The outline of the assigment is to build a deep neural network for a image classification task is as follows:
- Initialize the parameters for a L layer neural network.
- Implement the forward propagation module to compute the activation functions for L layers. We calculate RELU function for L-1 layers and Sigmoid function for the Lth layer. We store Z as a cache to be used in while calculating gradients in the backward propagation.
- Compute the loss.
- Implement the backward propagation module to compute the gradients of activation function and parameters.
- Update the parameters using gradient descent method.
- Superscript [l] denotes a quantity associated with the lth layer.
- Example: a[L] is the Lth layer activation. W[L] and b[L] are the Lth layer parameters.
- Superscript (i) denotes a quantity associated with the ith example.
- Example: x(i) is the ith training example.
- Lowerscript i denotes the ith entry of a vector.
- Example: a[l]_i denotes the ith entry of the lth layer's activations.
import time
import numpy as np
import h5py
import matplotlib.pyplot as plt
import scipy
from PIL import Image
from scipy import ndimage
%matplotlib inline
plt.rcParams['figure.figsize'] = (5.0, 4.0) # set default size of plots
plt.rcParams['image.interpolation'] = 'nearest'
plt.rcParams['image.cmap'] = 'gray'
%load_ext autoreload
%autoreload 2
np.random.seed(1)We write a helper function to initialize the parameters.
We store nl, the number of units in different layers in a variable layer_dims. For example, layer_dims= [2,4,1] is a neural network with 2 inputs, one hidden layer with 4 neurons, one output layer with one output unit.
def initialize_parameters_deep(layer_dims):
"""
Arguments:
layer_dims -- python array (list) containing the dimensions of each layer in our network
Returns:
parameters -- python dictionary containing your parameters "W1", "b1", ..., "WL", "bL":
Wl -- weight matrix of shape (layer_dims[l], layer_dims[l-1])
bl -- bias vector of shape (layer_dims[l], 1)
"""
parameters={}
L= len(layer_dims)
for l in range(1,L):
parameters['W'+str(l)]= np.random.randn(layer_dims[l], layer_dims[l-1])*0.01
parameters['b'+str(l)]= np.zeros((layer_dims[l],1))
assert(parameters['W' + str(l)].shape == (layer_dims[l], layer_dims[l-1]))
assert(parameters['b' + str(l)].shape == (layer_dims[l], 1))
return parametersAfter initializing the parameters, we will now do the forward propagation module. We will implement some basic functions that we will use later when implementing the model. We will complete three functions in this order:
- LINEAR
- LINEAR -> ACTIVATION where ACTIVATION will be either ReLU or Sigmoid.
- [LINEAR -> RELU] X (L-1) -> LINEAR -> SIGMOID (whole model)
The linear forward module (vectorized over all the examples) computes the following equations:
Z[l] = W[l]A[l-1] +b[l]
where A[0] = X.
def linear_forward(A,W,b):
"""
Implement the linear part of a layer's forward propagation.
Arguments:
A -- activations from previous layer (or input data): (size of previous layer, number of examples)
W -- weights matrix: numpy array of shape (size of current layer, size of previous layer)
b -- bias vector, numpy array of shape (size of the current layer, 1)
Returns:
Z -- the input of the activation function, also called pre-activation parameter
cache -- a python dictionary containing "A", "W" and "b" ; stored for computing the backward pass efficiently
"""
Z= np.dot(W,A)+b
assert(Z.shape==(W.shape[0],A.shape[1]))
cache= (A,W,b)
return Z, cacheWe use two activation functions:
def sigmoid(Z):
A = 1/(1+np.exp(-Z))
cache = Z
return A, cachedef relu(Z):
A = np.maximum(0,Z)
assert(A.shape == Z.shape)
cache = Z
return A, cachedef linear_activation_forward(A_prev,W,b,activation):
"""
Implement the forward propagation for the LINEAR->ACTIVATION layer
Arguments:
A_prev -- activations from previous layer (or input data): (size of previous layer, number of examples)
W -- weights matrix: numpy array of shape (size of current layer, size of previous layer)
b -- bias vector, numpy array of shape (size of the current layer, 1)
activation -- the activation to be used in this layer, stored as a text string: "sigmoid" or "relu"
Returns:
A -- the output of the activation function, also called the post-activation value
cache -- a python dictionary containing "linear_cache" and "activation_cache";
stored for computing the backward pass efficiently
"""
if activation=='sigmoid':
Z, linear_cache= linear_forward(A_prev,W,b)
A, activation_cache= sigmoid(Z)
elif activation=='relu':
Z, linear_cache= linear_forward(A_prev,W,b)
A, activation_cache= relu(Z)
assert(A.shape==(W.shape[0],A_prev.shape[1]))
cache= (linear_cache, activation_cache)
return A, cacheNow, we will create a function which implements the function linear_activation_forward created in the last step with RELU L-1 times and with Sigmoid one time for the Lth layer.
