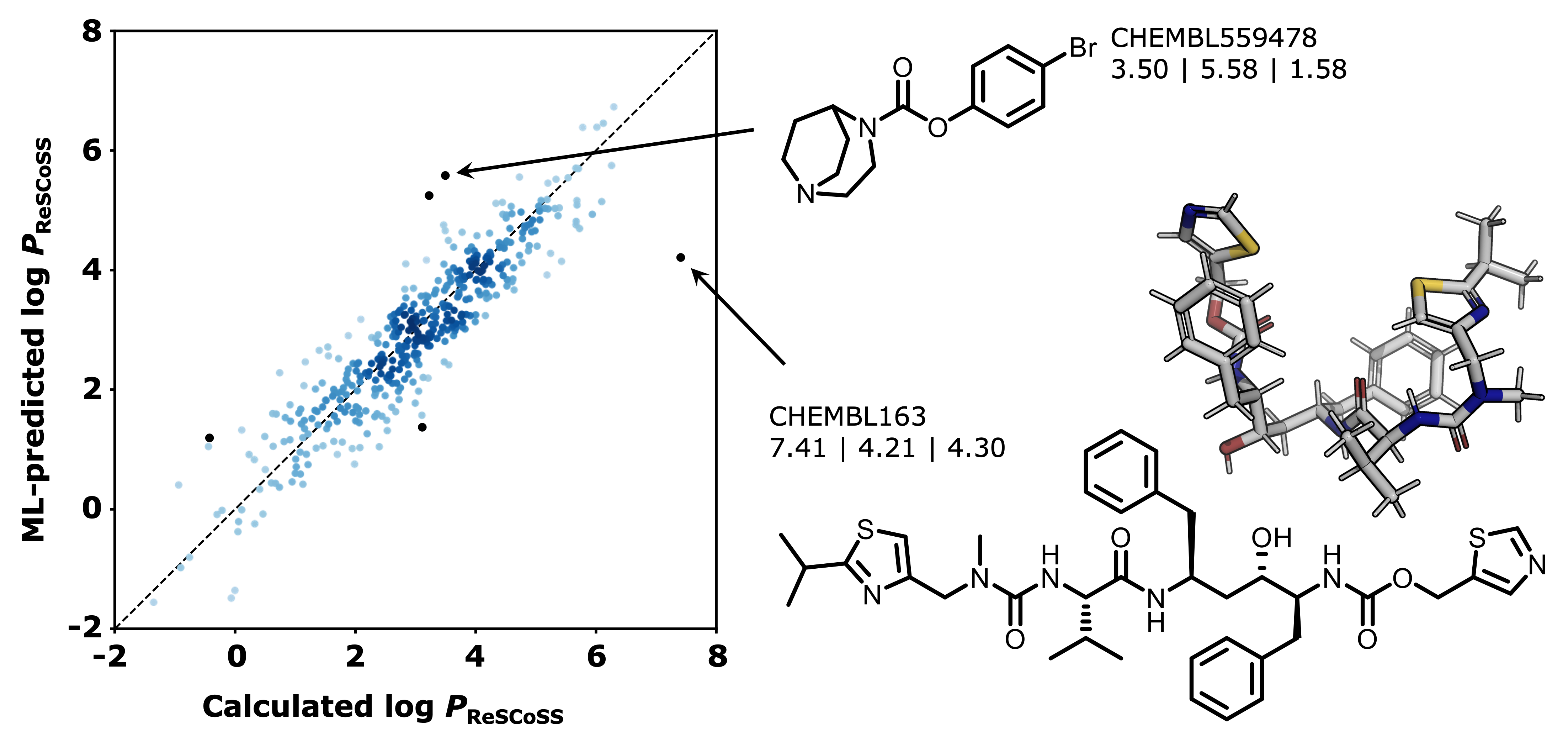
rescoss_logp_ml: Machine intelligence models for fast, quantum mechanics-based approximation of drug lipophilicity
Machine-learning models to predict ReSCoSS-calculated [1] log P values for drug-like molecules.
Details are described in our paper. Please cite if you use this work.
To setup and test everything:
git clone https://github.com/cisert/rescoss_logp_ml.git
cd rescoss_logp_ml
export PYTHONPATH="$(pwd):$PYTHONPATH"
makeTo use pre-trained models to make predictions:
from rescoss_logp_ml.predict_with_trained_model import predict_with_trained_model
from rescoss_logp_ml.utils import ROOT_PATH, MODEL_FILENAMES
import os
smiles_list = [
"O=C(C)Oc1ccccc1C(=O)O", # Aspirin
"O=C(O[C@H]1CCOC1)N[C@@H](Cc2ccccc2)[C@H](O)CN(CC(C)C)S(=O)(=O)c3ccc(N)cc3", # Amprenavir
"Cc1cccc(c1NC(=O)c2cnc(s2)Nc3cc(nc(n3)C)N4CCN(CC4)CCO)Cl", # Dasatinib
]
model_name = "chemprop" # alternatives: "rf", "lasso", "xgb"
split_type = "scaffold" # alternatives: "random"
target = "logp_rescoss" # alternatives: "logp_exp"
model_savepath = os.path.join(
ROOT_PATH,
"saved_models",
"az_set",
split_type,
target,
MODEL_FILENAMES[model_name]
)
preds = predict_with_trained_model(
smiles_list,
model_name=model_name,
target=target,
model_savepath=model_savepath
)
print(preds)See the Makefile for training/learning curve generation to replicate our results.
[1] Udvarhelyi, A., Rodde, S. & Wilcken, R. ReSCoSS: a flexible quantum chemistry workflow identifying relevant solution conformers of drug-like molecules. J. Comput. Aided. Mol. Des. 35, 399–415 (2021).