A tiny package for structure analysis of macromolecules. Atomic coordinates retrieved from a PDB file are stored in two formats: list and dictionary. So you can create your own methods on top of either format. This package provides structure analysis capability based on distance matrix.
pip install git+https://github.com/carbonscott/pyrotein --upgrade --user
If upgrading doesn't work as expected, run pip uninstall pyrotein before
installing it.
This package has only one dependency -- numpy.
You can create graphics using your preferred visualization tools. For
simplicity, it's a design choice not to include a specific visualization library
in pyrotein. However, examples of using Gnuplot and matplotlib are
included in the examples directory.
The colorschemes used in RMSD distance matrix and SVD analysis are inspired by this paper from Zhong Ren.
import pyrotein as prThe following code snippet loads a PDB file 6cmo.pdb under pdb directory.
import pyrotein as pr
import os
# Read coordinates from a PDB file...
drc = "pdb"
pdb = "6cmo"
fl_pdb = f"{pdb}.pdb"
pdb_path = os.path.join(drc, fl_pdb)
atoms_pdb = pr.atom.read(pdb_path)The method pr.atom.read returns molecular information encapsulated in a Python
List. However, a lookup table can be very handy for tasks such as accesssing a
particular atom CA from residue 1002 in chain A. The following example
shows how to achieve it using lookup table.
# Create a lookup table for this pdb...
atom_dict = pr.atom.create_lookup_table(atoms_pdb)
# Demo: Access atom `CA` from residue 1002 in chain A
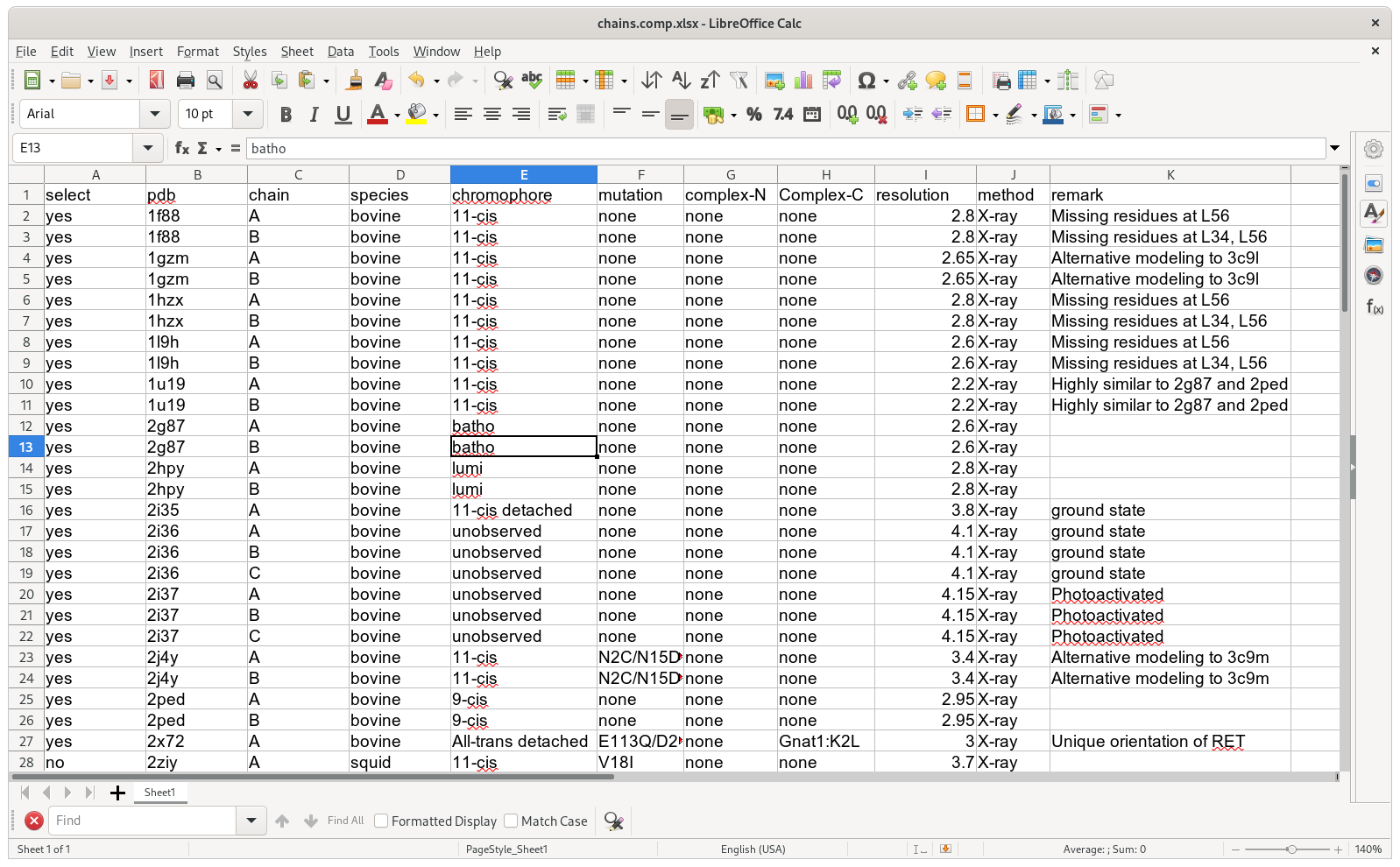
atom_dict["A"][1002]["CA"]A metadata table keeps track of protein information (PDB, ligand, method, etc).
It is vital in understanding clustering in the SVD analysis stage. Meanwhile,
the metadata table should be computer readable. For example, I keep metadata in
a xlsx file that can be read by a Python module openpyxl. If you decide to
use the same module, install it by pip install openpyxl --user. Check out
loaddata.py in examples directory to see how to load data from an xlsx
using openpyxl.
