A tool for the recovery of unassembled telomeres from soft-clipped read alignments.
Current version: 0.0.3
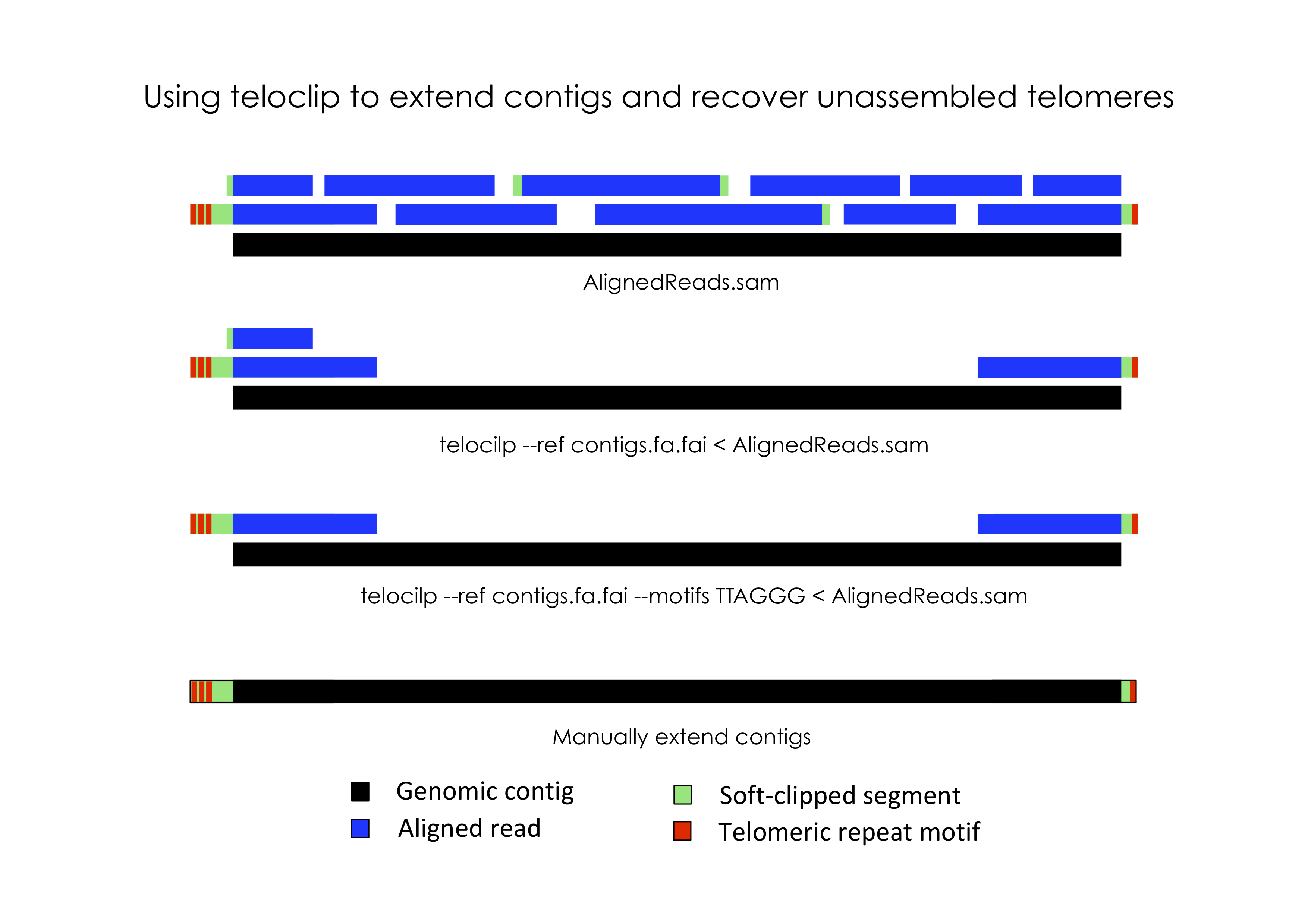
In most eukaryotic species, chromosomes terminate in repetitive telomeric sequences. A complete genome assembly should ideally comprise chromosome-level contigs that possess telomeric repeats at each end. However, genome assemblers frequently fail to recover these repetitive features, instead producing contigs that terminate immediately prior to their location.
