This package implements the LUC Calculus for reasoning about land use change trajectories. Based on a set of classified time series, we build expressions to answer specific questions, such as Which events of "Forest" areas were replaced by "Pasture"?
With package "lucCalculus" is possible to build questions using Allen's interval temporal logic relationships and also others extended from their study. I suggest the reader read (Allen 1983) and (Allen 1984) for more details. Besides, is possible to generate graphics with event information and plot maps with results. Using these events the user can to perform analysis on time series data to discover important land use changes.
- Git
- R
- Rstudio
- A set of classified GeoTIFF images by year
- The lucCalculus requires "devtools" package is available.
- Open RStudio
- Install devtools
install.packages("devtools") - Load devtools
library(devtools) - Install the lucCalculus package
install_github("e-sensing/lucCalculus")
-
This example was perfomed in Itanhanga municipality, in Mato Grosso state, Brazil.
-
Load the lucCalculus package
library(lucCalculus) -
Create a RasterBrick from a set of classified images
library(lucCalculus)
options(digits = 12)
#-----------------------
# 0. Open images and create a RasterBrick
#-----------------------
# create a RasterBrick from individual raster GeoTIFF classified previously
lucC_create_RasterBrick(path_open_GeoTIFFs = c(system.file("extdata/raster/rasterItanhanga", package = "lucCalculus")),
path_save_RasterBrick = getwd())
# ------------- define variables to use in metadata -------------
# open files
file <- paste0(getwd(),"/rasterItanhanga.tif")
file
# create timeline with classified data from SVM method
timeline <- c("2001-09-01", "2002-09-01", "2003-09-01", "2004-09-01", "2005-09-01", "2006-09-01",
"2007-09-01", "2008-09-01", "2009-09-01", "2010-09-01", "2011-09-01", "2012-09-01",
"2013-09-01", "2014-09-01", "2015-09-01", "2016-09-01")
timeline
# new variable with raster object
rb_class <- raster::brick(file)
# ------------- define variables to plot raster -------------
# original label - see QML file, same order
label <- c("Degradation", "Double_cropping", "Single_cropping", "Forest", "Pasture", "Pasture",
"Pasture", "Double_cropping", "Double_cropping", "Double_cropping", "Double_cropping",
"Double_cropping", "Single_cropping", "Single_cropping", "Water", "Water")
label
# original colors set - see QML file, same order
colors_1 <- c("#BEEE53", "#cd6155", "#e6b0aa", "#228b22", "#7ecfa4", "#afe3c8", "#64b376",
"#e1cdb6", "#b6a896", "#b69872", "#b68549", "#9c6f38", "#e5c6a0", "#e5a352",
"#0000ff", "#3a3aff")
colors_1
# plot rasterBrick
lucC_plot_raster(raster_obj = rb_class,
timeline = timeline, label = label,
custom_palette = TRUE, RGB_color = colors_1, plot_ncol = 4)
- Plotted RasterBrick rb_class
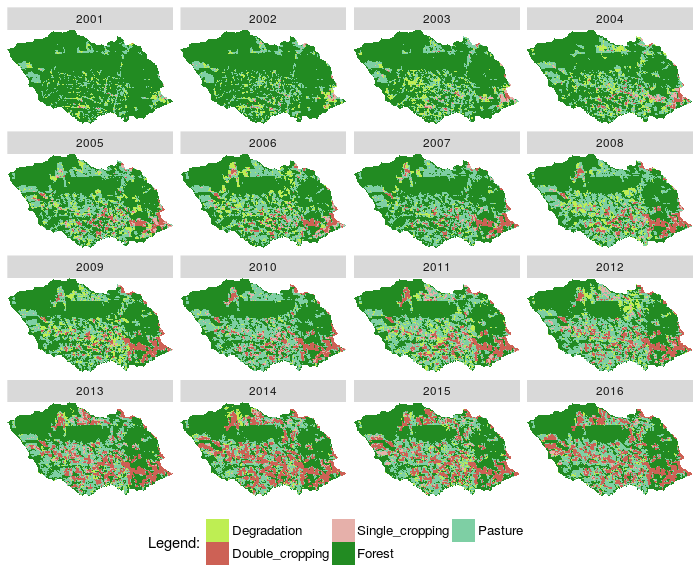
Fig. 1. Plot images classified from a RasterBrick |
- Apply lucC_pred_holds function to discover events of Degradation and Pasture in different time intervals. Parameter relation_interval = "equals" or "contains" produce different results to HOLDS. "Equals" says that all states/events must holds for each subinterval of time interval, whereas "Contains" says that one state/event can appear only once in entire time interval.