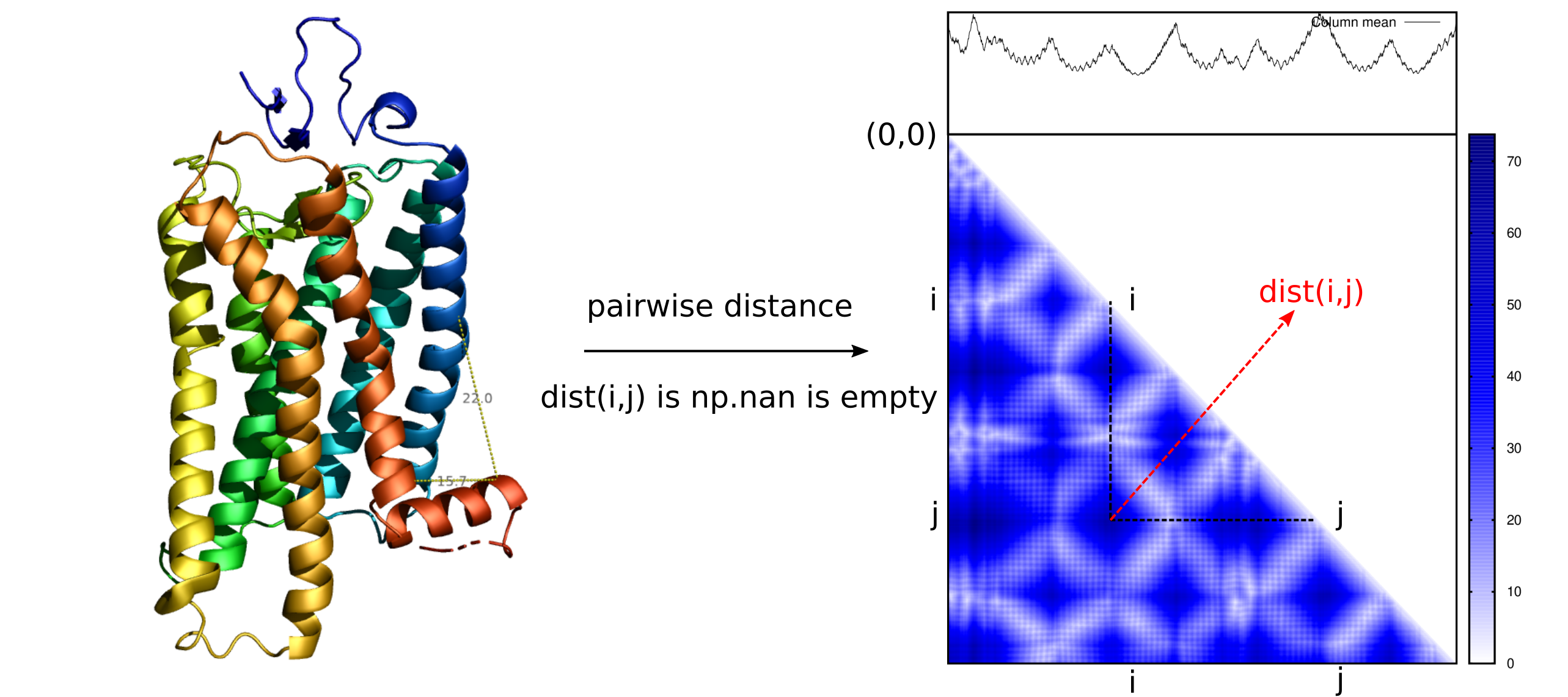
pyrotein is initially designed to perform distance matrix analysis of protein
structures. Distance matrix encodes pairwise atomic distance found in a protein
structure.
import os
import numpy as np
import pyrotein as pr
from loaddata import load_xlsx, label_TMs
from display import plot_dmat
# Specify chains to process...
fl_chain = "chains.comp.xlsx"
lines = load_xlsx(fl_chain)
drc = "pdb"
# Define atoms used for distance matrix analysis...
backbone = ["N", "CA", "C", "O"]
# Specify the range of atoms from adrenoceptor...
nterm = 1
cterm = 322
# The first element is to facilitate the indexing during assignment
len_segments = [ 0,
cterm - nterm + 1,
]
len_peptide = np.sum(len_segments) * len(backbone)
drc_dmat = "dmats"
pal = "set palette defined ( 0 '#F6FF9E', 0 'white', 0.5 'blue', 1 'navy' )"
for i_fl, line in enumerate(lines[-1]):
# Unpack parameters
_, pdb, chain, species = line[:4]
betatype = line[10]
# Read coordinates from a PDB file...
fl_pdb = f"{pdb}.pdb"
pdb_path = os.path.join(drc, fl_pdb)
atoms_pdb = pr.atom.read(pdb_path)
# Create a lookup table for this pdb...
atom_dict = pr.atom.create_lookup_table(atoms_pdb)
# Obtain the chain to process...
chain_dict = atom_dict[chain]
# Obtain coordinates...
xyzs = pr.atom.extract_xyz_by_atom(backbone, chain_dict, nterm, cterm)