Teloclip is designed to recover long-reads that can be used to extend draft contigs and resolve missing telomeres (short-read alignments may also be processed with teloclip). It does this by searching alignments of raw long-read data (i.e. Pacbio or ONP reads mapped with Minimap2) for 'clipped' alignments that occur at the ends of draft contigs. A 'clipped' alignment is produced where the end of a read is not part of its best alignment. This can occur when a read extends past the end of an assembled contig.
Information about segments of a read that were aligned or clipped are stored in SAM formatted alignments as a CIGAR string. Teloclip parses these strings to determine if a read has been clipped at one or both ends of a contig.
Optionally, teloclip can screen overhanging reads for telomere-associated motifs (i.e. 'TTAGGG' / 'CCCTAA') and report only those containing a match.
Teloclip is based on concepts from Torsten Seemann's excellent tool samclip. Samclip can be used to remove clipped alignments from a samfile prior to variant calling.
Teloclip hex-sticker was designed by @Super_Coleider.
Requires Python => v3.6
Clone from this repository and install as a local Python package.
% git clone https://github.com/Adamtaranto/teloclip.git && cd teloclip && pip install -e .Install from PyPi.
% pip install teloclipInstall from Bioconda.
% conda install -c bioconda teloclipTest installation.
# Print version number and exit.
% teloclip --version
teloclip 0.0.3
# Get usage information
% teloclip --helpFirst index the reference assembly
# Create index of reference fasta
% samtools faidx ref.fa
Reading alignments from SAM file
# Read input from file and write output to stout
% teloclip --ref ref.fa.fai in.sam
# Read input from stdin and write stdout to file
% teloclip --ref ref.fa.fai < in.sam > out.sam
# Filter alignments from BAM file, write sorted output to file
% samtools view -h in.bam | teloclip --ref ref.fa.fai | samtools sort > out.bam
Streaming SAM records from aligner
# Map PacBio long-reads to ref assembly, filter for alignments clipped at contig ends, write to sorted bam
% minimap2 -ax map-pb ref.fa pacbio.fq.gz | teloclip --ref ref.fa.fai | samtools sort > out.bam
# Map reads, exclude unmapped reads and non-primary/supplementary alignments. Report clipped alignments as sorted bam.
% minimap2 -ax map-pb ref.fa pacbio.fq.gz | samtools view -h -F 0x2308 | teloclip --ref ref.fa.fai | samtools sort > out.bam
Report clipped alignments containing target motifs
# Report alignments which are clipped at a contig end
# AND contain >=1 copy of the telomeric repeat "TTAGGG" (or its reverse complement "CCCTAA") in the clipped region.
% samtools view -h in.bam | teloclip --ref ref.fa.fai --motifs TTAGGG | samtools sort > out.bam
# Report alignments which are clipped at a contig end
# AND contain >=1 copy of the telomeric repeat "TTAGGG" (or its reverse complement "CCCTAA") ANYWHERE in the read.
% samtools view -h in.bam | teloclip --ref ref.fa.fai --motifs TTAGGG --matchAny | samtools sort > out.bam
# Compress homopolymers in query motifs and clipped regions to compensate for errors in raw PacBio or ONP data.
# i.e. The motif 'TTAGGGTTAGGG' becomes 'TAGTAG' and will match 'TTTTTAAAGGTTTAAGGG'.
% samtools view -h in.bam | teloclip --ref ref.fa.fai --motifs TTAGGGTTAGGGTTAGGGTTAGGGTTAGGG | samtools sort > out.bam
Extract clipped reads
# Find clipped alignments containing motif 'TTAGGG' and write reads to separate fasta files for each reference contig end.
# Clipped region of each read is masked as lowercase in output fasta files.
% samtools view -h in.bam | teloclip --ref ref.fa.fai --motifs TTAGGG | teloclip-extract --refIdx data/test.fna.fai --extractReads --extractDir SplitOverhangs
Additional filters
-
Consider pre-filtering alignments with "samtools view" to remove non-primary / low quality alignments.
-
Users may wish to exclude reads below a minimum length or read quality score to reduce the risk of incorrect alignments.
-
In some cases it may be useful to prioritise primary alignments that do not have the secondary alignment "SA" tag set.
# Exclude secondary alignments and primary alignments with SA tag set. % samtools view -h -F 0x100 in.sam | awk '!/SA:/ {print $0;}' | teloclip --ref ref.fa.fai > noSA.sam
Pre-corrected Data
Users may find improved specificity of alignments using pre-corrected long-read data.