# Forest holds from 2001 to 2007
a <- lucC_pred_holds(raster_obj = rb_class, raster_class = "Degradation",
time_interval = c("2001-09-01","2007-09-01"),
relation_interval = "contains", label = label, timeline = timeline)
head(a)
# Cerrado holds from 2009 to 2013
b <- lucC_pred_holds(raster_obj = rb_class, raster_class = "Pasture",
time_interval = c("2009-09-01","2013-09-01"),
relation_interval = "equals", label = label, timeline = timeline)
head(b)
# only locations where b occurs after a - Before Allen's Relation
c <- lucC_relation_before(a, b)
# plot results
# individual locations over time
lucC_plot_sequence_events(c, custom_palette = FALSE, show_y_index = FALSE)
# barplot with result quantified
lucC_plot_bar_events(c, custom_palette = FALSE, pixel_resolution = 232, side_by_side = TRUE)
# plot RasterBrick with individual states over time
lucC_plot_raster_result(raster_obj = rb_class, data_mtx = c, timeline = timeline,
label = label, custom_palette = TRUE, RGB_color = colors_1,
relabel = FALSE, shape_point = ".", plot_ncol = 4)
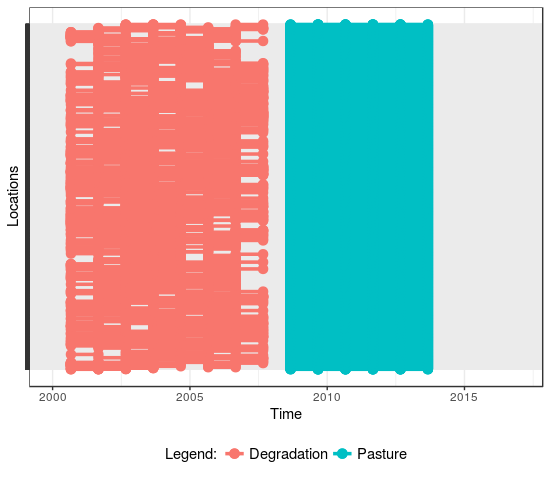
Fig. 2.(a) Locations over time |
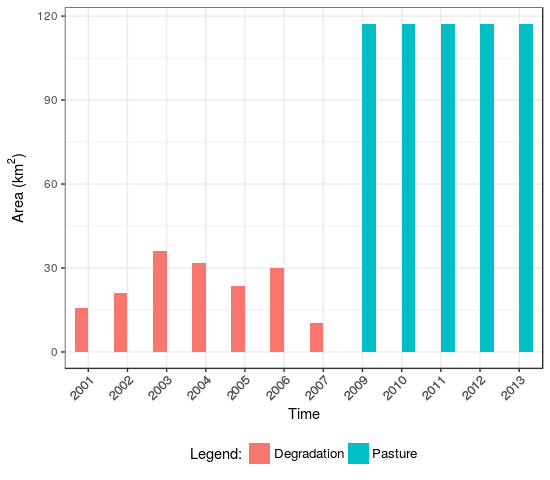
Fig. 2.(b) Barplot with number of states in km² |
- Plotted RasterBrick rb_class results
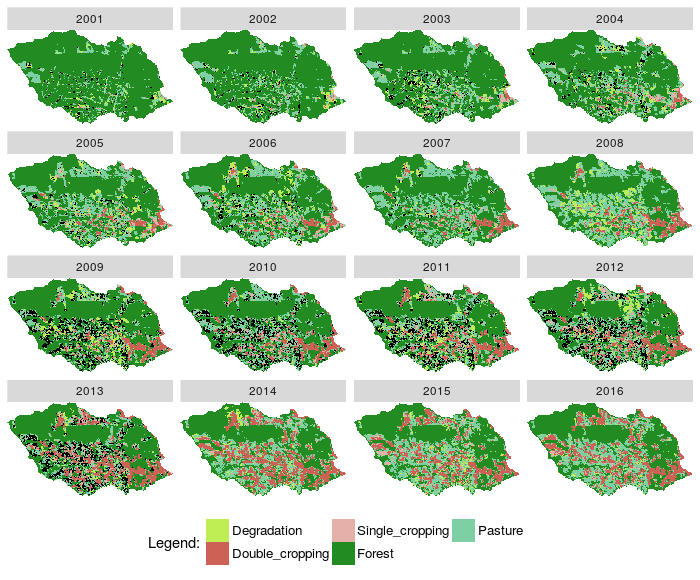
Fig. 3. Plot images classified from a RasterBrick and states from Before relation |
- Apply LUC Calculus to discover secondary vegetation from RasterBrick. We are insterested only in Forest class that RECUR after a non-sequential interval and Forest that EVOLVE after a different class in 2001. After this we update the original raster.
#----------------------------
# 1. RECUR predicate indicates a class that appear again
#----------------------------
forest_recur <- lucC_pred_recur(raster_obj = rb_class, raster_class = "Forest",
time_interval1 = c("2001-09-01","2001-09-01"),
time_interval2 = c("2002-09-01","2016-09-01"),
label = label, timeline = timeline,
remove_column = FALSE)
head(forest_recur)
#-------------------