In the code below, the variable AL denotes activation function for the Lth layer.
def L_model_forward(X, parameters):
caches=[]
A=X
L= len(parameters)//2
for l in range(1,L):
A_prev=A
A,cache= linear_activation_forward(A_prev, parameters['W'+str(l)],
parameters['b'+str(l)],
activation='relu')
caches.append(cache)
AL, cache= linear_activation_forward(A, parameters['W'+str(L)],
parameters['b'+str(L)],
activation='sigmoid')
caches.append(cache)
assert(AL.shape==(1, X.shape[1]))
return AL, cachesNow we calculate the cross-entropy loss, so as to check whether our model is learning or not. Our objective is to minimize the cost by optimizing the parameters W and b using gradient descent.
def compute_cost(AL,Y):
m= Y.shape[1]
cost= -(1/m)*np.sum(Y*np.log(AL)+(1-Y)*np.log(1-AL))
cost= np.squeeze(cost)
assert(cost.shape==())
return costSimilar to forward propagation module, we will create helper functions to caclulate gradients of the Loss functions with respect to the parameters. We will achieve this in 3 steps:
- LINEAR backward
- LINEAR -> ACTIVATION backward where ACTIVATION computes the derivative of either the ReLU or sigmoid activation
- [LINEAR -> RELU] X (L-1) -> LINEAR -> SIGMOID backward (whole model)
For a layer l, linear part is Z[l] = W[l]A[l-1] +b[l]. Suppose we have already calculated the derivative dZ[l]. The three outputs(dW[l], db[l], dA[l]) are calculated using dZ[l] using the following formulas:
def linear_backward(dZ, cache):
A_prev, W,b= cache
m= A_prev.shape[1]
dW= (1/m)*np.dot(dZ, A_prev.T)
db= (1/m)*np.sum(dZ, axis=1, keepdims=True)
dA_prev= np.dot(W.T, dZ)
assert(dA_prev.shape==A_prev.shape)
assert (dW.shape == W.shape)
assert (db.shape == b.shape)
return dA_prev, dW, dbIn order to calculate the gradients of the Loss function w.r.t parameters, we first need to calculate dZ[l]. We will two backward functions sigmoid_backward and relu_backward which calculates dZ[l] as .
def relu_backward(dA, cache):
"""
Implement the backward propagation for a single RELU unit.
Arguments:
dA -- post-activation gradient, of any shape
cache -- 'Z' where we store for computing backward propagation efficiently
Returns:
dZ -- Gradient of the cost with respect to Z
"""
Z = cache
dZ = np.array(dA, copy=True) # just converting dz to a correct object.
# When z <= 0, you should set dz to 0 as well.
dZ[Z <= 0] = 0
assert (dZ.shape == Z.shape)
return dZdef sigmoid_backward(dA, cache):
"""
Implement the backward propagation for a single SIGMOID unit.
Arguments:
dA -- post-activation gradient, of any shape
cache -- 'Z' where we store for computing backward propagation efficiently
Returns:
dZ -- Gradient of the cost with respect to Z
"""
Z = cache
s = 1/(1+np.exp(-Z))
dZ = dA * s * (1-s)
assert (dZ.shape == Z.shape)
return dZNow we can implement these functions with the linear_backward function created before to calculate dW[l], db[l] and dA[l]
def linear_activation_backward(dA, cache, activation):
linear_cache, activation_cache= cache
if activation=='relu':
dZ= relu_backward(dA, activation_cache)
dA_prev, dW, db= linear_backward(dZ, linear_cache)
elif activation=='sigmoid':
dZ= sigmoid_backward(dA, activation_cache)
dA_prev, dW, db= linear_backward(dZ, linear_cache)
return dA_prev, dW,dbNow we will implement backward function for the whole network. Remember, we stored (X,W,b,Z) in a value cache when we implemented L_model_forward. We will use these cache values while iterating through all hidden layers starting from layer L.
For Layer l, we know that . We need to calculate dA[L] which we will feed in the back propagation module for the whole network. dA[L] is calculated as
dAL = - (np.divide(Y, AL) - np.divide(1 - Y, 1 - AL)) # derivative of cost with respect to AL.
We will store dA, dW, db in a grads dictionary.
def L_model_backward(AL, Y, caches):
"""
Implement the backward propagation for the [LINEAR->RELU] * (L-1) -> LINEAR -> SIGMOID group
Arguments:
AL -- probability vector, output of the forward propagation (L_model_forward())
Y -- true "label" vector (containing 0 if non-cat, 1 if cat)
caches -- list of caches containing:
every cache of linear_activation_forward() with "relu" (it's caches[l], for l in range(L-1) i.e l = 0...L-2)
the cache of linear_activation_forward() with "sigmoid" (it's caches[L-1])
Returns:
grads -- A dictionary with the gradients
grads["dA" + str(l)] = ...
grads["dW" + str(l)] = ...
grads["db" + str(l)] = ...