# Calculate distance matrix...
dmat = pr.distance.calc_dmat(xyzs, xyzs)
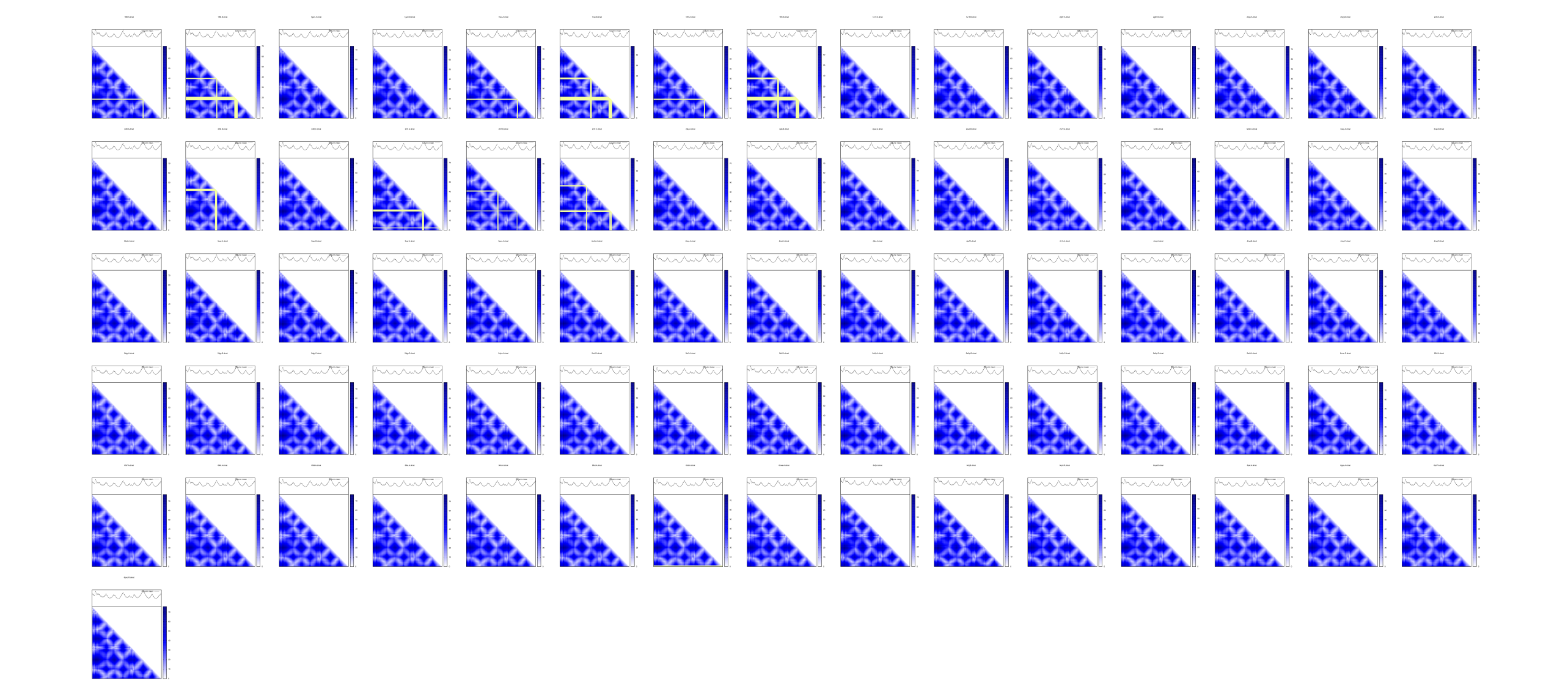
# It is a common practice to visualize a few distance matrix.
# Here is just a way how I use Gnuplot to do so, but you can use
# your favourite tool to visualize it.
fl_dmat = os.path.join(drc_dmat, f"{pdb}.{chain}.dmat")
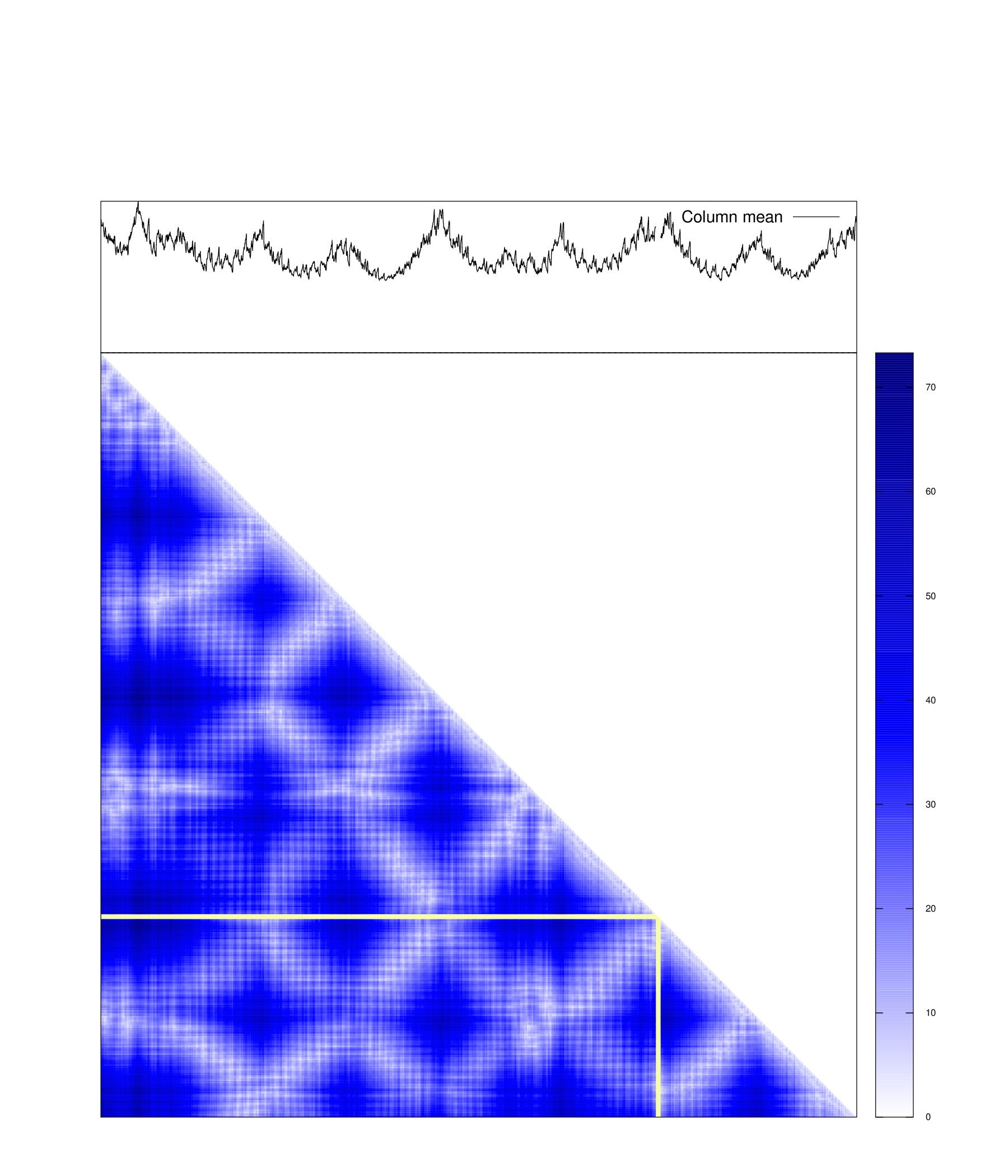
plot_dmat(dmat, fl_dmat, lbl = {}, palette = pal, NaN = 0)If all atoms in main chain and side chain are your interests, pyrotein is
capable of extracting coordinates and build up data correspondence by means of
sequence alignment result (You have to be careful about the result of sequecne
alignment. It can be problematic). Sequecne related functionalities are
supposed to be found in pyrotein.fasta submodule.