# plot some results from RECUR
# lucC_plot_sequence_events(forest_recur, custom_palette = FALSE, show_y_index = FALSE)
# lucC_plot_bar_events(forest_recur, custom_palette = FALSE)
lucC_plot_raster_result(raster_obj = rb_class, data_mtx = forest_recur,
timeline = timeline, label = label, custom_palette = TRUE,
RGB_color = colors_1, relabel = FALSE, shape_point = ".", plot_ncol = 4)
#-------------------
- Plotted RasterBrick rb_class results
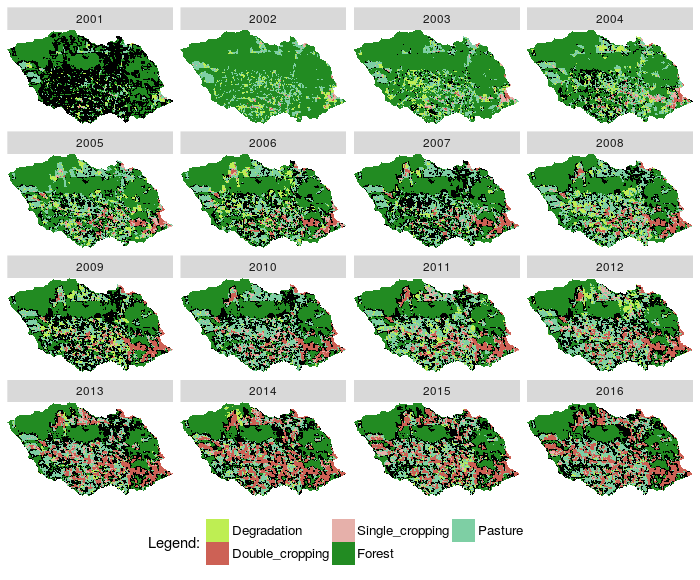

Fig. 4. Plot images classified from a RasterBrick and states from RECUR relation |
#----------------------------
# 2. EVOLVE to verify Forest class that occurs after a different class in 2001
#----------------------------
forest_evolve <- NULL
# classes without Forest based on original label
classes <- c("Degradation", "Double_cropping", "Single_cropping", "Pasture", "Pasture",
"Pasture", "Double_cropping", "Double_cropping", "Double_cropping", "Double_cropping",
"Double_cropping", "Single_cropping", "Single_cropping", "Water", "Water")
# percor all classes
for(i in seq_along(classes)){
print(classes[i])
temp <- lucC_pred_evolve(raster_obj = rb_class, raster_class1 = classes[i],
time_interval1 = c("2001-09-01","2001-09-01"), relation_interval1 = "equals",
raster_class2 = "Forest",
time_interval2 = c("2002-09-01","2016-09-01"), relation_interval2 = "contains",
label = label, timeline = timeline, remove_column = FALSE)
forest_evolve <- lucC_merge(forest_evolve, temp)
}
#-------------------
# plot some results from EVOLVE
# lucC_plot_sequence_events(forest_evolve, custom_palette = FALSE, show_y_index = FALSE)
# lucC_plot_bar_events(forest_evolve, custom_palette = FALSE, legend_text = "Legend:")
lucC_plot_raster_result(raster_obj = rb_class, data_mtx = forest_evolve,
timeline = timeline, label = label, custom_palette = TRUE,
RGB_color = colors_1, relabel = FALSE, shape_point = ".", plot_ncol = 4)
#-------------------
- Plotted RasterBrick rb_class results
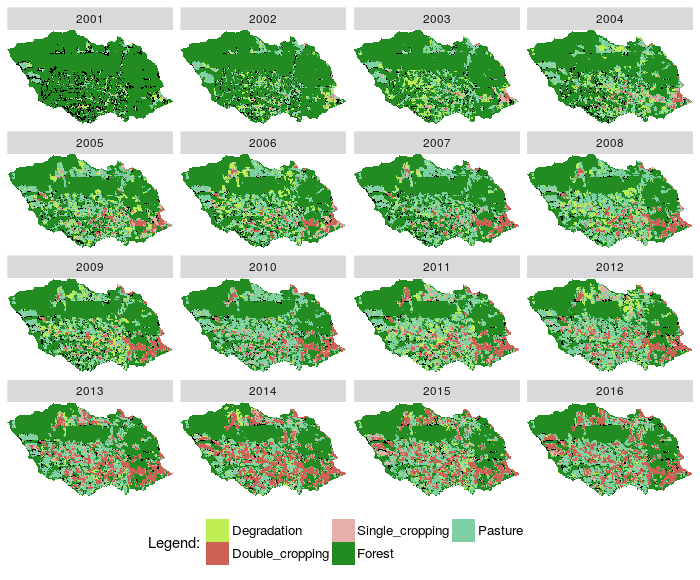
Fig. 5. Plot images classified from a RasterBrick and states from EVOLVE relation |