"""
grads={}
L= len(caches)
m= AL.shape[1]
Y.reshape(AL.shape)
dAL= - (np.divide(Y, AL) - np.divide(1 - Y, 1 - AL)) # derivative of cost with respect to AL
# Lth layer (SIGMOID -> LINEAR) gradients. Inputs: "AL, Y, caches". Outputs: "grads["dAL"], grads["dWL"], grads["dbL"]
current_cache= caches[-1]
grads['dA'+str(L)], grads['dW'+str(L)], grads['db'+str(L)]= linear_activation_backward(dAL, current_cache,
activation='sigmoid')
for l in reversed(range(L-1)):
# lth layer: (RELU -> LINEAR) gradients.
# Inputs: "grads["dA" + str(l + 2)], caches". Outputs: "grads["dA" + str(l + 1)] , grads["dW" + str(l + 1)] , grads["db" + str(l + 1)]
current_cache= caches[l]
dA_prev_temp, dW_temp, db_temp= linear_activation_backward(grads['dA'+str(l+2)], current_cache,
activation= 'relu')
grads['dA'+str(l)]= dA_prev_temp
grads['dW'+str(l+1)]= dW_temp
grads['db'+str(l+1)]= db_temp
return gradsWe will update the parameters of the model using gradient descent method as:
def update_parameters(parameters,grads, learning_rate):
L= len(parameters)//2
for l in range(L):
parameters['W'+str(l+1)]= parameters['W'+str(l+1)]-learning_rate*grads['dW'+str(l+1)]
parameters['b'+str(l+1)]= parameters['b'+str(l+1)]-learning_rate*grads['db'+str(l+1)]
return parametersCreating a function to predict the results of a L-layered deep neural network
def predict(X,y, parameters):
"""
This function is used to predict the results of a L-layer neural network.
Arguments:
X -- data set of examples you would like to label
parameters -- parameters of the trained model
Returns:
p -- predictions for the given dataset X
"""
m= X.shape[1]
L= len(parameters)//2
p= np.zeros((1,m))
probas, caches= L_model_forward(X, parameters)
for i in range(0, probas.shape[1]):
if probas[0,i]>0.5:
p[0,i]=1
else:
p[0,i]=0
print("Accuracy: " + str(np.sum((p == y)/m)))
return pWe will use all the functions created above to classify images whether they are cat or not. The data is in h5 format. We first create a function load_dataset to load the data and split it into training and testing. The dataset contains:
- a training set of m_train images labelled as cat (1) or non-cat (0)
- a test set of m_test images labelled as cat and non-cat
- each image is of shape (num_px, num_px, 3) where 3 is for the 3 channels (RGB).
def load_dataset():
train_dataset = h5py.File('datasets/train_catvnoncat.h5', "r")
train_set_x_orig = np.array(train_dataset["train_set_x"][:]) # your train set features
train_set_y_orig = np.array(train_dataset["train_set_y"][:]) # your train set labels
test_dataset = h5py.File('datasets/test_catvnoncat.h5', "r")
test_set_x_orig = np.array(test_dataset["test_set_x"][:]) # your test set features
test_set_y_orig = np.array(test_dataset["test_set_y"][:]) # your test set labels
classes = np.array(test_dataset["list_classes"][:]) # the list of classes
train_set_y_orig = train_set_y_orig.reshape((1, train_set_y_orig.shape[0]))
test_set_y_orig = test_set_y_orig.reshape((1, test_set_y_orig.shape[0]))
return train_set_x_orig, train_set_y_orig, test_set_x_orig, test_set_y_orig, classesLet's load the data and check the image for a random index.
train_x_orig, train_y, test_x_orig, test_y, classes = load_dataset()
index = 10
plt.imshow(train_x_orig[index])
print ("y = " + str(train_y[0,index]) + ". It's a " + classes[train_y[0,index]].decode("utf-8") + " picture.")y = 0. It's a non-cat picture.