import os
import numpy as np
import pyrotein as pr
from loaddata import load_xlsx
from display import plot_dmat
# [[[ OBTAIN THE CONSENSUS SEQUENCE ]]]
# Read the sequence alignment result...
# [WARNING] !!!sequence alignment is not trustworthy
fl_aln = 'seq.align.fasta'
seq_dict = pr.fasta.read(fl_aln)
# Obtain the consensus sequence (super seq)...
tally_dict = pr.fasta.tally_resn_in_seqs(seq_dict)
super_seq = pr.fasta.infer_super_seq(tally_dict)
# [[[ FIND SIZE OF DISTANCE MATRIX ]]]
# Get the sequence index (alignment) on the n-term side...
nseqi = pr.fasta.get_lseqi(super_seq)
# User defined range...
nterm, cterm = 1, 322
len_seg = cterm - nterm + 1
super_seg = super_seq[nseqi : nseqi + len_seg]
# Load constant -- atomlabel...
label_dict = pr.atom.constant_atomlabel()
aa_dict = pr.atom.constant_aminoacid_code()
# [[[ ANALYZE PDB ENTRIES ]]]
# Specify chains to process...
fl_chain = "chains.comp.xlsx"
lines = load_xlsx(fl_chain, sheet = "Sheet1")
drc = "pdb"
drc_dmat = "dmats.full"
pal = "set palette defined ( 0 '#F6FF9E', 0 'white', 0.5 'blue', 1 'navy' )"