#----------------------------
# 3. Merge both forest_recur and forest_evolve data sets
#----------------------------
forest_secondary <- lucC_merge(forest_evolve, forest_recur)
head(forest_secondary)
# plot
lucC_plot_bar_events(forest_secondary, custom_palette = FALSE, pixel_resolution = 232, legend_text = "Legend:")
# 4. Remove column 2001 because it' is not used to replace pixels's only support column
forest_secondary <- lucC_remove_columns(data_mtx = forest_secondary, name_columns = c("2001-09-01"))
head(forest_secondary)
# plot
lucC_plot_bar_events(forest_secondary, custom_palette = FALSE, pixel_resolution = 232, legend_text = "Legend:")
# 5. Plot secondary vegetation over raster without column 2001 because it' is not used to replace pixels's only support column
lucC_plot_raster_result(raster_obj = rb_class, data_mtx = forest_secondary, timeline = timeline,
label = label, custom_palette = TRUE, RGB_color = colors_1,
relabel = FALSE, shape_point = ".", plot_ncol = 4)
- Plotted RasterBrick rb_class results of secondary vegetation
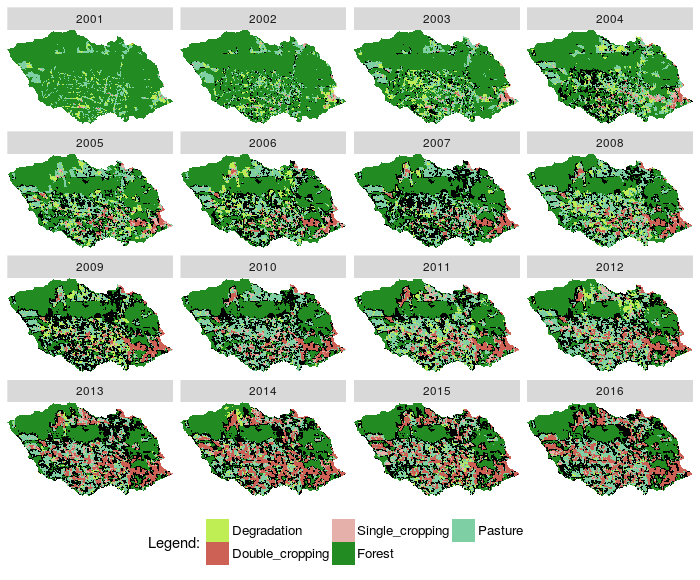

Fig. 6. Plot images classified from a RasterBrick and states from EVOLVE and RECUR relations, and also secondary vegetation locations |
- Update a raster with new value of pixel and open rasterBrick with new label.
#----------------------------
# 4. Update original raster to add new pixel value
#----------------------------
n_label <- length(label) + 1
# 1. update original RasterBrick with new class
rb_class_new <- lucC_raster_update(raster_obj = rb_class,
data_mtx = forest_secondary, # without 2001
timeline = timeline,
class_to_replace = "Forest", # only class Forest
new_pixel_value = n_label) # new pixel value
head(rb_class_new)
lucC_plot_bar_events(data_mtx = rb_class_new, pixel_resolution = 232, custom_palette = FALSE)
# 2. save the update matrix as GeoTIFF images
lucC_save_GeoTIFF(raster_obj = rb_class,
data_mtx = rb_class_new,
path_raster_folder = paste0(getwd(),"/rasterItanhangaSecVeg"),
as_RasterBrick = FALSE)
#------------
