undesirable result. The
log-transformed kernel density estimator that is produced by
logdensity only assigns positive probability to positive values, and
is thus bona fide in this estimation scenario.
Example 2
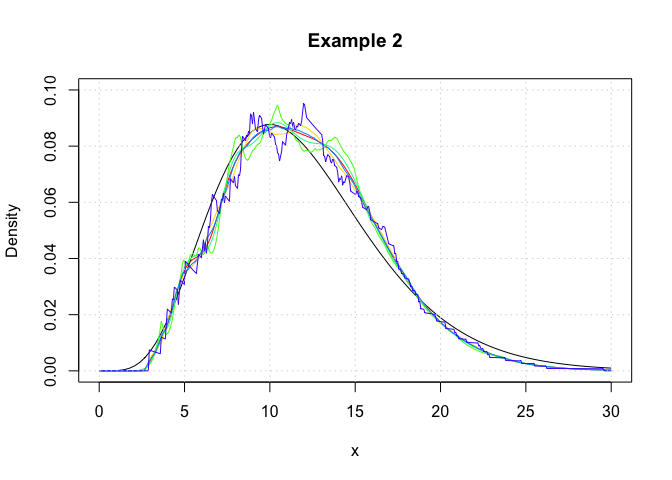
In this example, we showcase the variety of kernel functions that are
available in the package. Here, log-transformed kernel density
estimators are constructed using the logdensity function.
## Load 'logKDE' library.
library(logKDE)
## Set a random seed.
set.seed(1)
## Generate strictly positive data.
## Data are generated from a chi-squared distribution with 12 degrees of freedom.
x <- rchisq(100,12)
## Construct a log-KDE using the data, and using each of the available kernel functions.
logKDE1 <- logdensity(x,kernel = 'gaussian',from = 1e-6,to = 30)
logKDE2 <- logdensity(x,kernel = 'epanechnikov',from = 1e-6,to = 30)
logKDE3 <- logdensity(x,kernel = 'laplace',from = 1e-6,to = 30)
logKDE4 <- logdensity(x,kernel = 'logistic',from = 1e-6,to = 30)
logKDE5 <- logdensity(x,kernel = 'triangular',from = 1e-6,to = 30)
logKDE6 <- logdensity(x,kernel = 'uniform',from = 1e-6,to = 30)
## Plot the true probability density function of the generative model.
plot(c(0,30),c(0,0.1),type='n',xlab='x',ylab='Density',main='Example 2')
curve(dchisq(x,12),from = 0,to = 30,add = T)
## Plot each of the log-KDE functions, each in a different rainbow() colour.
lines(logKDE1$x,logKDE1$y,col = rainbow(7)[1])
lines(logKDE2$x,logKDE2$y,col = rainbow(7)[2])
lines(logKDE3$x,logKDE3$y,col = rainbow(7)[3])
lines(logKDE4$x,logKDE4$y,col = rainbow(7)[4])
lines(logKDE5$x,logKDE5$y,col = rainbow(7)[5])
lines(logKDE6$x,logKDE6$y,col = rainbow(7)[6])
## Add a grid for a visual guide.
grid()Example 3
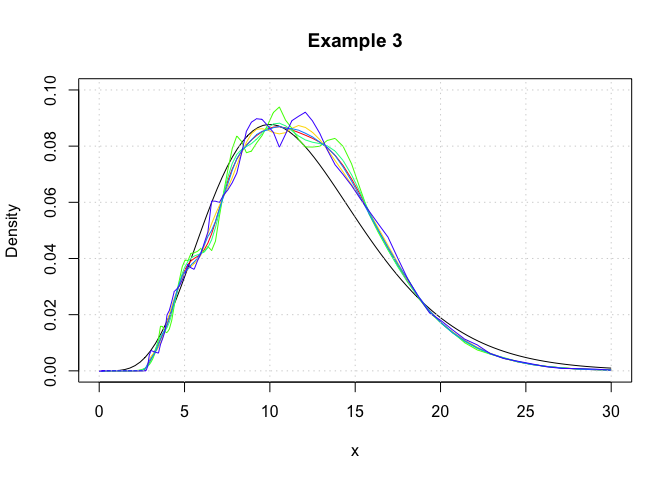
In this example, we show that logdensity and logdensity_ftt yield
nearly identical results. Here, log-transformed kernel density
estimators are constructed using the logdensity_ftt function.
## Load 'logKDE' library.
library(logKDE)
## Set a random seed.
set.seed(1)
## Generate strictly positive data.
## Data are generated from a chi-squared distribution with 12 degrees of freedom.
x <- rchisq(100,12)
## Construct a log-KDE using the data, and using each of the available kernel functions.
logKDE1 <- logdensity_fft(x,kernel = 'gaussian',from = 1e-6,to = 30)
logKDE2 <- logdensity_fft(x,kernel = 'epanechnikov',from = 1e-6,to = 30)
logKDE3 <- logdensity_fft(x,kernel = 'laplace',from = 1e-6,to = 30)
logKDE4 <- logdensity_fft(x,kernel = 'logistic',from = 1e-6,to = 30)
logKDE5 <- logdensity_fft(x,kernel = 'triangular',from = 1e-6,to = 30)
logKDE6 <- logdensity_fft(x,kernel = 'uniform',from = 1e-6,to = 30)