for i_fl, line in enumerate(lines[-1:]):
# Unpack parameters
_, pdb, chain, _ = line[:4]
# Read coordinates from a PDB file...
fl_pdb = f"{pdb}.pdb"
pdb_path = os.path.join(drc, fl_pdb)
atoms_pdb = pr.atom.read(pdb_path)
# Create a lookup table for this pdb...
atom_dict = pr.atom.create_lookup_table(atoms_pdb)
# Obtain the chain to process...
chain_dict = atom_dict[chain]
# Obtain the target protein by range...
tar_seq = seq_dict[f"{pdb}_{chain}"]
tar_seg = tar_seq[nseqi : nseqi + len_seg]
# Standardize sidechain atoms...
pr.atom.standardize_sidechain(atom_dict)
# Obtain coordinates...
xyzs = pr.atom.extract_xyz_by_atom(tar_seg, super_seg, chain_dict, nterm, cterm)
# Calculate distance matrix...
dmat = pr.distance.calc_dmat(xyzs, xyzs)
fl_dmat = os.path.join(drc_dmat, f"{pdb}.{chain}.dmat")
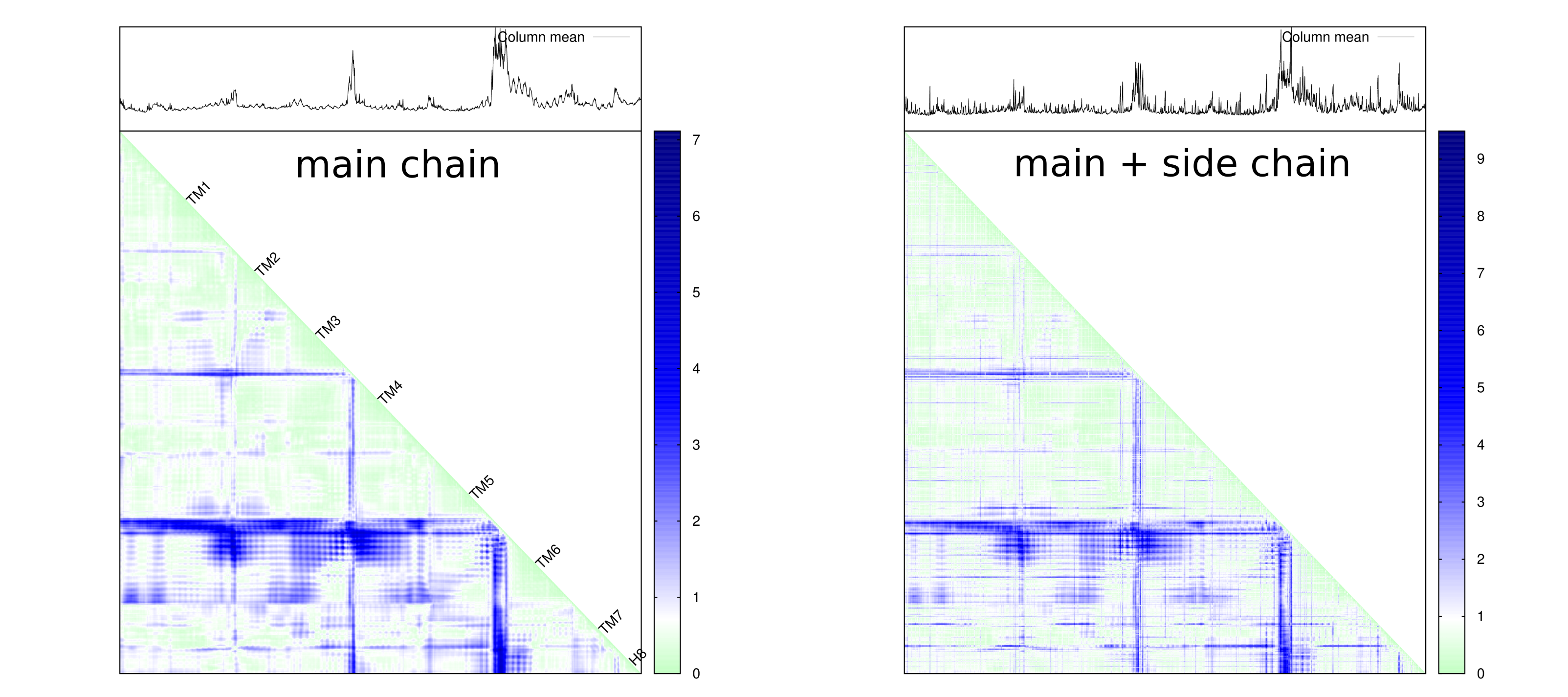
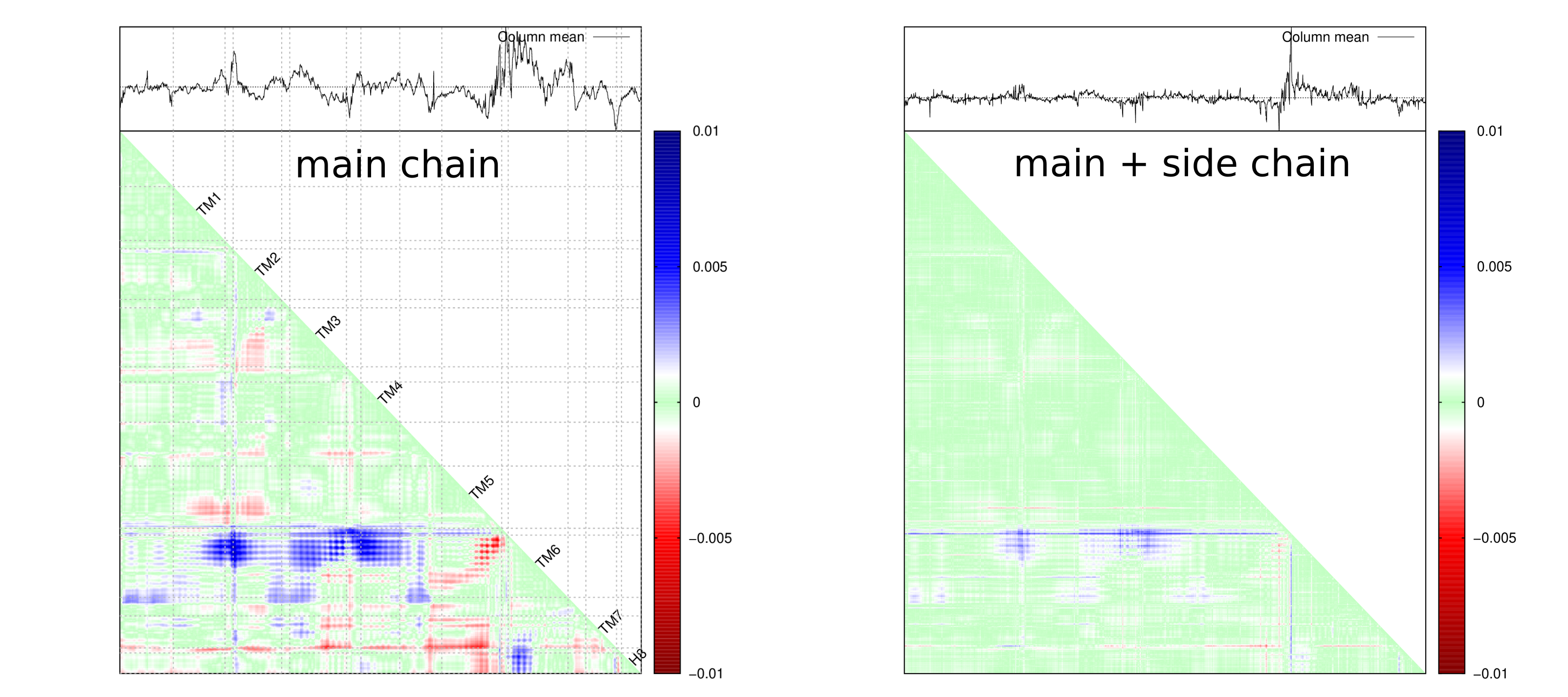
plot_dmat(dmat, fl_dmat, lbl = {}, palette = pal, NaN = 0)Distance matrix that depict both main chain and side chain.