- The genome assembler Canu preforms pre-correction of long-reads through iterative overlapping and correction prior to assembly. Corrected reads are trimmed based on coverage to remove low-confidence ends. The corrected reads are stored by Canu as PREFIX.correctedReads.fasta.gz, and the trimmed corrected reads as PREFIX.trimmedReads.fasta.gz.
- Single molecule long-reads can also be corrected with tools such as LoRDEC or HALC.
Note: Long reads may loose ends containing telomeres as a result of trimming. Give it a go, try the raw reads if unsuccessful.
Extending contigs
- Before using terminal alignments identified by teloclip to extend contigs, inspect alignments in a genome browser that displays information about clipped reads, such as IGV. Check for conflicting clipped sequences.
- After manually extending contigs the revised assembly should be re-polished using available long and short read data to correct indels present in the raw long-reads.
- Validate the final assembly by re-mapping long-read data and checking for alignments that extend into revised contig ends.
Illumina data
- Teloclip will also work fine with aligned short read data, which has a far lower error rate than single-molecule long-read data. However, there are obvious limits to the distance that a contig may be extended with shorter reads.
Hybridising existing assemblies
- You may have assemblies for your genome generated with different assemblers/configurations or data types (i.e. 10X linked-reads, Illumina, PacBio, ONP) which vary in their success in assembling individual telomeres.
- These alternative assemblies can be treated as long reads and aligned to a reference using Minimap2. Teloclip will identify aligned contigs that can be used to extend those in the reference set.
- Be cautious of short contigs that may align to may repetative sub-telomeric regions.
Align alternative assembly contigs to reference and report overhang alignments.
% minimap2 -ax asm5 ref.fa asm.fa | samtools view -h -F 0x2308 | teloclip --ref ref.fa.fai | samtools sort > asm2ref.bam
Circularising Mitochondrial / Bacterial genomes
- Using default settings, teloclip will report alignments with clipped regions extending past linear contig ends.
- Reads can be extracted from these alignments using circlator's bam2reads and re-aligned to an assembly graph in Bandage to help identify uncircularised contigs.
Run teloclip --help to view the programs' most commonly used options:
Usage: teloclip [-h] [--version] --refIdx REFIDX [--minClip MINCLIP] [--maxBreak MAXBREAK]
[--motifs MOTIFS] [--noRev NOREV] [--noPoly NOPOLY] [--matchAny MATCHANY]
[samfile]
Required:
--refIdx REFIDX Path to fai index for reference fasta. Index fasta using `samtools faidx FASTA`
Positional arguments:
samfile Input SAM can be added as the first positional argument after flagged options.
If not set teloclip will read from stdin.
Optional:
--minClip Require clip to extend past ref contig end by at least N bases.
Default: 1
--maxBreak Tolerate max N unaligned bases at contig ends.
Default: 50
--motifs If set keep only reads containing given motif/s from a comma delimited list
of strings. By default also search for reverse complement of motifs.
i.e. TTAGGG,TTAAGGG will also match CCCTAA,CCCTTAA
Default: None
--noRev If set do NOT search for reverse complement of specified motifs.
Default: Find motifs on both strands.
--noPoly If set collapse homopolymer tracks within motifs before searching overhangs.
i.e. "TTAGGGTTAGGGTTAGGGTTAGGGTTAGGG" -> "TAGTAGTAGTAGTAG".
Useful for PacBio or ONP long reads homopolymer length errors. Defaut: Off.
--matchAny If set motif match may occur in unclipped region of alignment.
Defaut: False
--version Show program's version number and exit.
Run teloclip-extract --help to view the programs' most commonly used options:
Usage: teloclip-extract [-h] --refIdx REFIDX [--prefix PREFIX]
[--extractReads] [--extractDir EXTRACTDIR]
[--minClip MINCLIP] [--maxBreak MAXBREAK] [--version]
[samfile]
positional arguments:
samfile If not set, will read sam from stdin.
optional arguments:
-h, --help Show this help message and exit
--refIdx Path to fai index for reference fasta. Index fasta
using `samtools faidx FASTA`
--prefix Use this prefix for output files. Default: None.
--extractReads If set, write overhang reads to fasta by contig.
--extractDir
Write extracted reads to this directory. Default: cwd.
--minClip Require clip to extend past ref contig end by at least
N bases.
--maxBreak Tolerate max N unaligned bases at contig ends.
--version Show program's version number and exit
Submit feedback to the Issue Tracker
Software provided under MIT license.