# create a RasterBrick from individual raster GeoTIFF, case saved as separate layers in lucC_save_GeoTIFF, as_RasterBrick = FALSE
lucC_create_RasterBrick(path_open_GeoTIFFs = paste0(getwd(),"/rasterItanhangaSecVeg"),
path_save_RasterBrick = getwd())
# open file RasterBrick
file <- c(paste0(getwd(),"/rasterItanhangaSecVeg.tif"))
# create timeline with classified data from SVM method
timeline <- c("2001-09-01", "2002-09-01", "2003-09-01", "2004-09-01", "2005-09-01", "2006-09-01",
"2007-09-01", "2008-09-01", "2009-09-01", "2010-09-01", "2011-09-01", "2012-09-01",
"2013-09-01", "2014-09-01", "2015-09-01", "2016-09-01")
# new variable with raster object
rb_class2 <- raster::brick(file)
# ------------- define variables to plot raster -------------
# original label - see QML file, same order and new class secondary vegetation
label2 <- c("Degradation", "Double_cropping", "Single_cropping", "Forest", "Pasture", "Pasture",
"Pasture", "Double_cropping", "Double_cropping", "Double_cropping", "Double_cropping",
"Double_cropping", "Single_cropping", "Single_cropping", "Water", "Water",
"Secondary_vegetation")
# original colors set - see QML file, same order
colors_2 <- c("#BEEE53" , "#cd6155", "#e6b0aa", "#228b22", "#7ecfa4", "#1e174d", "#afe3c8",
"#64b376", "#e1cdb6", "#b6a896", "#b69872", "#b68549", "#9c6f38", "#e5c6a0",
"#e5a352", "#0000ff", "#3a3aff")
# plot rasterBrick
lucC_plot_raster(raster_obj = rb_class2,
timeline = timeline, label = label2,
custom_palette = TRUE, RGB_color = colors_2, plot_ncol = 4)
- Plotted RasterBrick rb_class results with new class Secondary Vegetation
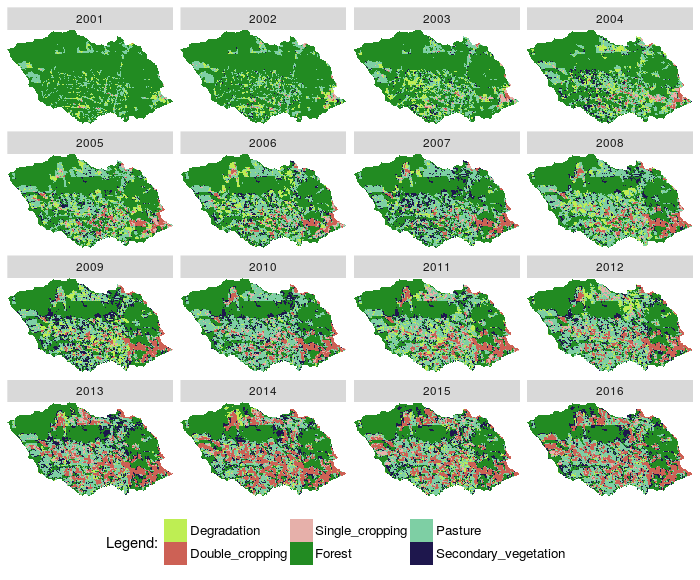
Fig. 7. Plot a RasterBrick and states with new secondary vegetation class |
- Difference between Forest and Secondary Vegetatation
#----------------------------
# 5. Discover Forest and Secondary vegetation - LUC Calculus
#----------------------------
secondary.mtx <- lucC_pred_holds(raster_obj = rb_class2, raster_class = "Secondary_vegetation",
time_interval = c("2001-09-01","2016-09-01"),
relation_interval = "contains", label = label2, timeline = timeline)