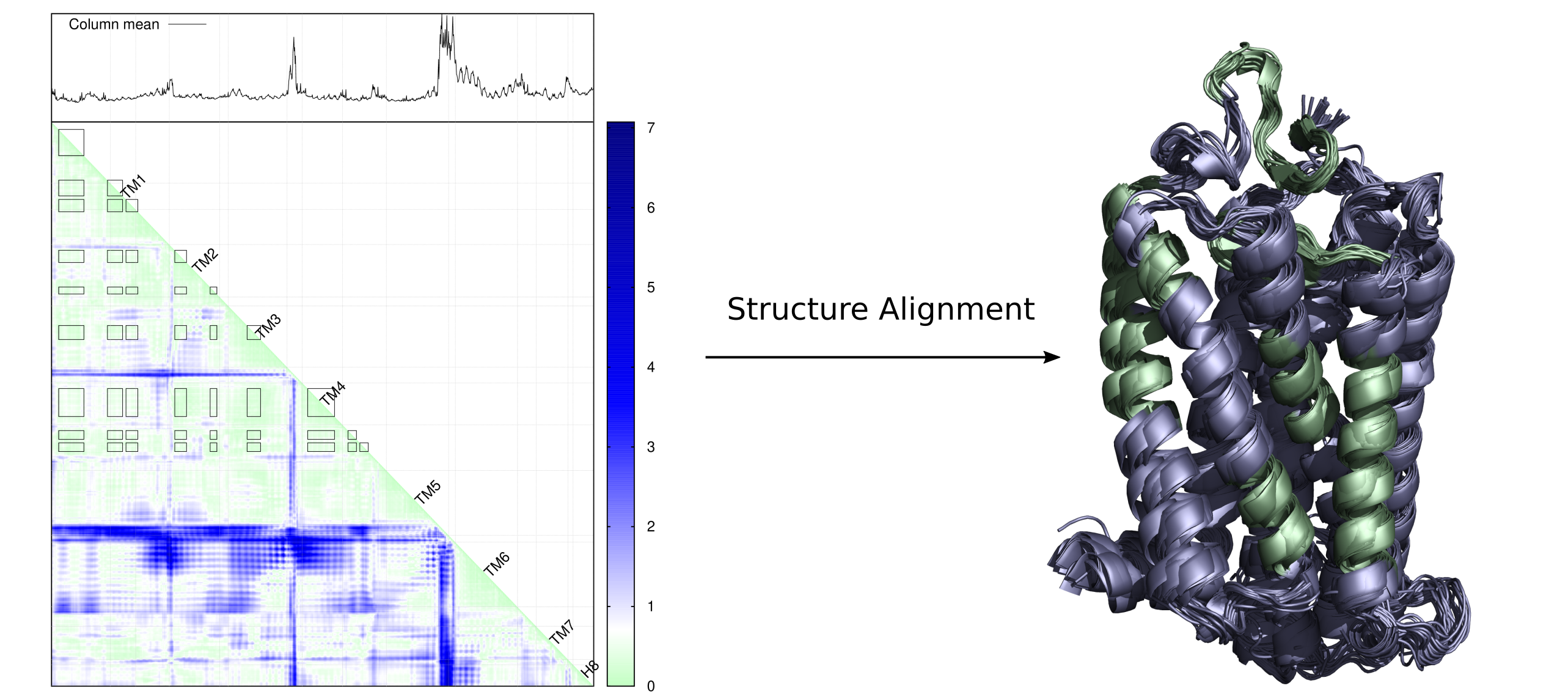
What can we do by putting all distance matrix together like below?
If you consider a distance matrix is a picture full of pixels, RMSD distance matrix encodes RMSD of all pixel values at a specific location. RMSD distance matrix enables the understanding of rigidity of a protein, that is to say, smaller deviation means more rigid, and vice versa. So it can provide a so-called rigid protein framework for structure alignment.
Of course, the analysis can be carried out on both main chain only or main + side chain scenarios. You can see how much details are captured while considering both main chain and side chain for each residue.
A significant protein structure analysis pyrotein enables is SVD analysis. An
example code to enable it:
import numpy as np
import pyrotein as pr
import os
from loaddata import load_xlsx
# [[[ OBTAIN THE CONSENSUS SEQUENCE ]]]
# Read the sequence alignment result...
# [WARNING] !!!sequence alignment is not trustworthy, need to check manually
fl_aln = 'seq.align.fasta'
seq_dict = pr.fasta.read(fl_aln)
# Obtain the consensus sequence (super seq)...
tally_dict = pr.fasta.tally_resn_in_seqs(seq_dict)
super_seq = pr.fasta.infer_super_seq(tally_dict)
# [[[ FIND SIZE OF DISTANCE MATRIX ]]]
# Get the sequence index (alignment) on the n-term side...
nseqi = pr.fasta.get_lseqi(super_seq)
# User defined range...
nterm, cterm = 1, 322
len_seg = cterm - nterm + 1
super_seg = super_seq[nseqi : nseqi + len_seg]
# Load constant -- atomlabel...
label_dict = pr.atom.constant_atomlabel()
aa_dict = pr.atom.constant_aminoacid_code()
# Calculate the total length of distance matrix...
len_dmat = np.sum( [ len(label_dict[aa_dict[i]]) for i in super_seg ] )
# [[[ ANALYZE PDB ENTRIES ]]]
# Specify chains to process...
fl_chain = "chains.comp.xlsx"
lines = load_xlsx(fl_chain, sheet = "Sheet1")
drc = "pdb"
## dmats = np.zeros((len(lines), len_dmat, len_dmat))
len_lower_tri = (len_dmat * len_dmat - len_dmat) // 2
dmats = np.zeros((len(lines), len_lower_tri))
# Process each entry...
for i_fl, line in enumerate(lines):
# Unpack parameters
_, pdb, chain, species = line[:4]
print(f"Processing {pdb}_{chain}")
# Read coordinates from a PDB file...
fl_pdb = f"{pdb}.pdb"
pdb_path = os.path.join(drc, fl_pdb)
atoms_pdb = pr.atom.read(pdb_path)
# Create a lookup table for this pdb...
atom_dict = pr.atom.create_lookup_table(atoms_pdb)
# Obtain the chain to process...
chain_dict = atom_dict[chain]
# Obtain the target protein by range...
tar_seq = seq_dict[f"{pdb}_{chain}"]
tar_seg = tar_seq[nseqi : nseqi + len_seg]
# Standardize sidechain atoms...
pr.atom.standardize_sidechain(atom_dict)
# Obtain coordinates...
xyzs = pr.atom.extract_xyz_by_seq(tar_seg, super_seg, chain_dict, nterm, cterm)
# Calculate distance matrix...
dmat = pr.distance.calc_dmat(xyzs, xyzs)
# Convert dmat into one-dimensional array and keep it in dmats...
dmats[i_fl, :] = pr.utils.mat2tril(dmat, offset = -1)
# Replace np.nan with mean across samples...
pr.utils.fill_nan_with_mean(dmats.T, axis = 1)