## Plot the true probability density function of the generative model.
plot(c(0,30),c(0,0.1),type='n',xlab='x',ylab='Density',main='Example 3')
curve(dchisq(x,12),from = 0,to = 30,add = T)
## Plot each of the log-KDE functions, each in a different rainbow() colour.
lines(logKDE1$x,logKDE1$y,col = rainbow(7)[1])
lines(logKDE2$x,logKDE2$y,col = rainbow(7)[2])
lines(logKDE3$x,logKDE3$y,col = rainbow(7)[3])
lines(logKDE4$x,logKDE4$y,col = rainbow(7)[4])
lines(logKDE5$x,logKDE5$y,col = rainbow(7)[5])
lines(logKDE6$x,logKDE6$y,col = rainbow(7)[6])
## Add a grid for a visual guide.
grid()We observe that the logdensity_fft outputs are noticiably smoother
than those of logdensity. This is because fast Fourier transformations
(FFT) only yield kernel density estimates at discrete points, and the
regions between these discrete points are approximated via a linear
approximator, namely using the approx function. This is the same
evaluation technique as that which is used in the function density.
Additionally the FFT approximation points are evenly space on the real
line, whereas those used for logdensity are evenly spaced on a log
scale.
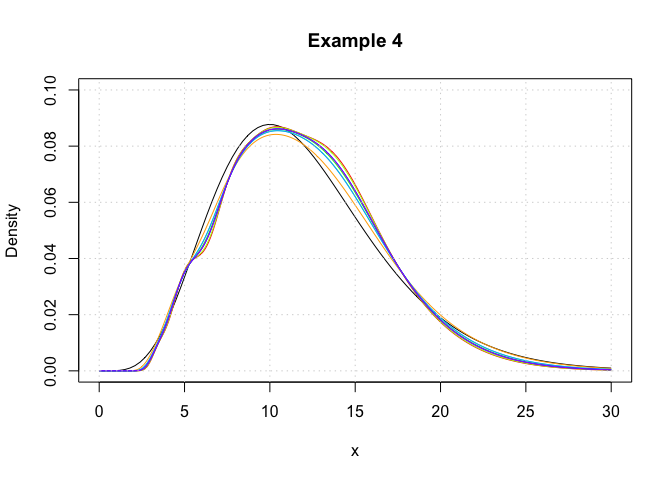
Example 4
In this example, we showcase the variety of plugin bandwidth estimators
that are available in the package. Here, log-transformed kernel density
estimators are constructed using the logdensity function.
## Load 'logKDE' library.
library(logKDE)
## Set a random seed.
set.seed(1)
## Generate strictly positive data.
## Data are generated from a chi-squared distribution with 12 degrees of freedom.
x <- rchisq(100,12)
## Construct a log-KDE using the data, and using each of the available kernel functions.
logKDE1 <- logdensity(x,bw = 'nrd0',from = 1e-6,to = 30)
logKDE2 <- logdensity(x,bw = 'logcv',from = 1e-6,to = 30)
logKDE3 <- logdensity(x,bw = 'logg',from = 1e-6,to = 30)
logKDE4 <- logdensity(x,bw = 'nrd',from = 1e-6,to = 30)
logKDE5 <- logdensity(x,bw = 'ucv',from = 1e-6,to = 30)
#> Warning in stats::bw.ucv(log(x)): minimum occurred at one end of the range
logKDE6 <- logdensity(x,bw = 'bcv',from = 1e-6,to = 30)
#> Warning in stats::bw.bcv(log(x)): minimum occurred at one end of the range
logKDE7 <- logdensity(x,bw = 'SJ-ste',from = 1e-6,to = 30)
logKDE8 <- logdensity(x,bw = 'SJ-dpi',from = 1e-6,to = 30)
## Plot the true probability density function of the generative model.
plot(c(0,30),c(0,0.1),type='n',xlab='x',ylab='Density',main='Example 4')
curve(dchisq(x,12),from = 0,to = 30,add = T)
## Plot each of the log-KDE functions with different choices of bandwidth, each in a different rainbow() colour.
lines(logKDE1$x,logKDE1$y,col = rainbow(9)[1])
lines(logKDE2$x,logKDE2$y,col = rainbow(9)[2])
lines(logKDE3$x,logKDE3$y,col = rainbow(9)[3])
lines(logKDE4$x,logKDE4$y,col = rainbow(9)[4])
lines(logKDE5$x,logKDE5$y,col = rainbow(9)[5])
lines(logKDE6$x,logKDE6$y,col = rainbow(9)[6])
lines(logKDE7$x,logKDE7$y,col = rainbow(9)[7])
lines(logKDE8$x,logKDE8$y,col = rainbow(9)[8])
## Add a grid for a visual guide.
grid()Unit testing
Using the package testthat, we have conducted the following unit test
for the GitHub build, on the date: 06 August, 2018. The testing files
are contained in the
tests
folder of the respository.
## Load 'logKDE' library.
library(logKDE)
## Load 'testthat' library.
library(testthat)
## Test 'logKDE'.
test_package('logKDE')
#> ══ testthat results ════════════════════════════════════════════════════════════════════════════════════════════════════
#> OK: 74 SKIPPED: 0 FAILED: 0Bug reporting and contributions
Thank you for your interest in logKDE. If you happen to find any bugs
in the program, then please report them on the Issues page
(https://github.com/andrewthomasjones/logKDE/issues). Support can also
be sought on this page. Furthermore, if you would like to make a
contribution to the software, then please forward a pull request to the
owner of the repository.