head(secondary.mtx)
forest.mtx <- lucC_pred_holds(raster_obj = rb_class2, raster_class = "Forest",
time_interval = c("2001-09-01","2016-09-01"),
relation_interval = "contains", label = label2, timeline = timeline)
head(forest.mtx)
Forest_secondary.mtx <- lucC_merge(secondary.mtx, forest.mtx)
head(Forest_secondary.mtx)
# plot results
lucC_plot_bar_events(data_mtx = Forest_secondary.mtx, custom_palette = TRUE,
RGB_color = c("black", "gray60"), #c("#228b22", "#7ecfa4"),
pixel_resolution = 231.656, side_by_side = TRUE,
relabel = TRUE, original_labels = c("Forest", "Secondary_vegetation"),
new_labels = c("Forest", "Secondary vegetation"))
# Compute values
measuresFor_Sec <- lucC_result_measures(data_mtx = Forest_secondary.mtx, pixel_resolution = 231.656)
measuresFor_Sec
#------------
Years Classes Pixel_number Area_km2 Cumulative_Sum Relative_Frequency Cumulative_Relative_Frequency
1 2001 Forest 46647 2503.2880404674 2503.2880404674 10.463261831648 10.463261831648
2 2002 Forest 43152 2315.7306048031 4819.0186452705 9.679307877447 20.142569709096
3 2003 Forest 38585 2070.6448226346 6889.6634679050 8.654896515835 28.797466224931
4 2004 Forest 33751 1811.2306183423 8700.8940862474 7.570595109653 36.368061334583
5 2005 Forest 29849 1601.8317302273 10302.7258164746 6.695348091257 43.063409425841
6 2006 Forest 27909 1497.7225956954 11800.4484121700 6.260191962173 49.323601388013
7 2007 Forest 27226 1461.0697405999 13261.5181527700 6.106990087861 55.430591475875
8 2008 Forest 25062 1344.9397575448 14606.4579103148 5.621589127377 61.052180603252
9 2009 Forest 24618 1321.1127185076 15927.5706288225 5.521996693711 66.574177296963
10 2010 Forest 24134 1295.1390993770 17222.7097281995 5.413431968723 71.987609265685
11 2011 Forest 22913 1229.6147420248 18452.3244702243 5.139552776139 77.127162041824
12 2012 Forest 21345 1145.4688023619 19597.7932725862 4.787838956343 81.915000998167
13 2013 Forest 20781 1115.2020230444 20712.9952956306 4.661329648712 86.576330646880
14 2014 Forest 20236 1085.9548692713 21798.9501649019 4.539082179459 91.115412826339