## Explore the dataset
m_train = train_x_orig.shape[0]
num_px = train_x_orig.shape[1]
m_test = test_x_orig.shape[0]
print ("Number of training examples: " + str(m_train))
print ("Number of testing examples: " + str(m_test))
print ("Each image is of size: (" + str(num_px) + ", " + str(num_px) + ", 3)")
print ("train_x_orig shape: " + str(train_x_orig.shape))
print ("train_y shape: " + str(train_y.shape))
print ("test_x_orig shape: " + str(test_x_orig.shape))
print ("test_y shape: " + str(test_y.shape))Number of training examples: 209
Number of testing examples: 50
Each image is of size: (64, 64, 3)
train_x_orig shape: (209, 64, 64, 3)
train_y shape: (1, 209)
test_x_orig shape: (50, 64, 64, 3)
test_y shape: (1, 50)
# Reshape the training and test examples
train_x_flatten = train_x_orig.reshape(train_x_orig.shape[0], -1).T # The "-1" makes reshape flatten the remaining dimensions
test_x_flatten = test_x_orig.reshape(test_x_orig.shape[0], -1).T
# Standardize data to have feature values between 0 and 1.
train_x = train_x_flatten/255.
test_x = test_x_flatten/255.train_x's shape: (12288, 209)
test_x's shape: (12288, 50)
We will create a function L_layer_model using all the helper functions created above to predict whether an image is of a cat or not.
def L_layer_model(X, Y, layers_dims, learning_rate = 0.0075, num_iterations = 3000, print_cost=False):#lr was 0.009
"""
Implements a L-layer neural network: [LINEAR->RELU]*(L-1)->LINEAR->SIGMOID.
Arguments:
X -- data, numpy array of shape (number of examples, num_px * num_px * 3)
Y -- true "label" vector (containing 0 if cat, 1 if non-cat), of shape (1, number of examples)
layers_dims -- list containing the input size and each layer size, of length (number of layers + 1).
learning_rate -- learning rate of the gradient descent update rule
num_iterations -- number of iterations of the optimization loop
print_cost -- if True, it prints the cost every 100 steps
Returns:
parameters -- parameters learnt by the model. They can then be used to predict.
"""
np.random.seed(1)
costs = [] # keep track of cost
# Parameters initialization.
### START CODE HERE ###
parameters = initialize_parameters_deep(layers_dims)
### END CODE HERE ###
# Loop (gradient descent)
for i in range(0, num_iterations):
# Forward propagation: [LINEAR -> RELU]*(L-1) -> LINEAR -> SIGMOID.
### START CODE HERE ### (≈ 1 line of code)
AL, caches = L_model_forward(X, parameters)
### END CODE HERE ###
# Compute cost.
### START CODE HERE ### (≈ 1 line of code)
cost = compute_cost(AL,Y)
### END CODE HERE ###
# Backward propagation.
### START CODE HERE ### (≈ 1 line of code)
grads = L_model_backward(AL, Y, caches)
### END CODE HERE ###
# Update parameters.
### START CODE HERE ### (≈ 1 line of code)
parameters = update_parameters(parameters, grads, learning_rate)
### END CODE HERE ###
# Print the cost every 100 training example
if print_cost and i % 100 == 0:
print ("Cost after iteration %i: %f" %(i, cost))
if print_cost and i % 100 == 0:
costs.append(cost)
# plot the cost
plt.plot(np.squeeze(costs))
plt.ylabel('cost')
plt.xlabel('iterations (per tens)')
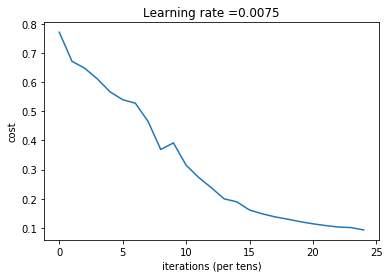
plt.title("Learning rate =" + str(learning_rate))
plt.show()
return parametersLet's create a 5 layer model. Take layers_dims= [12288, 20, 7, 5, 1]
parameters = L_layer_model(train_x, train_y, layers_dims, num_iterations = 2500, print_cost = True)Cost after iteration 0: 0.771749
Cost after iteration 100: 0.672053
Cost after iteration 200: 0.648263
Cost after iteration 300: 0.611507
Cost after iteration 400: 0.567047
Cost after iteration 500: 0.540138
Cost after iteration 600: 0.527930
Cost after iteration 700: 0.465477
Cost after iteration 800: 0.369126
Cost after iteration 900: 0.391747
Cost after iteration 1000: 0.315187
Cost after iteration 1100: 0.272700
Cost after iteration 1200: 0.237419
Cost after iteration 1300: 0.199601
Cost after iteration 1400: 0.189263
Cost after iteration 1500: 0.161189
Cost after iteration 1600: 0.148214
Cost after iteration 1700: 0.137775
Cost after iteration 1800: 0.129740
Cost after iteration 1900: 0.121225
Cost after iteration 2000: 0.113821
Cost after iteration 2100: 0.107839
Cost after iteration 2200: 0.102855
Cost after iteration 2300: 0.100897
Cost after iteration 2400: 0.092878
pred_train = predict(train_x, train_y, parameters)Accuracy: 0.9856459330143539
pred_test = predict(test_x, test_y, parameters)Accuracy: 0.8