# SVD...
# Column as example
# Row as feature
u, s, vh = np.linalg.svd( dmats.T, full_matrices = False )
# Export data for downstream analysis...
np.save("dmats.full.npy" , dmats)
np.save("u.full.npy" , u)
np.save("s.full.npy" , s)
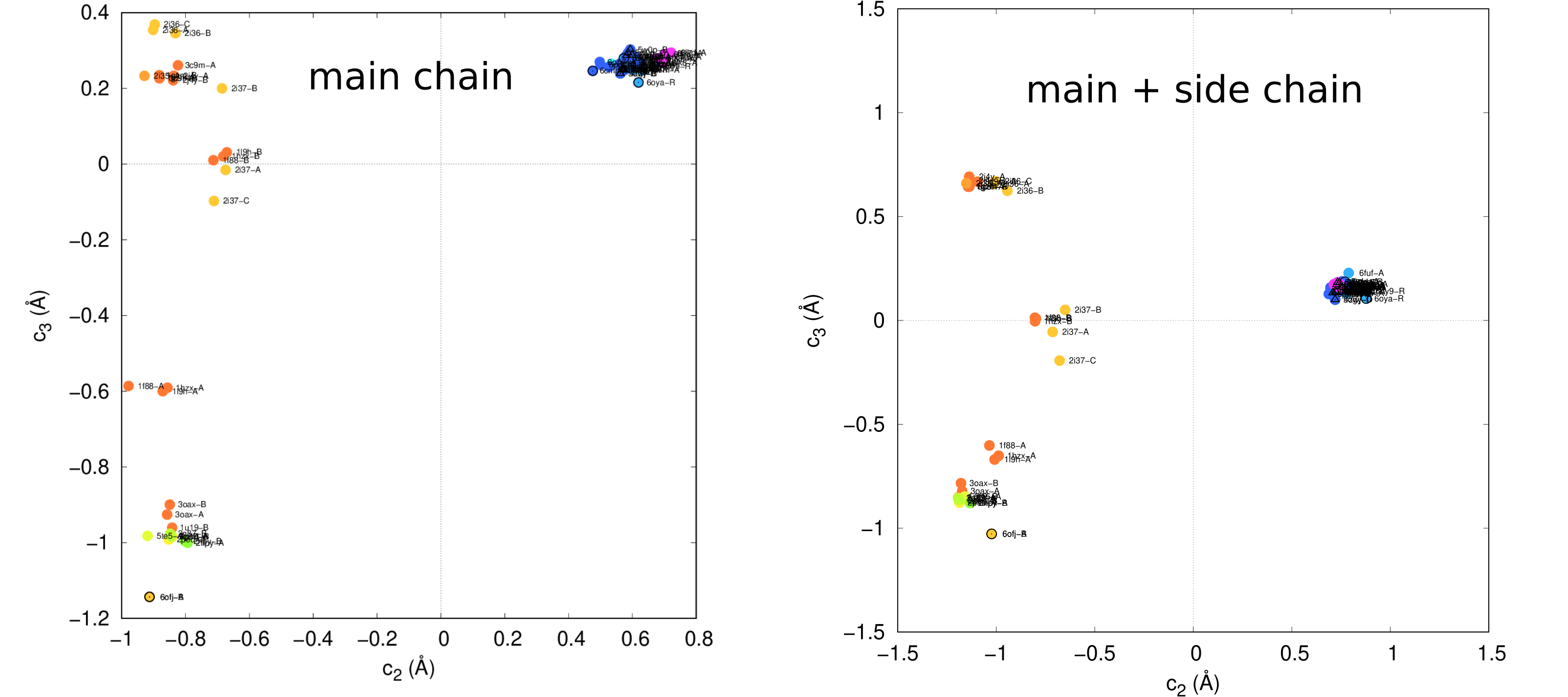
np.save("vh.full.npy", vh)A comparison of left singular values is shown below.
It also distinguishes entries in the analysis better in main + side chain scenario. Check out the follow scatter plot. (Rotation might require to make them look better.)
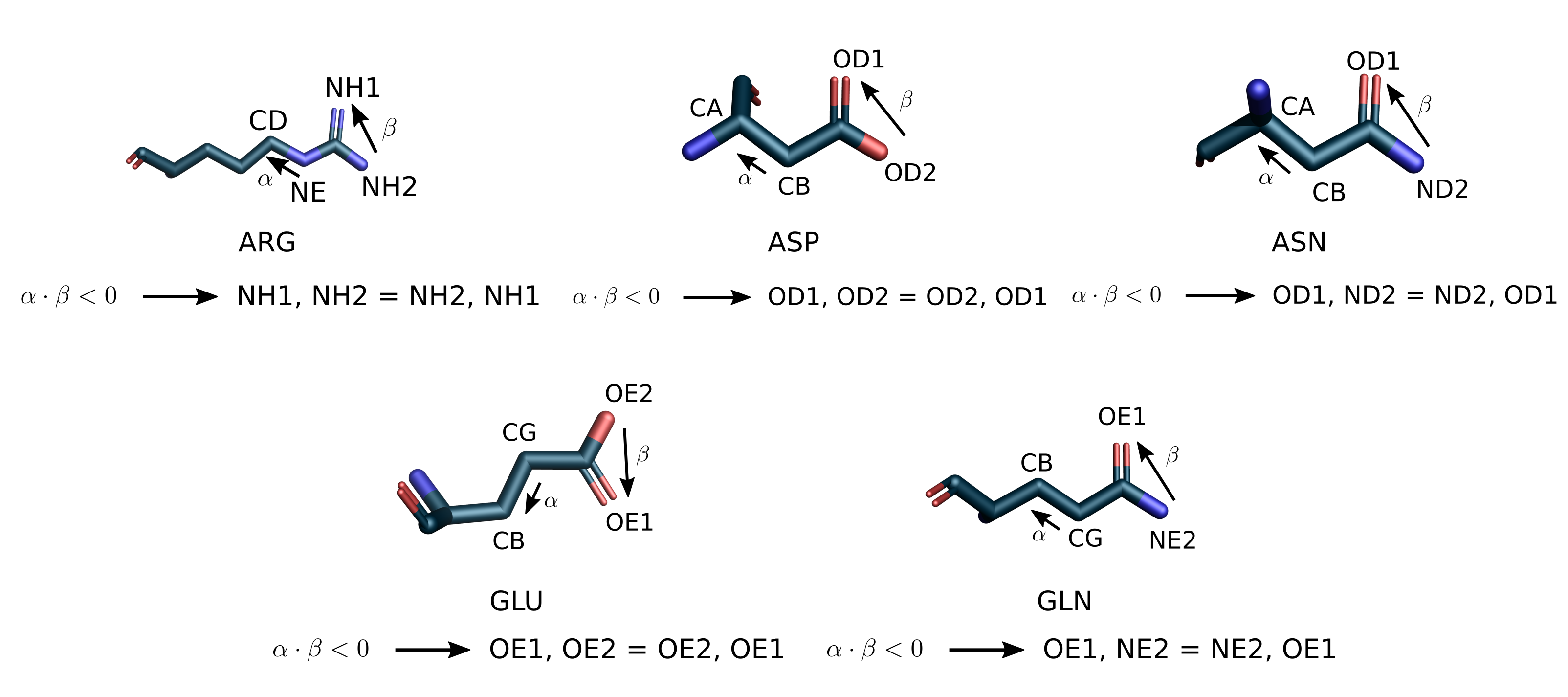
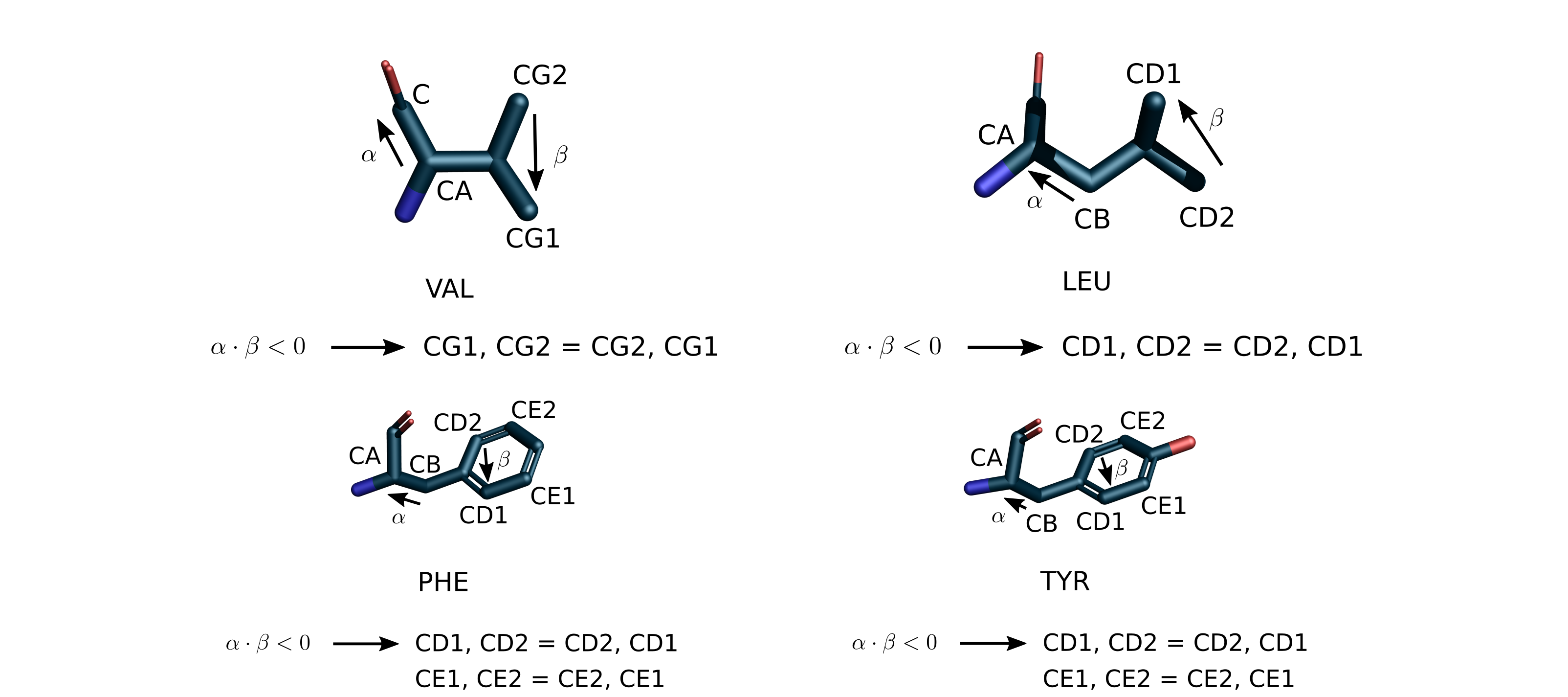
Sidechain atoms can be interchangeable, such as NH1 and NH2 in ARG, OD1
and ND2 in ASN. The figure below illustrates the possible ambiguous
scenarios and specifies the swapping rules leading to a standard ordering.
Sample code to consider sidechain atom standardization.
import pyrotein as pr
import numpy as np
import os
# Read atomic information...
pdb = "1f88"
chain = "A"
drc_pdb = "pdb"
fl_pdb = f"{pdb}.pdb"
path_pdb = os.path.join(drc_pdb, fl_pdb)
atom_list = pr.atom.read(path_pdb)
atom_dict = pr.atom.create_lookup_table(atom_list)
# Standardize sidechain atoms...
pr.atom.standardize_sidechain(atom_dict)The examples directory contains two examples about distance matrix and RMSD
distance matrix. Two visualization choices are provided via Gnuplot and
matplotlib.
cat pdb/*.fasta > step1.all.fasta./fasta.step1.pyto extract chains defined inxxx.db.xlsx; It returnsstep2.interest.fastacat step2.interest.fasta | pbcopyand paste it into Clustal Omegafasta.check.pyto check if the defaultnseqiandcseqiare good enough otherwise determinenseqiandcseqimanually- Check if terribly placed insertion will confuse sequence alignment.
- Clustal Omega might return residue 'X'. Repalce it with '-'. Otherwise, the string-match-based method to identify lower bound residue ID will fail.
cseqican be manually determined if the defaultcseqiis too large.- PyMol fetched
fastafile might contain extra residue that might not have a coordinate available in PDB. Replace those residue with '-' too.
The warning RuntimeWarning: Mean of empty slice is triggered by np.nanmean
when the input array has nothing but np.nan values.
If RMSD analysis gives very smeared intensities, you need to check sequence
similarities. How different are they? In groups or just a few outliers.
Does the sequence alignment make sense? (It doesn't do a good job for a minority
of entries that have a long insertion.)