15 2015 Forest 19879 1066.7966419373 22865.7468068392 4.459004479416 95.574417305756
16 2016 Forest 19730 1058.8006310893 23924.5474379285 4.425582694244 100.000000000000
17 2001 Secondary_vegetation 0 0.0000000000 0.0000000000 0.000000000000 0.000000000000
18 2002 Secondary_vegetation 624 33.4866494577 33.4866494577 0.870256474624 0.870256474624
19 2003 Secondary_vegetation 1244 66.7586409060 100.2452903636 1.734934382104 2.605190856728
20 2004 Secondary_vegetation 2781 149.2409809964 249.4862713601 3.878498807581 6.483689664310
21 2005 Secondary_vegetation 2858 153.3731476763 402.8594190364 3.985886225123 10.469575889433
22 2006 Secondary_vegetation 4259 228.5571154490 631.4165344854 5.939779367669 16.409355257102
23 2007 Secondary_vegetation 6334 339.9109577962 971.3274922816 8.833661074153 25.243016331255
24 2008 Secondary_vegetation 3902 209.3988881151 1180.7263803967 5.441892249976 30.684908581231
25 2009 Secondary_vegetation 7631 409.5138173260 1590.2401977227 10.642511470929 41.327420052160
26 2010 Secondary_vegetation 5969 320.3234144436 1910.5636121663 8.324616822169 49.652036874329
27 2011 Secondary_vegetation 4785 256.7846436778 2167.3482558440 6.673360947241 56.325397821570
28 2012 Secondary_vegetation 5551 297.8916524671 2465.2399083112 7.741656555514 64.067054377083
29 2013 Secondary_vegetation 6753 362.3963842750 2827.6362925862 9.418015982595 73.485070359678
30 2014 Secondary_vegetation 6068 325.6362001748 3153.2724927610 8.462686359009 81.947756718687
31 2015 Secondary_vegetation 6417 344.3651114901 3497.6376042511 8.949416342412 90.897173061099
32 2016 Secondary_vegetation 6527 350.2682067471 3847.9058109982 9.102826938901 100.000000000000
- Quantity of Forest and Secondary vegetation after LUC Calculus application
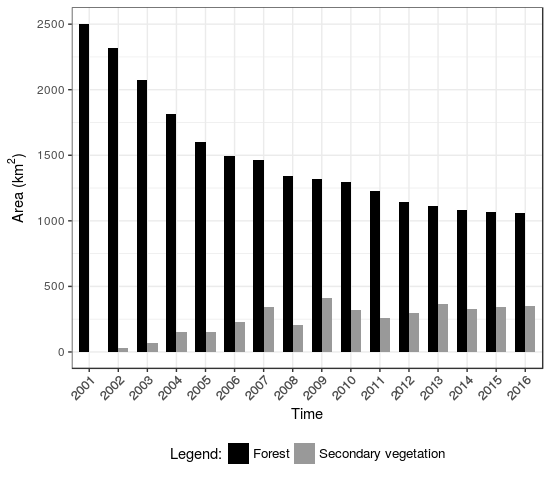
Fig. 8. Number of Forest and Secondary Vegetation |
- Difference between Forest and Secondary Vegetatation
#----------------------------
# 5. Discover direct land use transitions
#----------------------------
class1 <- c("Forest")
classes <- c("Degradation", "Double_cropping", "Pasture", "Single_cropping")
direct_transi.df <- NULL
# along of all classes
for(x in 2:length(timeline)){
t_1 <- timeline[x-1]
t_2 <- timeline[x]
cat(paste0(t_1, ", ", t_2, sep = ""), "\n")
# moves across all classes
for(i in seq_along(classes)){
cat(classes[i], collapse = " ", "\n")
temp <- lucC_pred_convert(raster_obj = rb_class2, raster_class1 = class1,
time_interval1 = c(t_1,t_1), relation_interval1 = "equals",
raster_class2 = classes[i],
time_interval2 = c(t_2,t_2), relation_interval2 = "equals",
label = label2, timeline = timeline)
direct_transi.df <- lucC_merge(direct_transi.df, temp)
}
cat("\n")
}
Forest_others <- direct_transi.df
head(Forest_others)
str(Forest_others)
# plot results
lucC_plot_frequency_events(data_mtx = Forest_others,
pixel_resolution = 232, custom_palette = FALSE)
lucC_plot_bar_events(Forest_others, custom_palette = FALSE, pixel_resolution = 232,
legend_text = "Legend:", side_by_side = FALSE)
# replace classes by new legend
Forest_others[c(3:ncol(Forest_others))] <-
ifelse(Forest_others[c(3:ncol(Forest_others))] == "Degradation", "Forest_Degradation",
ifelse(Forest_others[c(3:ncol(Forest_others))] == "Double_cropping", "Forest_Double_cropping",
ifelse(Forest_others[c(3:ncol(Forest_others))] == "Pasture", "Forest_Pasture",
ifelse(Forest_others[c(3:ncol(Forest_others))] == "Single_cropping", "Forest_Single_cropping"
, ""))))
lucC_plot_bar_events(Forest_others, pixel_resolution = 231.6465, custom_palette = TRUE,
RGB_color = c("#6e8b3d", "#a2cd5a", "#006400", "#b4eeb4"),
legend_text = "Land use transitions:", relabel = TRUE,
original_labels = c("Forest_Degradation", "Forest_Double_cropping", "Forest_Pasture", "Forest_Single_cropping"),
new_labels = c("Degradation", "Forest to Double Cropping", "Forest to Pasture", "Forest to Single Cropping"),
side_by_side = FALSE)
- Direct land use transition from Forest class to others in Itanhangá municipality
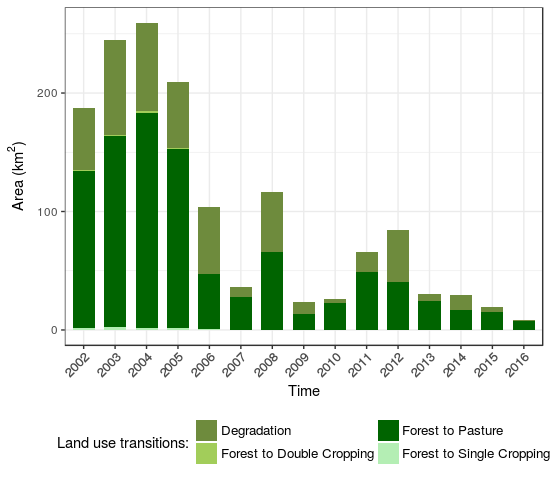
Fig. 9. Quantity of direct land use transitions |